The Neotropical woody bamboo clade is a lineage comprising three subtribes (Arthrostylidiinae, Chusqueinae, and Guaduinae) within the Bambuseae tribe, which belongs to the Bambusoideae subfamily of the Poaceae family (Sungkaew et al. 2009, BPG 2012, Kelchner et al. 2013, Clark et al. 2015). Olmeca Soderstr. is one of the six genera belonging to the Guaduinae subtribe, alongside Apoclada McClure, Eremocaulon Soderstr. & Londoño, Guadua Kunth, Otatea (McClure & E.W. Sm.) C.E. Calderón & Soderstr., and Tibisia C.D. Tyrrell, Londoño & L.G. Clark. It includes five described species (Ruiz-Sanchez et al. 2011, Tyrrell et al. 2018, Ruiz-Sanchez et al. 2021).
The geographical distribution of Olmeca ranges from east-central Mexico (Veracruz) to northern Honduras (Ruiz-Sanchez et al. 2011, 2020). Four of the five Olmeca species are endemic to Mexico (Ruiz-Sanchez et al. 2020). Two species, O. recta Soderstr. and O. reflexa Soderstr., are distributed in southern Veracruz, Oaxaca, and Chiapas. They inhabit tropical perennial forests and produce bacoid caryopses (Figure 1; Rzedowski 1978, Soderstrom 1981, Ruiz-Sanchez et al. 2011, Ruiz-Sanchez & Sosa 2015, Ruiz-Sanchez et al. 2017, 2020). Both species have been recorded in sympatry (Ruiz-Sanchez et al. 2011), in the region known as Uxpanapa in southern Veracruz. The other three Olmeca species- O. clarkiae (Davidse & R.W. Pohl) Ruiz-Sanchez, Sosa & Mejía-Saulés, O. fulgor (Soderstr.) Ruiz-Sanchez, Sosa & Mejía-Saulés, and O. zapotecorum Ruiz-Sanchez, Sosa & Mejía-Saulés-are distributed in Veracruz, Oaxaca, and Chiapas in Mexico, with O. clarkiae also found in Chiapas and Lempira, Honduras. They inhabit montane cloud forests and produce typical caryopsis fruits (Figure 2; Rzedowski 1978, Ruiz-Sanchez et al. 2011).
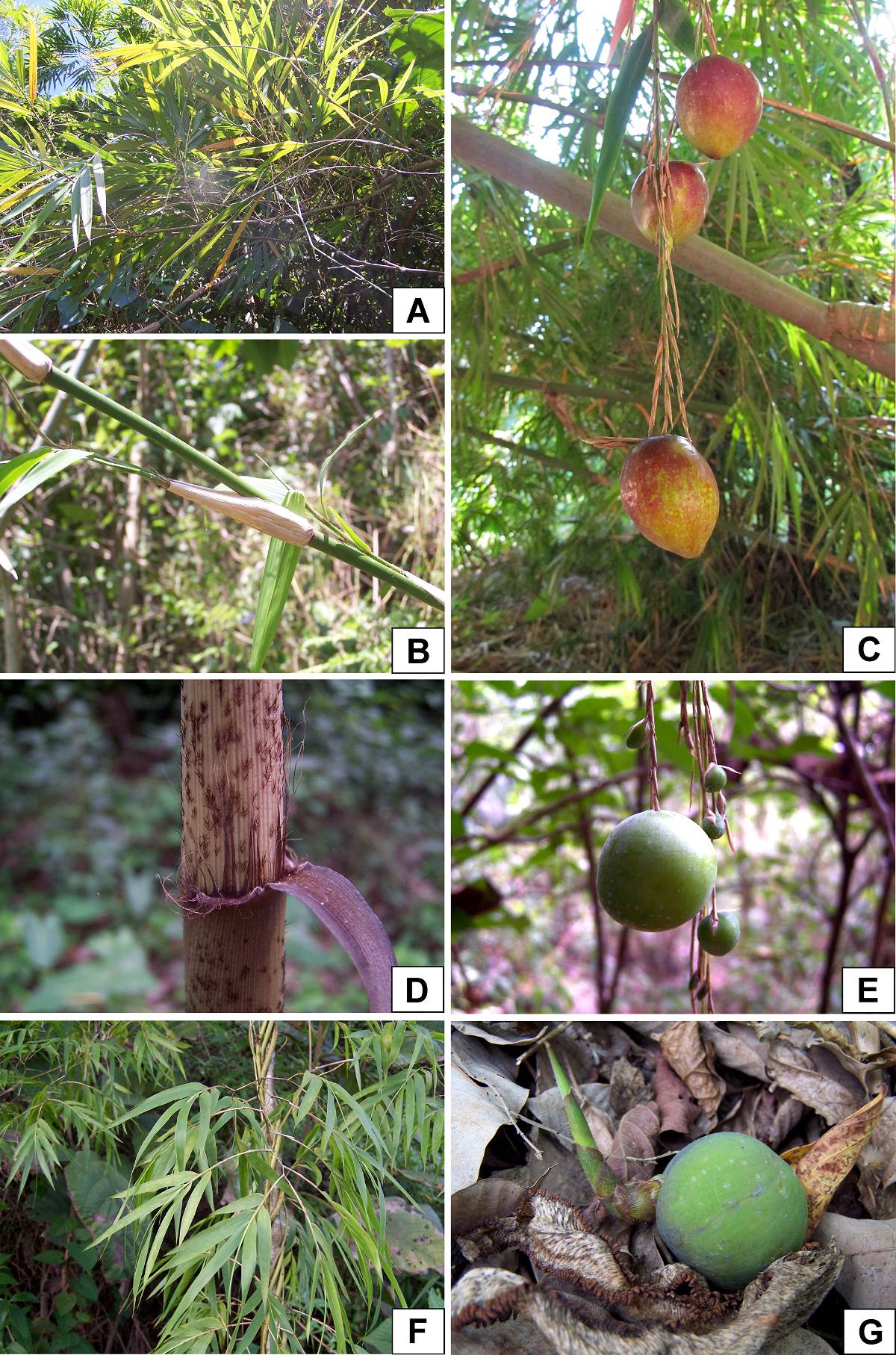

