Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Similars in
SciELO
Similars in
SciELO
Share
Tropical and subtropical agroecosystems
On-line version ISSN 1870-0462
Trop. subtrop. agroecosyt vol.13 n.1 Mérida Jan. 2011
Artículos de investigación
Molecular characterization and identification of biocontrol isolates of Trichoderma harzianum from Embu District, Kenia
Caracterización molecular e identificación de aislamientos de biocontrol de Trichoderma harzianum del Distrito de Embu, Kenia
E. N. Siameto1, S. Okoth2*, N. O. Amugune2, and N. C. Chege2
1 School of Sciences, Narok University College, P. O Box 861, Narok * Corresponding author E-mail: dorisokoth@yahoo.com Tel: 02-444904 Ext 2483.
2 School of Biological Sciences, University of Nairobi, P.O Box 30197, Nairobi.
Submitted February 15, 2010
Accepted May 25, 2010
Revised received June 11, 2010
Abstract
Species in the genus Trichoderma are important commercial source of several enzymes, biofungicides and growth promoters. The most common biological control agents of the genus are strains of T. harzianum, T .viride and T. viriens. In this study, sixteen selected isolates of T. harzianum from different land use types in Embu, Kenya were tested for antagonistic action against five soil borne phytopathogenic fungi (Rhizoctonia solani, Pythium sp, Fusarium graminearum, F. oxysporum f sp phaseoli and F. oxysporum f sp Lycopersici) using dual culture assay and through production of non-volatile inhibitors. Seven isolates were further characterized using RAPD-PCR procedure to determine genetic variability. All T. harzianum isolates had considerable antagonistic effect on mycelial growth of the pathogens in dual cultures compared to the control. Maximum inhibitions occurred in Pythium sp-055E interactions (73%).The culture filtrates obtained from Czapek's liquid medium reduced the dry weight (mg) of the mycelia significantly while those from the potato dextrose broth showed minimum inhibition growth. Pythium sp. was most sensitive compared to other pathogens. Genetic similarities generated using Jaccard's coefficient of similarity ranged from 0.231 to 0.857 for isolates 055E, 011E, 010E and 015E. Since all T. harzianum isolates evaluated were effective in controlling colony growth of the soil borne pathogens both in dual cultures and in culture filtrates they could be tried as a broad spectrum biological control agent in the green house and under field conditions.
Key words: Trichoderma harzianum, growth antagonism, genetic similarity RAPDs.
INTRODUCTION
Trichoderma harzianum is an asexually reproducing filamentous fungus and a species aggregate. It is grouped on the basis of conidiophores branching patterns with short side branches, short inflated phialides, smooth and small conidia (Rifai 1969). These characteristics allow for the relatively easy identification of Trichoderma as a genus, but the species concept is difficult to interpret and there is considerable confusion over the application of specific names (Rifai 1969; Papavizas 1985). This disparity of criteria makes it difficult to search for, and above all characterize new biocontrol agents (BCAs) within the species and to reidentify them in natural environment once they are present in a pathosystem (Papavizas 1985).
The advent of polymerase chain reaction (PCR) has allowed the analysis of small numbers of fungal cells or even single spores, dried herbarium material (Carlies & Watkinson 1994), or extinct organisms (Golenberg et al. 1990).The selection of universal oligonucleotide primers specific to fungi (Van Belkum et al. 1993; Sandhu et al. 1995) has provided easy access to nucleotide sequences. These techniques have been proven to be valuable tools in fungal taxonomy and their application has led to the reconsideration of several genera (Sherriff et al., 1994)
Hence, Random markers as products of the PCR-RAPD technique have been used for taxonomy of Trichoderma to discriminate species (Hadrys et al., 1992; Williams et al., 1990).
