Introduction
Maize genetic diversity is very important and farmers use it to create new varieties (Kato et al. 2009). One maize variant that can be used for this purpose is the polyembryony phenomenon (PEm), which was generated through a natural mutation, because of its high productive potential and grain nutritional quality (Castro and Rodriguez 1979). These benefits are obtained under the hypothesis that two or more embryos per seed favors the increase of production, both in the number of plants and ears, allowing to increase grain storage of nutrients of quality because of its greater number of embryos (Espinoza et al. 1998, González-Vázquez et al. 2011). Polyembryony was discovered by Leewenhoek in 1719 (Batygina and Vinogradova 2007), who observed formation of two seedlings from the same seed in citrus Moreover, formation of additional embryos may result from differentiation and development of various tissues, zygotes, synergies, antipodes, nucella or tegument (Peter 2009).
The Mexican Maize Institute at Universidad Autónoma Agraria Antonio Narro (IMM-UAAAN) has performed different researches on polyembryony since the 1970s development of two populations that have this characteristic with PEm on the average frequencies from 55 to 65% (González-Vázquez et al. 2011, Espinoza-Velázquez et al. 2012). The inheritance of this mutant has been identified as the one corresponding to two loci with epistatic interaction of the dominant double type, which segregates in F2 in a proportion of 15 cases that germinate in individual plant: 1 case of double or higher number of plants (polyembryonic) associated to the phenomenon of incomplete penetrance (Rebolloza-Hernández et al. 2011). From this study, it can be assumed that the PEm population could contain some type of apomictic reproduction since polyploidy and polyembryony regularly accompany this type of asexual reproduction. In a study carried out by Avendaño et al. (2015) using the IMM-UAAAN polyembrynic populations, it was found similarity between DNA sequences of the internal transcribed spacers (ITS) of the female parent and one of the PEm progenies, which might suggest a probable relationship between polyembryony and apomixis. If this hypothesis is corroborated in one of the PEm populations, genetic material could be available for studies and special reproductive applications in maize.
Correlation of phenotypes with molecular markers (MM) provides reference points for elucidation of genetic variation in plants (Kumar et al. 2014). There are several MM that allow to determine structure, dynamics, relation and genetic variability of a population, and to estimate the degree of divergence between individuals. The most useful MM have been those based on the polymerase chain reaction (PCR) such as amplification of short-sequence DNA repeats (SSR) hypervariable regions, internal transcribed spacers (ITS) and intergenic spacers (IGS), among others (Rodriguez-Tovar et al. 2004, Kirst et al. 2005). The ITS and IGS regions have been used mainly in cases of biparental inheritance, and are suitable for comparisons between closely related species and genera (Alvarez and Wendel 2003). On the other hand, the SSR have applications in plant genetic studies, such as genetic traces of crops, diversity studies and germplasm selection. The aim of this work was to analyze and validate the genetic similarity, through the use of hypervariable regions and SSR microsatellites among members of maize families with PEm nature to obtain indicators of molecular variation and clustering relationship, with the purpose of obtaining evidence to corroborate the hypothesis of probable relation between apomixis and polyembryony.
Materials and methods
Plant material
Genetic material was obtained from four families of S1 inbred lines (D-Si) and a family of open pollination (D-PL), derived from the UAAAN-IMM-BAP maize population, which is dwarf (br-2 brachytic gene) with an average of polyembryony of 60% generated in the spring-summer/2010 cycle, in Buenavista, Saltillo, Mexico. Seed samples from three families (D-S1 - 03, D-S1-05 and D-PL-13) were germinated during spring/summer-2013 to obtain foliar tissue and DNA samples. The other two families (D-Si-03* and D- Si-07*) correspond to the remaining DNA sample, which was extracted by Avendano et al. (2015).
Each family was composed of the female parent and three progeny plants (one individual, and one pair of polyembryonic PEm plants, these originated from a single seed). The families D-S1 -03 and D- S1 -03* came from the same female parent, sampled at different time. For molecular variance and principal coordinate analyses, as reference material of the common corn (No-PEm), the high inbreeding line (AN-Tep-3) and the Tuxpeno-HOC population (high oil corn) were used because of their genetic distance in relation to the PEm genotypes.