Figure 1 A-C. Olmeca recta. A. Foliage leaves. B. Young culm showing branching. C. Bacoid fruits. D-G. Olmeca reflexa. D. Culm leaf, showing reflexed blade. E. Bacoid fruits in distinct maturation periods. F. Foliage leaves. G. Germinating bacoid fruit.
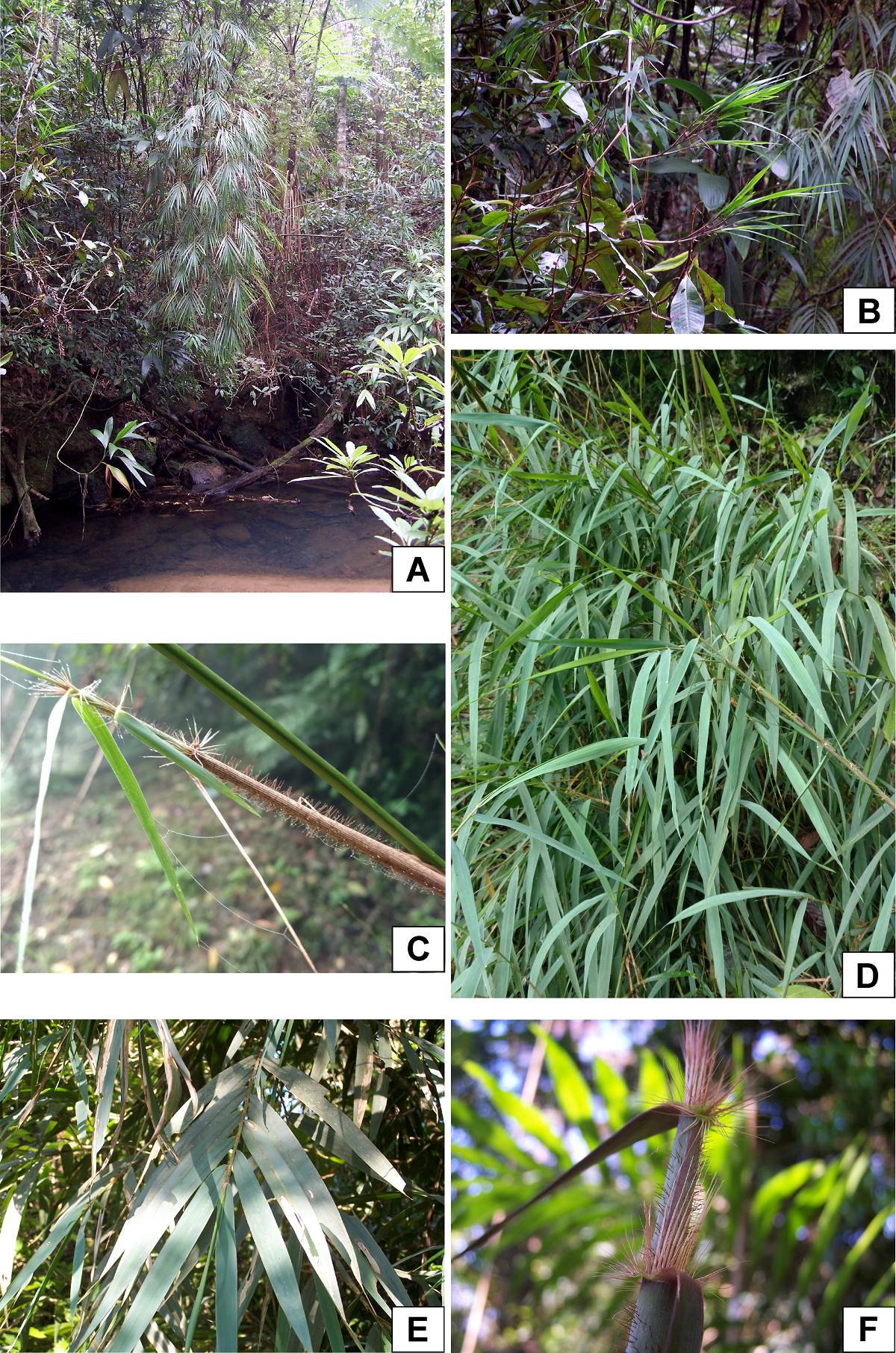

Figure 2 A-B. Olmeca clarkiae. A, Arching culm showing branches and foliage leaves. B. Foliage leaves. C-D. Olmeca fulgor. Main culm branching, showing culm leaves and blades. D. Foliage leaves. E-F. Olmeca zapotecorum. E. Foliage leaves. F. New shoot showing culm leaves and blades.
Previous molecular phylogenetic hypotheses based on plastid markers have shown that the two Olmeca species (O. recta and O. reflexa) form a clade, albeit without strong nodal support. This clade was closely related to Aulonemia clarkiae Davidse & R.W. Pohl, A. fulgor Soderstr., and an undescribed species of Aulonemia (Ruiz-Sanchez et al. 2011). Based on those phylogenetic results and morphological characters, Ruiz-Sanchez et al. (2011) proposed the inclusion of the three Aulonemia species into the genus Olmeca. However, it is important to note that Ruiz-Sanchez et al. (2011) did not achieve good phylogenetic resolution among the branches of the three former Aulonemia species. The relationships between O. clarkiae, O. fulgor, and O. zapotecorum remain unresolved, likely due to the limited resolution provided by the five plastid markers used (matK, rbcL, psbK-psbI, rpl16, and psbA-trnH). The sister relationship between Olmeca and Otatea (McClure & E.W. Sm.) C.E. Calderón & Soderstr. has been consistently supported by studies using a few chloroplast markers or complete plastomes (Ruiz-Sanchez et al. 2011, Ruiz-Sanchez & Sosa 2015, Saarela et al. 2018, Gallaher et al. 2022).