Trichoderma species have been investigated for over 70 years (Hjeljord & Tronsmo 1998). They have been used as biological control agents (BCAs) and their isolates have become commercially available of late (Freeman et al., 2004). This development is largely the result of a change in public attitude towards the use of chemical pesticides and fungicides such as methyl bromide (Elad et al. 1980; Basim et al. 1999). In this respect Trichoderma species have been studied as BCAs against soil-borne plant pathogenic fungi (Henis 1984; Chet and Inbar, 1994). Replacement or reduction of chemical application can be achieved through use of biologically based fungicides, a concept included in the broad definition of biological control proposed by Cook & Baker (1983). The commercial use of Trichoderma BCAs must be preceded by precise identification, adequate formulation, and studies about the synergistic effects of their mechanisms of biocontrol.
In this study the in vitro antifungal activity of T. harzianum isolates were evaluated against selected soil borne plant pathogens (Rhizoctonia solani, Pythium sp, Fusarium graminearum, F .oxysporum f. sp phaseoli and F. oxysporum f. sp Lycopersici). We aimed also at characterizing T. harzianum isolates using RAPD-PCR technique to establish the degree of genetic variation and determine any relationship between molecular variation and antifungal activity.
MATERIAL AND METHODS
Isolation and identification of Trichoderma species
Trichoderma sp were isolated using dilution plate technique (Johnson et al, 1959) and soil washing methods (Gams et al, 1987; Bills & Polishook 1994) on malt extract agar (MEA) and cornmeal agar (CMA) with 2% dextrose) both with streptomycin 50mg/L and cyclcosporin 10mg/L. The colonies were counted and identified and transferred to Petri dishes containing potato dextrose agar (PDA) and incubated at 15, 25, 30 and 35°C for further identification to species level.
Genus identification of green fungus was undertaken using the method of Domsch et al. (1980). Trichoderma isolates were identified to species level following the taxonomic key of Samuels et al. (2004). Microscopic examination was carried out by mounting the culture in lactophenol cotton blue but for size measurements KOH and water was used as the mounting fluid. A small amount of material was placed in a drop of 3% KOH on a slide and then replaced with water. The isolates were preserved in sterile soil and stored at 4°C.
Isolation of phytopathogenic fungi.
Diseased plant tissues were obtained from the University of Nairobi, College of Agriculture and Veterinary Medicine (CAVM), Upper Kabete campus field station which is 10 Km north-west of Nairobi. Rhizoctonia solani was isolated from Spinach (Spinacea olerácea), Fusarium oxysporum f. sp Lycopersici from tomato (Lycopersicum esculetum), F. oxysporum f. sp phaseoli from beans (Phaseolus vulgaris) and F. graminearum from maize cob (Zea mays). Diseased plant tissues were washed under running tap water to remove surface soil, dust and other contaminants. Tissue pieces were cut out from the leading edge of lesion, and placed in one percent sodium hypochlorite for five minutes, then washed in sterile distilled water and dried on sterile filter paper. The dried pieces were cut into approximately one centimeter pieces, plated onto PDA and incubated at 25°C. Pythium sp was obtained by planting beet root seeds in water logged soil. The seedlings that emerged were infected by dumping- off. The seedlings stems were cut into one centimeter pieces and inoculated onto PDA plates.
Evaluation of Dual culture on agar plates
Plates of PDA were inoculated with five millimeter disc obtained from five-day-old cultures of the phytopathogens ten millimeter from the edge of the plate. After two days a five millimeter disc of the Trichoderma harzianum cultures was placed in the same plate at a distance of fifty five millimeter from the phytopathogens disc. Pythium sp and the biocontrol strains were inoculated at the same time. The paired cultures were incubated at room temperature for six days. The growth of the fungi was recorded by measuring the radial growth of the pathogens. The percentage growth of the pathogens was calculated as follows:
Percent growth=Radius of the growth in the direction of the test strain ÷ radius of the growth in the absence of the test strain x100 (Edington et al, 1971).--------- Equation 1
Evaluation of dual culture using Slide method
For Rhizoctonia solani and Pythium sp -Trichoderma interaction, a clean sterile glass slide was placed in nine centimeter diameter plates. A small amount of melted PDA was spread over the slide to make a thin film on the slide. Five millimeter discs of one week old growing colonies cut from the margin of each pathogen and Trichoderma isolates were placed on the opposite sides of the slide three centimeter apart on the PDA surface. A two milliliter of sterilized distilled water was added to the plate to prevent drying and incubated 25 ± 1°C for a week. At the end of incubation period, point of contact between Trichoderma—Pathogen hyphae was stained with lactophenol in cotton blue and observed under a light microscope for the presence of mycelial penetration and for cell wall disintegration.