DNA isolation and ITS amplification
DNA extraction was performed from leaf tissue of 17-day old PEm maize seedlings using the method described by Doyle and Doyle (1987) with some modifications, such as: addition of 0.5% (v/v) 2- mercaptoethanol to the CTAB extraction buffer. Concentration and quality of DNA was measured in and EpochTM Micro-Volume System Spectrophotometer. Extracted DNA was visualized by electrophoresis in 1% (w/v) agarose gels and stained with ethidium bromide (0.5 pg mL-1). The ITS region of the 18S-26S ribosomal DNA was amplified to determine genetic similarity using the primers pair: N18L-F (5’-AAGTCGTAACAAGGTTTCCGTAGGTG-3’) and C26A-R (5’-TTTCTTTTCCTCCGCT-3’) (Eldenäs et al. 1998). Furthermore, the IGS region of ribosomal DNA was amplified with the primers pair: IGS- F (5’-CTGAACGCCTCTAAGTCAG-3’) and IGS-R (5’- GAGACAAGCATATGACTACTG-3’) (Kim et al. 2001).
Amplification of ITS and IGS regions of the four D-S1 and one D-PL families was carried out in a total volume of 25 µL, composed of 14.5 and 17.0 µL of ddH2O, 1X and 0.4X reaction buffer added with MgCl2, respectively, 0.2 mM of dNTPs, 0.8 µM of each primer and 2.0 µL of DNA. PCR amplification was performed in a Px2 Thermal cycler using the following program: 5 min of initial denaturation at 94 °C; 35 cycles of 1 min at 94 °C, 1 min at 50.3 °C for ITS and 51.3 °C for IGS, and 1 min at 72 °C; and a final extension of 5 min at 72 °C. The amplified products were visualized in 1% (w/v) agarose gels and fragment size was estimated using a 100 µb DNA Ladder (Invitrogen®).
Sequence analysis. Sequencing of PCR products were performed on a Perkin Elmer Applied Biosystem Model 3730, using the fluorescence terminator method (Taq FS Dye Terminator Cycle Sequencing Fluorescence-Based Sequencing). Alignments were carried out in BioEdit version 7.1.3 software, and the percentage of invariant sites (monomorphic) and G+C content were determined using DnaSP (DNA sequence polymorphism version 5.10.01) (Librado and Rozas 2009), by pair comparison of individuals in each family including the female parent plant and three descendents (one individual and a pair of polyembryonic plants PEm).
SSR analysis
In the SSR analysis, ten maize microsatellite loci, one of each chromosome, were obtained from the MaizeGDB database (Table 1). Each locus was selected because of the high PIC (polymorphism information content) presented in a study by Warburton et al. (2002). Reactions were performed in a cocktail composed as following: 1.5 to 3X buffer with MgCl2, 0.25 mM of dNTPs, 0.66 to 2 pM of each primer, 6.6% (v/v) of DMSO. 0.1 U of PaqDNA Polymerase, 1.5 pL of DNA and ddH2O up to a final volume of 15 pL. Warburton et al. 2002 described the reference amplification protocol with an initial denaturation step of 2 min at 94 °C; 30 cycles of 30 s at 94 °C, 1 min of annealing (Table 1) and 1 min of polymeration at 72 °C; followed of final extension step at 72 °C for 5 min.