Ruiz-Sanchez & Sosa (2015) expanded the sampling of taxa and molecular markers (including atpB-rbcL, matK, rps16, rps16-trnQ, trnC-rpoB, trnD-trnT, trnL-trnF, trnT-trnL, GBSSI, and ITS) to investigate the origin and evolution of fleshy fruits in woody bamboos. Their analysis included all five known Olmeca species and confirmed that Olmeca is monophyletic. They found that O. reflexa is sister to the rest of the genus, followed by O. recta, and then a clade comprising O. fulgor as sister to the clade formed by O. clarkiae and O. zapotecorum. This finding suggests that fleshy fruits (bacoid caryopses) evolved once within Olmeca, as both O. recta and O. reflexa produce bacoid caryopses and then reverted to a typical caryopsis in the ancestor of the clade with O. fulgor, O clarkiae, and O. zapotecorum. Additionally, O. fulgor is not closely related to O. zapotecorum, despite their morphological similarities. Ruiz-Sanchez & Sosa (2015) estimated that the divergence between Olmeca and Otatea occurred approximately 7.2 million years ago (Ma). Similarly, Gallagher et al. (2022) estimated this divergence to be around 8 Ma.
Approximately 67 species across 12 genera within two tribes (Arundinarieae and Bambuseae) produce bacoid caryopses (Ruiz-Sanchez et al. 2017). Bacoid caryopses have evolved independently at least seven times within the Bambusoideae subfamily since the late Miocene (Ruiz-Sanchez & Sosa 2015). Ruiz-Sanchez & Sosa (2015) found that current climatic variables and soil parameters were not significantly correlated with the appearance of this type of fruit. However, Liu et al. (2024) demonstrated that topography and climate are correlated with the presence of bacoid caryopses in Paleotropical woody bamboos. Van der Pijl (1982) hypothesized that bacoid caryopses in Bambusoideae are related to the retention of water and nutrients, favoring rapid germination and growth, thereby producing vigorous seedlings. De la Cruz Aparicio (1985) and Ruiz-Sanchez et al. (2017) reported seedlings of Olmeca recta and O. reflexa, respectively, growing up to 1 meter tall within two months. These findings align with van der Pijl's (1982) hypothesis.