Determination of antifungal properties of T. harzianum culture filtrates
To harvest non-volatile antibiotics produced by Trichoderma isolates, the fungus was grown on potato dextrose broth (PDB) and Czapek's liquid medium (CLM) Each T. harzianum isolate was inoculated into 100ml PDB incubated at 20OC then filtered through 0.22 mm Millipore filters after 10 days. 2ml aliquots of these filtrates were placed in sterile Petri dishes and 25ml of PDA added. Five millimeter wide mycelial discs of the pathogen were placed at the center of the solidified agar plates and incubated at room temperature for 6 days. Colony diameters were measured daily and inhibition percentage obtained using the formula:
Inhibition percent = [(C2-C1) ÷ C2) x 100] ((Edington etal. 1971).----------------Equation 2
Where: C1 means growth of the pathogens in the presence of antagonist and C2 means the growth of control. Each experiment had three replicates and complete randomized design was adopted .PDA without the culture filtrates served as the control.
To test for non-volatile antibiotics from Czapek's liquid medium the method of Tianhui (1994) was amended. Flasks containing 50 ml of CLM were inoculated with a five millimeter disc of T. harzianum isolates and incubated at 25°C in the dark for two weeks. Culture liquids were filtered through filter paper and sterilized by bacterial filtration (Acrodisc nitrocellophane membranes, pore size 0.22|am). The filtrates were diluted with PDB to a concentration of 70%. PDB lacking the filtrate of Trichoderma served as a control. Two agar discs containing the pathogen were inoculated in each flask and then incubated at 25°C and mycelia filtered through filter paper after eight days, dried and weighed. The percent dry weight was then obtained using the following formula:
Percent dry weight = dry weight of control - dry weight of pathogen ÷ dry weight of control x 100.-------------------Equation 3.
Each treatment was replicated three times.
Isolation of DNA from filamentous fungi
Mycelia for DNA extraction were cultured in 50 ml of yeast potato dextrose broth (YPDB) at 20°C with rotary shaking at 120 revolutions per minute (rpm), harvested by filtration through a filter paper and washed with distilled water after two days. The mycelia were freeze-dried and ground in the presence of sand and stored at -20°C. One hundred milligrams of the mycelial powder was transferred into eppendorf tubes. 500μ1 of 2xCTAB buffer (equilibrated to 65°C) and 1.0% of p-mercaptoethanol were added and the tubes heated at 65°C for 30 minutes. 500μ1 of chloroform-isoamyl alcohol (24:1) was added to the tubes and then vortexed for 30 seconds then centrifuged at 12,000rpm for 15 minutes. The upper portion of the aqueous phase was recovered and 1/5 volume of 5% CTAB and mix. Another chloroform-isoamyl alcohol (24:1) extraction was performed. Equal volume of CTAB precipitation buffer was mixed with the recovered supernatant and left to stand on ice for 20 minutes. The mixture was centrifuged for 15 minutes at 12,000rpm and the supernatant discarded. The pellet was rehydrated with high salt TE buffer (heat at 65°C for 5-10 minutes). DNA was precipitated by the addition of an equal volume of cold absolute ethanol with incubation at -20°C for 10 min. DNA was collected by centrifugation at 12,000rpm for 10 minutes, washed with 70% ethanol, dried, and re-suspended in lOOul of 1XTE buffer and stored at -20°C. The quality of the DNA was checked by use of 0.7% agarose gel and quantified spectrophotometrically.