Table 1: Simple Sequence Repeats (SSR) markers used in present study, with their genomic position, annealing temperature and sequence.
| Primer name | Bin† | PIC †† | Alignment temperature (°C) | Sequence (5’ ( 3’) |
| phi064 f | 1.11 | 0.79 | 56.0 | CCGAATTGAAATAGCTGCGAGAACCT |
| phi064 r | ACAATGAACGGTGGTTATCAACACGC | |||
| phi96100 f | 2.0 | 0.75 | 50.0 | AGGAGGACCCCAACTCCTG |
| phi96100 r | TTGCACGAGCCATCGTAT | |||
| phi053 f | 3.05 | 0.70 | 60.0 | CTGCCTCTCAGATTCAGAGATTGAC |
| phi053 r | AACCCAACGTACTCCGGCAG | |||
| phi072 f | 4.01 | 0.59 | 56.0 | ACCGTGCATGATTAATTTCTCCAGCCTT |
| phi072 r | GACAGCGCGCAAATGGATTGAACT | |||
| zct118 f | 5.07 | 0.76 | 56.0 | CTTCCAGCCGCAACCCTC |
| zct118 r | CCAACAACGCGGACGTGA | |||
| phi452693 f | 6.06 | 0.57 | 52.4 | CAAGTGCTCCGAGATCTTCCA |
| phi452693 r | CGCGAACATATTCAGAAGTTTG | |||
| phi328175 f | 7.04 | 0.67 | 60.0 | GGGAAGTGCTCCTTGCAG |
| phi328175 r | CGGTAGGTGAACGCGGTA | |||
| phi420701 f | 8.01 | 0.56 | 60.0 | ATGTTTCAAAACCACCCAGA |
| phi420701 r | ATGGCACGAATAGCAACAGG | |||
| phi032 f | 9.04 | 0.48 | 56.0 | CTCCAGCAAGTGATGCGTGAC |
| phi032 r | GACACCCGGATCAATGATGGAAC | |||
| umc1152 f | 10.01 | 0.74 | 60.0 | CCGAAGATAACCAAACAATAATAGTAGG |
| umc1152 r | ACTGTACGCCTCCCCTTCTC |
†Bin indicates the chromosomal location. †† PIC is polymorphic information content.
Data analyses
The amplification pattern of PCR products was transformed into a binary matrix, where 1 indicates presence and 0 = absence. From these results, the homozygous and heterozygous condition of each loci was determined. Pattern of similarity was sought between the female parent plant and each progenie seedling, one of the individual type and the polyembryonic pair (PEm). An analysis of molecular variance (AMOVA) was performed, where five groups were considered, four belonging to each of the PEm families (D-Si-03, D-PL-13, D-Si-03* and D-Si-07*) and the fifth group corresponding to the reference material. A Principal Coordinate Analysis (PCoA) and a dendrogram were performed in order to observe the variability and to group the studied genotypes. All the statistical analyzes were carried out using INFOGEN software (v. 2011).
Results
Comparison of the hypervariable regions nucleotide sequences in families D-S1 (first selffertilization), and D-PL (open pollination)
Frequencies invariable sites in the ITS region of D-S1 families did not show cases of identical sequences in the six possible comparisons per maize family, but the D-S1 -07* family had very high similarity values (96 to 99%). D-S1 -05 Family showed the highest variability in the sites, indicating absence of identical sequences among its members, being discarded for the rest of analyzes. In the series of comparisons of D-S1 -03 and D-S1 -03* families, the highest percentage of similarity was showed between the pair of polyembryonic sister plants (PEm1 and PEm2). In general, the families analyzed had widely divergent values, from 39 to 99%, where the highest values were mostly for the comparison between PEm sisters plants, and not for the female parent (M) with one of the PEm daughters (Figure 1), which is a contrary condition from a previous report. As in the ITS region of the D-S1 -03 and D-S1 -03* families, the highest value of monomorphic sites in the IGS region of the D-S1-07* family was presented in comparison between the pair of polyembryonic sister plants (PEm1 and PEm2) (Figure 2).