In recent years, genome subsampling techniques, such as restriction site-associated DNA sequencing (RADseq; Baird et al. 2008), have proven effective for resolving phylogenetic issues in woody bamboos. These methods have successfully clarified phylogenetic relationships at various taxonomic levels, from genus to more complex groups. Specifically, these techniques have been applied to resolve phylogenetic relationships at the genus and subtribe levels in temperate woody bamboos (Wang et al. 2017, Ye et al. 2019), paleotropical woody bamboo species (PWB) (Liu et al. 2020, 2024), and the Neotropical genus Guadua (Ruiz-Sanchez et al. 2025). Notably, Liu et al. (2024) reconstructed the relationships at the subtribe and generic levels among PWB species. They confirmed the monophyly of the four major subtribes of PWB, contrasting with the eight subtribes proposed by Wong et al. (2016).
Phylogenetic hypotheses based on a few plastid markers (Ruiz-Sanchez et al. 2011, Ruiz-Sanchez & Sosa 2015) have shown poor phylogenetic resolution (Ruiz-Sanchez et al. 2011), weak nodal support among Olmeca species (Ruiz-Sanchez & Sosa 2015), or non-monophyly (Tyrrell et al. 2018). This raises the compelling questions: could RADseq data better resolve the internal relationships within Olmeca? Will O. recta and O. reflexa form a clade, given that both species share the ability to produce bacoid caryopsis? Will the remaining three Olmeca species form a second clade, with O. fulgor as the sister species to O. zapotecorum due to their morphological similarities?
Material and methods
Sample collection. In this study, we sampled all five Olmeca species (Table 1). Leaf tissue from living specimens was collected from wild populations. Voucher specimens were deposited at the University of Guadalajara Herbarium (IBUG) and the Instituto de Ecología, A.C. Herbarium (XAL) (Thiers 2024).
Table 1 Sample species, geographical distribution and voucher records.
| Species | Distribution | Collection |
|---|---|---|
| Olmeca. clarkiae (Davidse & R.W. Pohl) Ruiz-Sanchez, Sosa & Mejía-Saulés | D | A. Vázquez 504 (XAL) |
| O. fulgor (Soderstr.)Ruiz-Sanchez, Mejía-Salues & Sosa | A,C | E. Ruiz 519 (IBUG) |
| O. recta Soderstr. | B | E. Ruiz 132 (XAL) |
| O. reflexa Soderstr. | B | T. Mejía-Saulés 2045 (XAL) |
| O. zapotecorum Ruiz-Sanchez, Sosa & Mejía-Saulés | C | E. Ruiz 864 (IBUG) |
| Otatea acuminata (Munro) C.E. Claderón & Soderstr. | A | E. Ruiz 644 (IBUG) |
A = Transmexican Volcanic Belt province, B = Veracruzan province, C = Sierra Madre del Sur Province, D = Chiapas Highland province.
DNA extraction and sequencing. Total DNA was extracted from leaf tissues using the CTAB procedure (Doyle & Doyle 1987, Cota-Sánchez et al. 2006). DNA quality was assessed by visualization on 1 % agarose gels, and quantification was performed using a Nanodrop (Thermo Scientific, USA) and a Qubit 2.0 fluorometer (Life Technologies, Carlsbad, CA, USA).
Library preparation was based on a modified double-digest restriction site-associated DNA approach (3RAD; Bayona-Vásquez et al. 2019). Genomic DNA was digested using the EcoRI-HF and NheI restriction enzymes. Sequencing was conducted on an Illumina HiSeq 2500 platform (Illumina, San Diego, CA, USA), producing paired-end reads of 150 bp. Library preparation and sequencing were carried out in the Department of Environmental Health Science at the University of Georgia, USA.
Samples were assembled into orthologous loci using ipyrad v.0.9.85 (Eaton & Overcast 2020) with a reference-guided approach. Specifically, the reference genome of Dendrocalamus latiflorus (GCA_017311315.1, Zheng et al. 2022) was used for this purpose. On average, we recovered 1.54 million (±570,922 SD) raw reads per sample. The proportion of mapped reads varied across samples, with a mean of 39.7 % (SD = 4.78 %). Notably, coverage was relatively uniform among clusters of mapped reads, with a mean of 79,434 clusters (SD = 15,156). Ultimately, the assembly yielded a final number of 54,614 non-overlapping loci, averaging 9,102 loci per sample (SD = 3,533). Further detailed information is available in the Supplementary material. Sequences were uploaded to GenBank PRJNA1267969.