RAPD analysis and PCR conditions
Amplification reactions were performed in 0.6ml microcentrifuge tubes in a 25 μ1 reaction volumes containing 5ng of template DNA, Taq buffer, 2.5mM MgC12, 0.8mM each dNTP, 1.15ng/μ1 Primer and lunit Taq DNA polymerase. Amplification reactions were performed in a Perkin-Elmer, Gene amp PCR system 2400 thermal Cycler programmed for 35 cycles of denaturation at 94°C for 30 seconds, low stringency annealing temperature at 31°C for 1.0 minute and polymerization at 72°C for 1.0 minute with a final extension step at 72°C for 10 minutes. PCR products were separated on 2.5 % agarose/ IX TBE gels. A 50bp DNA molecular size marker was loaded on the first well and used for comparison. The banding pattern was visualized on Ultraviolet ttansilluminator and documented by MultiDoc digital imaging system.
Data analysis
Bands were manually scored 1 for presence and 0 for absence and the binary data used for statistical analysis using the software R-command version 2.1.1. The size of the fragments (molecular weight in base pairs) was estimated by using 50 bp ladder marker, which was run along with the amplified products. A genetic dissimilarity matrix was calculated according to Squared Euclidean Distance which estimated all pair-wise differences in the amplification product and Cluster analysis was done by Wards method using a minimum variance algorithm (Ward, 1963).
Genetic Similarity (GS) was analyzed using the equation (Jaccard, 1908):
GS = (Nab)/(Na+Nb-Nab) ---------- Equation 4.
Where Nab is the number of shared fragments between isolates a and b, Na is the number of scored fragments of isolate a, and Nb the number of scored fragments of isolate b. Genetic distance (GD) was then calculated as GD = 1-GS. The data for growth inhibition measurement were arcsine transformed then subjected to analysis using the Statistica version 6 software.
RESULTS
Inhibition of growth of pathogenic fungi by T. harzianum on culture media
As single cultures Phythium sp grew actively and colonized the entire agar surface within two days, whereas F. oxysporum took two weeks, and F. graminearum and Rhizoctonia sp filled the plate in six days. Dual culture assays provided evidence that T. harzianum isolates reduced growth of the pathogens. Nine out of the sixteen isolates tested were able to inhibit the growth of three pathogens each by more than 50%.Three of the T. harzianum isolates inhibited four pathogens. Isolates 015E and 05 IE were superior to others since they inhibited the growth of five pathogens tested.
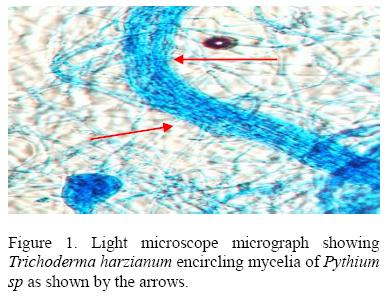
T. harzianum showed parasitic behavior against Pythium (Fig 1) by coiling round the host hyphae and degrading it. The dual cultures plates showed initial rapid growth of the host which stopped at the point of contact with the parasite. T. harzianum over grew the pathogen resulting into complete degradation of the latter and sporulation of the former over the entire plate. The isolate 055E gave the lowest percentage inhibition of 31.76% against T. harzianum while 015E gave the highest inhibition of 73.33 %.