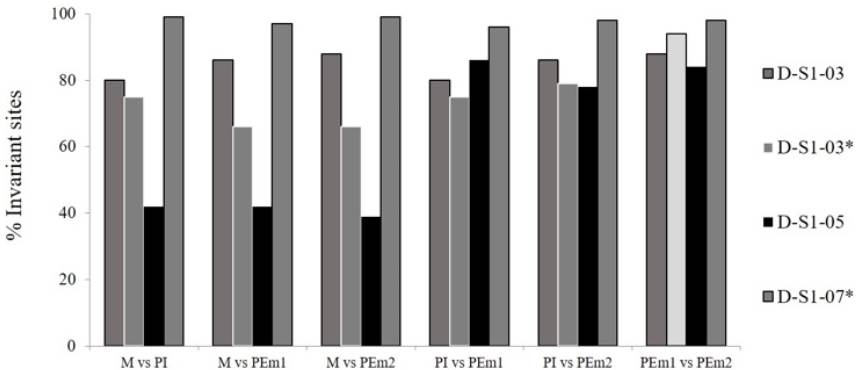
Figure 1: Percentage of invariant sites in ITS region sequence in D-S1 families. M: female parent plant, PEm: each of the two polyembryonic plant, PI: individual plant.

Figure 2: Percentage of invariant sites of IGS region sequence in D-S1-07* family. M: female parent plant, PEm: polyembryonic plant, PI: individual plant.
The D-PL-13 family, generated by open- pollination (PL) showed high percentages of monomorphic sites in the six comparisons of the ITS region (Figure 3). However, the comparison analysis of the IGS region in the genotypes of this family were contrary to those obtained in the ITS analysis (Figure 4). Morphology (phenotype) of the polyembryonic seedlings corresponding to the D-PL-13 family consisted of two mesocotyls and two completely separate radicles.

Figure 3: Percentage of invariant sites in ITS region sequence of D-PL-13 family. M: female parent plant, PEm: polyembryonic plant.

Figure 4: Percentage of invariant sites in IGS region sequence of D-PL-13 family. M: female parent plant, PEm: polyembryonic plant, PI: individual plant.
SSR analysis
The genotypes included in this study were the two families that showed high sequence similarity (99 to 100%) in the ITS region. Results of the similarity analysis in those families are presented in Table 2. Families with the highest loci similarity were D-Si- 03, D-Si-03* and D-PL-13. In the D-Si-03 family, the female parent plant and the individual daughter showed 100% homozygous loci, likewise in the D- S1 -03* family, comparisons among the three sister plants (the possible comparisons among PI, PEm1 and PEm2) presented 100% homozygous loci, and the female parent plant only 67%. Similarly, the D-PL- 13 family showed high similarity (100%) in only the two polyembryonic sister plants (PEm1 and PEm2), and the percentage of homozygous loci of the female parent plant was equal or less to 50%.
Table 2: Similarity analysis of homozygous and heterozygous loci in maize families, within the context of polyembryony.
| Comparisons | D-S1-03 (%) | D-PL-13 (%) | (Avendaño et al 2015) | |
| D-S103* (%) | D-S1-07* (%) | |||
| M vs PI | 100 | 25 | 67 | 50 |
| M vs PEm1 | 83 | 50 | 67 | 67 |
| M vs PEm2 | 83 | 50 | 67 | 83 |
| PI vs PEm1 | 83 | 50 | 100 | 83 |
| PI vs PEm2 | 83 | 50 | 100 | 67 |
| PEm1 vs PEm2 | 83 | 100 | 100 | 83 |
M: mother plant, PEm: polyembryonic plant, PI: individual plant.
Results of the molecular variance analysis (AMOVA) showed significant differences (p < 0.01) among the five analyzed groups (Table 3). The highest percentage of genetic variation was due to difference within members of each group, and the restwas due to differences among groups. The ordering of polyembryonic and exotic maize samples that yielded the Principal Coordinate Analysis (PCoA), performed with the Anderberg distance (cophenetic correlation of 0.966), is presented in Figure 5. The ordering consisted of formation of four groups, one containing the AN-Tep-3 and Tuxpeno HOC genotypes, both representing common or normal maize genotypes, and genetically distant from PEm-like families. The second group consisted of the D-S1 -03 family and a member of the D-PL-13; the third group included the D-S1 -07* family and a member of DPL-13 family and finally, the fourth group consisted of the D-S1 -03* family and the rest of the individuals in the D-PL-13 family.