Phylogenetic Analysis. We employed maximum likelihood (ML) to infer phylogenetic trees using five ingroup accessions. To root the trees, we included a specimen of Otatea acuminata (Munro) C.E. Calderón & Soderstr. Initially, we filtered all assembled loci to retain those with a minimum coverage of at least 25 % of all samples, subsequently removing any samples that contained data for less than 10 % of all sites. This resulted in a filtered dataset comprising 411 loci, which we concatenated using the window extracter module in the ipyrad-analysis toolkit to generate a final alignment. For the main reconstruction, we utilized RAxML v. 8.2.12 (Stamatakis 2014), employing the GTRCAT evolutionary model and executing 100 repetitions for bootstrap support. Additionally, we conducted a cross-validation reconstruction using I-QTREE v. 2.2.0.3 (Minh et al. 2020) with the TVM+F+G4 model, as suggested by ModelFinder (Kalyaanamoorthy et al. 2017). This involved running 1,000 replicates to obtain ultrafast bootstrap (BS) support values (Hoang et al. 2018) and 1,000 replicates to calculate Shimodaira-Hasegawa-like approximate likelihood ratio test (SH-aLRT) support (SH-aLRT, Anisimova et al. 2011).
With the aim of exploring possible incomplete lineage sorting or reticulate relationships among Olmeca species, we inferred a species tree based on a multispecies coalescent approach using ASTRAL v. 5.7.1 (Zhang et al. 2018). To acquire the required data for this analysis, we first determined how many informative gene trees could be obtained by splitting scaffolds of the reference genome into non-overlapping 2 Mb windows and concatenating assembled RAD loci within each window. This was performed using the treeslider module in the ipyrad-analysis toolkit, which also applied a filter to each alignment to remove sites with missing data across the ingroup samples. This yielded a total of 1,336 windows. We further refined this set by requiring more than 10 SNPs per window. RAxML was then used to infer a maximum likelihood tree for each of the remaining 102 alignments, which were subsequently used in our ASTRAL reconstruction.
Time tree and historical biogeography. Conventional Bayesian methods for estimating divergence times face statistical and computational challenges when applied to RADseq datasets. These challenges primarily arise from the abundance of short, unidentified RAD loci, which complicate their integration into phylogenetic partitioning strategies. To address this, we adopted the strategy of Donoghue et al. (2022) to calibrate our tree using RelTime (Tao et al. 2020) in MEGA 11 (Tamura et al. 2021). RelTime redistributes molecular branch lengths into branch-specific segments of both geological time and evolutionary rates while adhering to user-specified constraints. We used Otatea acuminata as an outgroup and calibrated the divergence between Olmeca and Otatea using the Timetree of Life tool (Kumar et al. 2022), which provides a median estimate of 7.4 Ma (95 % confidence interval [CI]: 2.9-15.7 Ma) using a normal prior to incorporate the confidence interval.
For the historical biogeography analysis, we reconstructed the ancestral species ranges for Olmeca species using the Statistical Dispersal-Vicariance Analysis (S-DIVA) approach (Yu et al. 2015). S-DIVA statistically evaluates alternative ancestral ranges at each node in a tree, accounting for phylogenetic uncertainty and uncertainty in DIVA optimization. S-DIVA was executed using RASP v. 4 (Yu et al. 2020). This method requires a phylogeny and discrete range data as input. For the phylogeny, we used the RAxML tree. For discrete range data, we coded the Olmeca species according to their geographical distribution (Ruiz-Sanchez et al. 2020), by dividing it into four areas based on the biogeographical regionalization of the Neotropical Region proposed by Morrone (2014): (A) Transmexican Volcanic Belt province; (B) Veracruzan province; (C) Sierra Madre del Sur province; and (D) Chiapas Highland province (Table 1).
Results
Phylogenetic Analysis. The final alignment consisted of 4,292,771 base pairs (bp) and included 9,592 single nucleotide polymorphisms (SNPs) with 66.2 % missing data. Maximum likelihood analyses (RAxML and IQ-TREE) and coalescent-based analyses produced identical topologies (Figure 3A). Olmeca was confirmed to be monophyletic with strong support across all metrics: 100 % bootstrap support (BS), SH-aLRT of 100, and posterior probabilities (PP) of 1.0. Two well-supported clades were identified (100 % BS, 100 SH-aLRT, 1.0 PP). The first, informally named the Reflexa clade, includes O. recta and O. reflexa (Figure 3A). The second, informally named the Fulgor clade, consists of O. fulgor and O. zapotecorum as sister species (100 % BS, 100 SH-aLRT, 1.0 PP), with O. clarkiae sister to this pair (Figure 3A).