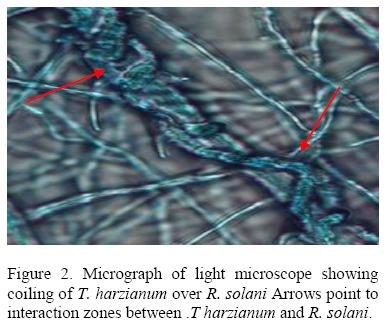
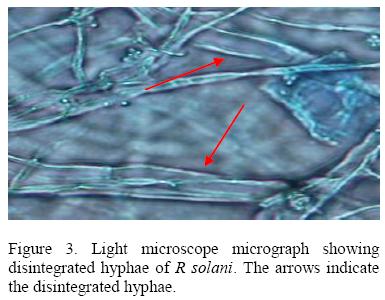
All isolates of T. harzianum that were tested inhibited growth of R .solani with isolate 029E giving the highest percentage inhibition (61.55 %) while 063E the lowest (25.88%). The inhibition process was parasitic (Fig. 2), where the parasite penetrated the host cell wall directly suggested that there was some mechanical activity that led to the disintegration of the hyphae (Fig 3).
Inhibition of growth of pathogenic fungi by T. harzianum through production of non-volatile compounds
Slight colony inhibition of the pathogens (F. oxysporum f.sp Lycopersici, F. oxysporum f. spp. phaseoli, Pythium spp., Fusarium graminearum and Rhizoctonia solani) was observed when exposed to the culture filtrates of T. harzianum (Table 1). There was no inhibition of radial growth by all tested isolates on Pythium sp. Isolate 014E inhibited F. oxysporum growth by 80%. Isolates 057E, 021E, 055E, 015E, 0051E, and 010E had no effect on F. graminearum. The culture filtrates from isolates 042E, 063E, 030E, 029E, 046E, 044E, 011E, 049E, and 014E inhibited the growth of R. solani.
The different culture filtrate from the T. harzianum isolates caused significant (P =0.05) effect on the dry weight of the phytopathogens: F. oxysporum f. sp Lycopersici, F. oxysporum f. sp phaseoli Pythium sp, Fusarium graminearum and Rhizoctonia solani. The culture filtrates of the isolates suppressed the growth of the pathogens and reduced the dry weight (mg) mycelia (Fig 4). Pythium sp was the most influenced as compared to the control. However some isolates' filtrates seems to have enhanced the accumulation of mycelia since the percentage weight obtained of the mycelia was more than 100%. Pythium sp for example was enhanced by the isolates 042E, 049E, 046E, 045E, 02 IE and 05 IE while the rest (014E, 029E, 010E, 044E, 055E, 011E, 015E, 057E, and 031E) greatly suppressed the accumulation of the mycelia of Pythium by more than 50%. The filtrate from the isolate 011E was superior to others since the dry weight of the mycelia of four (F. oxysporum f. sp Lycopersici, F. oxysporum f. sp phaseoli Pythium sp and F. graminearum) phytopathogens was less than 50%. The Fusarium pathogens were always susceptible to the culture filtrates from all the T. harzianum isolates.
DNA polymorphism analysis of the T. harzianum isolates
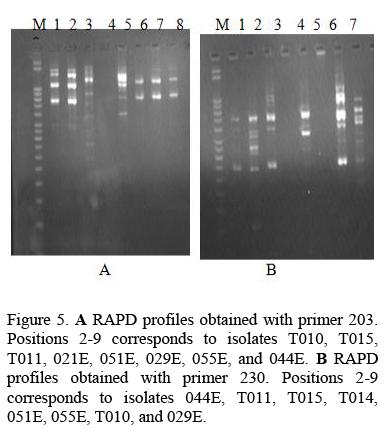
In the study four random primers namely 203, 230, 220, Op 13 were used which gave bands ranging from 350bp to 2000bp as shown in Table 2. All the primers produced intense bands totaling to81 bands were produced. The 7 samples used for DNA polymorphism were antagonistic to the phytopathogens used. In dual cultures the isolates 015E and 05 IE were superior to others since they inhibited the growth of five pathogens by more than 50% and 029E inhibited the growth of four pathogens. In the culture filtrate activity the isolates 011E and 055E had a wide range of activity by suppressing the growth of five pathogens while isolates 044E and 010E suppressed the growth of four pathogens each.