Table 3: Analysis of Molecular Variance (AMOVA) within and between polyembryonic and exotic families.
| Variation source | DF | SS | MS | Variation Percentage | p-value | Number of Iterations |
| Groups | 4 | 95.1 | 23.78 | 26.2 | 0.0075 | 400 |
| Within groups | 13 | 138.0 | 10.6 | 73.8 | 0.0025 | 400 |
| Total | 17 | 233.1 | 13.71 |

Figure 5: Principal Coordinate Analysis of polyembryonic and exotic genotypes. M: female parent plant, PEm: polyembryonic plant, PI: individual plant, *: material generated by Avendaño et al. (2015), AN-Tep-3 y Tuxp HOC: exotic materials Tepalcingo and Tuxpeño, respectively.
The dendrogram to identify the grouping of the evaluated genotypes was obtained with the UP- GMA method (Unweighted Pair-Group Methods with Arithmetic Mean), which is shown in Figure 6. The most remote group corresponded to the AN-Tep-3 and Tuxpeno HOC genotypes. The second subgroup was formed with the two polyembryonic plants of the same family (D-S1 -03 PEm1 and D-S1 -03 PEm2). In the third group, two polyembryonic plants of D-S1 - 07 family were found to form a subgroup (D-S1 -07 PEm1* and D-S1 -07 PEm2*), with a distance of 0.50 and in the fourth group, a subgroup with a distance of 0.32 between a single plant and a polyembryonic (D-Si-03 PI* and D-Si-03 PEm1*) was formed, the genotypes being considered more closely related.
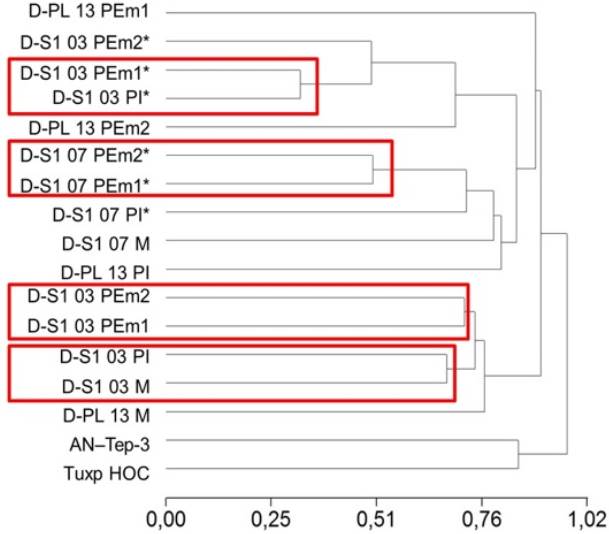
Figure 6: Dendrogram of polyembryonic and exotic maize genotypes based on Anderberg distance measurements. M: female parent plant, PEm: polyembryonic plant, PI: individual plant, *: material generated by Avendaño et al. (2015), AN-Tep-3 and Tuxp HOC: exotic materials Tepalcingo and Tuxpeño, respectively.
Discussion
The high levels of similarity among members comparison of the D-S1-07* family in the ITS region suggested the possibility of corroborating the results with an analysis of the IGS region; however, application of the analysis showed significant divergences (Figure 2), with values well below those obtained in the ITS region. A possible explanation is the rapid evolution of these regions, where it is common manifestation of polymorphism (Zhou et al. 1996). In a review by Willson and Burley (1983), referred to simple polyembryony as the ability of the female game-tophyte to present more than one egg cell with the potential to produce an embryo after being fertilized by the same or different male, that will depend on the source of pollen grains (Tudge 2006). With the values obtained in the present research for monomorphic sites in the IGS region, it can be supported the fact that PEm1 and PEm2 plants are not considered as identical, and therefore not come from the same origin, even in the case when pollination is controlled and the degree of similarity could come from the accumulation of homozygous genes generated by self-fertilization in an expected proportion of 50%, as is the case of the S1 families.