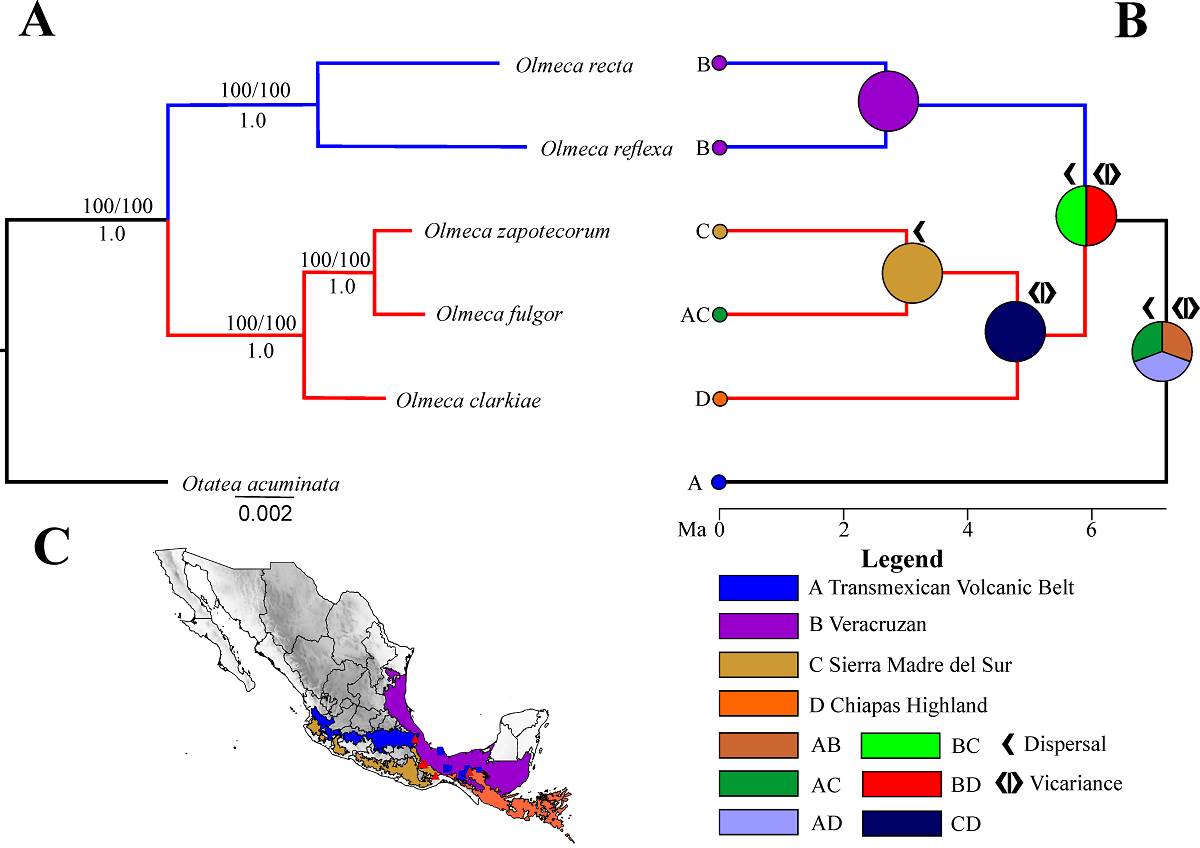
Figure 3 A. Maximum likelihood tree inferred using RAxML, numbers above the branches are bootstrap support values (left) and SH-aRLT support values (right); numbers below the branches are posterior probabilities. Blue branches indicate the informal Reflexa clade and red branches the informal Fulgor clade. B. RelTime (MEGA 11) calibrated chronograph derived RAxML tree based on SNPs and ancestral range inferences. Circles at nodes represent ancestral area optimizations. C. Map showing biogeographical regions according to Morrone (2014) and legend of the biogeographical areas used.
Time tree and historical biogeography. The time-tree estimation analysis indicates that the divergence between Olmeca and Otatea occurred at a mean age of 7.2 million years ago (Ma) (95 % confidence interval [CI] = 6-13.4 Ma). The crown age of Olmeca was estimated at 5.89 Ma (95 % CI = 4.69-7.4 Ma). The Reflexa clade was dated to 2.68 Ma (95 % CI = 1.48-4.84 Ma), while the Fulgor clade was estimated to be 4.8 Ma (95 % CI = 3.98-5.77 Ma). The clade formed by O. fulgor and O. zapotecorum had an estimated age of 3.01 Ma (95 % CI = 2.03-4.48 Ma) (Figure 3B).
The biogeographic analysis using the S-DIVA model identified two possible combined ancestral areas for Olmeca: Veracruzan + Chiapas Highland provinces (50 %) or Veracruzan + Sierra Madre del Sur provinces (50 %). For the Reflexa clade, the ancestral area was identified as the Veracruzan province. In the Fulgor clade, a combined ancestral area of Chiapas Highland + Sierra Madre del Sur provinces was determined (100 %). Finally, the clade formed by O. fulgor and O. zapotecorum was found to have the Sierra Madre del Sur province as its ancestral area (100 %). Additionally, the analysis supports a series of events throughout the evolutionary history of Olmeca, including three dispersal and three vicariance events (Figure 3B, C).
Discussion
Our phylogenetic hypothesis supports that Olmeca is a monophyletic group comprising two well-supported clades: the Reflexa and Fulgor clades (Figure 3A). Previous phylogenetic studies using a limited number of chloroplast markers suggested that Olmeca was non-monophyletic (Ruiz-Sanchez et al. 2011, Tyrrell et al. 2018). In contrast, Ruiz-Sanchez & Sosa (2015), using a combined nuclear and chloroplast dataset, recovered Olmeca as monophyletic. However, their results showed that O. recta and O. reflexa did not form a clade. Despite containing 66.2 % missing data, our analysis successfully resolved the phylogenetic relationships withinOlmeca. This aligns with findings by Tripp et al. (2017), who reconstructedPetalidiumNees (Acanthaceae) phylogeny with 90 % missing data. The use of SNPs from RADseq provided significantly more molecular characters than previous phylogenetic studies where Ruiz-Sanchez et al. (2011) had only 97 and Tyrrell et al. (2018) had only 265 parsimony informative characters (vs. ~9,000 here), resulting in a more resolved phylogenetic tree with strong nodal support. The Reflexa clade comprises two sister species, Olmeca recta and O. reflexa, both strongly supported by high nodal values (Figure 3A). Both O. recta and O. reflexa inhabit tropical perennial forests. According to Ruiz-Sanchez et al. (2020), few Mexican woody bamboo species occur in this vegetation type, including Rhipidocladum bartettlii (McClure) McClure, Chusquea simpliciflora Munro, Guadua tuxtlensis Londoño & Ruiz-Sanchez, and G. longifolia (E. Fourn.) R.W. Pohl.
The Fulgor clade consists of O. fulgor as sister to O. zapotecorum, with O. clarkiae sister to that pair (Figure 3A). Ruiz-Sanchez et al. (2011) recovered the same clade but noted a lack of internal resolution (i.e., a polytomy). Ruiz-Sanchez & Sosa (2015) also identified this clade but with a conflicting topology, placing O. fulgor as sister to O. clarkiae under weak support. Our new phylogeny, based on nuclear SNPs, resolves these relationships with strong support and well-resolved clades.
We found that the crown age of Olmeca was 5.89 Ma, while Ruiz-Sanchez & Sosa (2015) estimated it at 6.05 Ma. Both time estimates are very similar and fall within the Late Miocene period. However, the topologies of the two phylogenetic trees differ significantly. The estimated crown age for Reflexa clade, the clade that includes two species with bacoid fruits, was 2.68 Ma (Figure 3B), which corresponds to the Late Pliocene period. In contrast, Ruiz-Sanchez & Sosa (2015) found a paraphyletic relationship between these two species and reported an older divergence time. Our time estimate for these bacoid fruit-producing species is similar to those of Dinochloa Buse and Melocalanna Trin., two Asian genera that also produce bacoid fruits (Liu et al. 2024). However, the estimated age is younger than that of Melocalamus Link & Otto (7.66 Ma), another Asian genus producing bacoid fruits (Liu et al. 2024). Additionally, Alvimia C.E. Calderón ex Soderstr. & Londoño, a Neotropical woody bamboo genus that produces bacoid fruits, shows a divergence time similar to that of Olmeca (Ruiz-Sanchez & Sosa 2015). According to Liu et al. (2024), bacoid fruits may have evolved independently in the Arundinarieae and Bambuseae tribes, following similar evolutionary patterns as adaptations to comparable environments.
The Fulgor clade has an estimated crown age of 4.89 Ma (Figure 3B), corresponding to the middle Pliocene. This clade is older than the Reflexa clade. The three species that compose this clade inhabit montane cloud forests (Ruiz-Sanchez et al. 2011, 2020). Montane cloud forests are among the richest ecosystems in Mexico, accounting for 10 % of the entire Mexican flora (Ornelas et al. 2013). This ecosystem harbors nearly 50 % of Mexico’s total woody bamboo diversity (Ruiz-Sanchez et al. 2015, 2020), and most of its woody bamboo species fall into some risk category (Ruiz-Sanchez et al. 2020). Notably, the montane cloud forest is one of the most threatened ecosystems in Mexico (Ornelas et al. 2013). Olmeca clarkiae and O. zapotecorum have been classified as Critically Endangered, while O. fulgor is listed as Endangered. In contrast, O. recta is categorized as Near Threatened, and O. reflexa as Vulnerable (Ruiz-Sanchez et al. 2020).