High percentages of monomorphic sites in the ITS region of an open-pollination (PL) generated family were found. That is to say, as if the nucleotide sequence of each of the members, female parent plant and its progenies, were the same and some type of asexual reproduction (apomixis) could exist since the progeny, product of open pollination does not contain the forced accumulation of gene pairs in homozygous by effect of self-fertilization, a condition that, when observed, would increase the likelihood of similarity, at least among the DNAs of sister plants. But, it can be concluded that existence of some type of apomictic reproduction is not confirmed according to comparison analysis of the IGS region.
Morphology of polyembryonic seedling was similar to one of the three polyembryonic origin versions described by Erdelska (1996), where multiple individual embryos were produced closely together, but separated by an epidermis, with independent plumules and radicles, generated by the phenomenon called polyspermia, where the embryos originate from the fertilization of multiple egg cells. One of the probable causes that may inflict the sequence similarity in the ITS region in this study could be due to similar genetic condition of the pollen source, with common genetic basis. In addition, the percentage of G+C content in all the studied families was presented in a range of 46 to 76%, which coincides with that reported for maize composition, in a range from 45 to 75% (Carels and Bernardi 2000).
Regarding SSR analysis, homozygous loci increased by the reduction of heterozygous sites because of self-fertilization, as was in the case of three Si families. On the other hand, it was expected that in the genotypes with open pollination (PL), the progeny would present the majority of loci in heterozygous condition with respect to the female parent plant. However, this was not observed in the D-PL-13 family, which may be explained because families derived from the D population contained a common genetic base. Moreover, asexual reproduction is strongly correlated with hybridization, since most species that are apomictic emerged from the hybridization of two genomes, having heterozygous loci and, because of epigenetic changes, heterozygous was perpetuated (Beck et al. 2012, Lovell et al. 2013).
According to the molecular variance analysis, higher genetic variation was found within members of the five analysed groups. In a study by Ramakrishnan et al. (2014) in bud explants of 17 maize genotypes, they determined an amount of variation of 93% within genotypes and 3% between genotypes, concluding that the female parent and the seedlings generated in vitro have a 100% genetic similarity. Based on the results of this study, it is possible to indicate that there was variation among members of the families, and even more in the group that corresponded to the exotic genotypes, corresponding to the open-pollinated Tuxpeno HOC population and the inbred line AN-Tep-3, which correspond to the common maize type, or Non-PEm.
In the Principal Coordinate Analysis, the D-Si- 03 and D-S1 -07* families had a better ordering, which was consistent with the close relationship of the percentages of invariant sites in the ITS region. On the other hand, the individuals that were less ordered were those of the D-PL-13 family, which was originated by open pollination. Results of the three performed analyzes in the D-S1 -03 and D-S1 -07 families, either PCoA, dendrogram or sequence comparison of the ITS region, coincided in detecting the close relationship that exists between the members of the families and even more so between the two polyembryonic sister plants (PEm1 and PEm2). On the other hand, the D-PL-13 family corroborated the divergence among the members that formed it, even between the two PEm plants generated in the same seed, a result that suggests that this pair of plants, initially considered as “twins”, differ in some degree in their genetic condition, that is, they are non-identical sisters.
Conclusions
The use of molecular techniques applied in this research to genotypes derived from UAAAN-IMM- BAP maize population allowed to demonstrate that reproduction of the studied maize plants is of a sexual type, and that based on the molecular markers used, no evidence was obtained about the probable relationship of a common genetic basis between polyembryony and apomixis. The high similarity detected in the analysis of the ITS hyper-variable regions is not sufficient evidence to support an identical genetic condition between the female parent plant and one of the PEm progenies, and even among the PEm sister plants pair. However, the highest but not complete similarity was observed in all cases between the pair of polyembryonic plants, emerged from a single seed. Sequencing of the ITS and IGS regions, as well as the use of SSR microsatellites of different chromosomes, was a practical and economical tool for the assess ment of similarity between genotypes.











 text new page (beta)
text new page (beta)