As mentioned earlier, the Reflexa clade includes two species that inhabit tropical perennial forests, while the Fulgor clade comprises three species that inhabit montane cloud forests. A similar pattern of two clades was found in the phylogeny of Tridimeris Baill., a perennial tree from the Annonaceae family (Ortiz-Rodriguez et al. 2024). Tridimeris is now taxonomically classified into two sections: Tridimeris sect. Tridimeris and T. sect. Zoque Ortiz-Rodr. & Couvreur. Species in sect. Tridimeris inhabit montane cloud forests, with some occurring in sympatry with species from the Fulgor clade. Conversely, species in sect. Zoque inhabit tropical perennial forests, with some coexisting with Olmeca recta and O. reflexa in Los Tuxtlas, Uxpanapa, and La Pera (Ortiz-Rodriguez et al. 2024). Unfortunately, the phylogeny of Tridimeris has not been dated, but it would be interesting to determine whether both genera share similar diversification ages.
Our ancestral area reconstruction identified two possible ancestral regions for Olmeca, each with equal probability (50 %): Veracruzan + Chiapas Highland provinces or Veracruzan + Sierra Madre del Sur provinces. The reconstruction suggests that, in the first scenario, a dispersal event occurred from Veracruzan to Sierra Madre del Sur, whereas in the second scenario, dispersal occurred from Veracruzan to Chiapas Highland. However, in both cases, a vicariance event took place, leading to the separation of the ancestral Veracruzan province from the other two areas. This implies a division between montane cloud forests and tropical lowland ecosystems.
A distinctive feature of montane cloud forests is their location at elevations higher than tropical lowland ecosystems (such as tall or medium evergreen and semi-evergreen forests) but lower than the temperate forests found in mountainous regions (e.g., Pine, Pine-Oak, or Oak forests). In addition to species exclusive to these forests, montane cloud forests also host species typical of both lowland humid forests and temperate forests from colder mountain regions, where the combination of altitude, humidity, and temperature fosters their coexistence (Villaseñor 2010).
The crown age of the Fulgor clade was estimated at 4.89 Ma, while the Reflexa clade dates to 2.68 Ma - representing a 2.21 Myr divergence. This extended temporal gap may reflect the time required for environmental stabilization, particularly the re-establishment of tropical perennial forest conditions that potentially facilitated the evolution of bacoid-type fruits in both species. These findings align with Ruiz-Sanchez & Sosa (2015) and Liu et al. (2024), who reported that bacoid fruits have independently evolved in different bamboo lineages within tropical forests. Moreover, they highlighted that this type of caryopsis represents an evolutionary novelty not only in bamboo species but also among grasses in general.
The crown age ofOlmecawas estimated at 5.89 Ma, corresponding to the Late Miocene when the divergence between the Fulgor and Reflexa clades occurred. According to Graham (1975), tropical perennial forests were replaced by warm-temperate oak forests during this period. This finding was further supported by Palacios-Chávez & Rzedowski (1993), who documented the replacement of tropical perennial forests with montane cloud forests in northwestern Chiapas. These paleoenvironmental changes suggest that periods of environmental instability followed by stabilization may have driven the divergence of theseOlmecaclades in southern Mexico. It is important to note thatOlmeca clarkiaeis geographically isolated from other Fulgor clade species by the Isthmus of Tehuantepec. This lowland valley, which separates the Sierra Madre del Sur from the Chiapas Highlands, formed approximately 6 Ma (Barrier et al. 1998). The Isthmus serves as a classic example of vicariance through habitat fragmentation (Ornelas et al. 2010, 2013, Gutiérrez-Ortega et al. 2020, 2024).
Our results suggest that RADseq data can effectively resolve the internal relationships within Olmeca, providing a well-supported phylogenetic framework for the genus. The analyses recovered O. recta and O. reflexa as a monophyletic group, supporting the hypothesis that the ability to produce a bacoid caryopsis is a shared, derived trait. Additionally, the remaining three Olmeca species formed a second clade, with O. fulgor as the sister species to O. zapotecorum, consistent with their morphological similarities. These findings highlight the power of RADseq data in resolving complex phylogenetic relationships and contribute to a better understanding of the evolutionary history of Olmeca.
Supplementary material
Supplemental data for this article can be accessed here: https://doi.org/10.17129/botsci.3676











 nueva página del texto (beta)
nueva página del texto (beta)