The DNA profiles obtained for Trichoderma harzianum isolates were scored (Fig 5) and a Dendrogram or Dissimilarity matrix developed using Squared Euclidean Distance and Clustering based on Wards method (Fig 6).
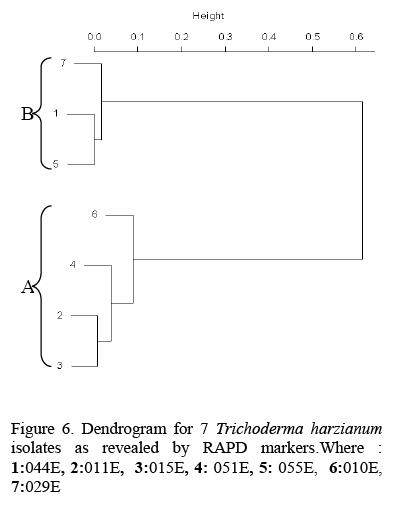
In the Dendrogram, all the 7 isolates were distinctly divided into two major clusters 'A' and 'B' at 20 units. Isolate 05 IE and 029E spanned the extremes of the entire Dendrogram. Genetic dissimilarity ranged from a lowest of 0.143 (between T010 and T015) to a highest of 0.857 (between 055E and 051E).Isolate 051E, T011, T015, and T010 were assigned to cluster 'A'. Genetic dissimilarity among the entries in this cluster ranged from a lowest of 14.3 percent (between T015and T010) to a highest of 35.7 percent (between TOlOand 05 IE). The other cluster 'B' comprised of three accessions. In this cluster isolate 044E 055E and 029E were grouped together. The genetic dissimilarity in this group was ranging from 33.3 percent between 055E and 029E to a high of 75 percent between 044E and 029E.
DISCUSSION
T. harzianum inhibited the growth of the target organisms through its ability to grow much faster than the pathogenic fungi thus competing efficiently for space and nutrients. Starvation was the most common cause of death for microorganisms, so that competition for limiting nutrients resulted in biological control of fungal phytopathogens.
A second mechanism of pathogen control was mycoparasitism. Microscopic observation of the interaction region between R. solani and Pythium spp. with T. harzianum showed that the mycelia of T. harzianum grew on the surface of the pathogens always coiling round their mycelia and later penetrating their cell walls directly without formation of appresorium structures. The pathogen mycelia then disintegrated suggesting enzyme action. Lorito et al. (1998), Metcalf et al. (2001) and Sharon et al., 2001 demonstrated possible role of chitinolytic and/or glucanases enzymes in bio-control by Trichoderma. These enzymes function by breaking down the polysaccharides, chitin, and glucans that are responsible for the rigidity of fungal cell walls, thereby destroying cell wall integrity limiting the growth of the pathogen. A mixture of several enzymes might be necessary for efficient cell wall lysis. T. harziunum has been reported to apply high chitinase and β-1, 3-glucanase activities (Sivan et al., 1984).
A third mechanism of pathogen control by Trichoderma was through antibiosis. Culture filtrates of the isolates had an inhibitory effect on the radial growth of the pathogens and mycelial accumulation suggesting action of non-volatile antibiotics in the filtrates. This agrees with the findings of Sivan et al., (1984) who noted that culture filtrates of T. harzianum strongly inhibited the growth of Pythium aphanidermatum whereas T. hamatum filtrates caused only minor inhibition of growth. Antibiotic inhibitions have been documented by Claydon et al. (1987), Dubey and Suresh (2006), Kucuk and Kivanc (2003) and Lynch (1990). Claydon et al. (1987) reported inhibition due to antibiotics trichodermin, harzianum A and harzianolide. Dubey and Suresh (2006) found that non-volatile substances produced by T. harzianum are inhibitory to F. oxysporum f. sp. ciceris causing chickpea wilt.
Culture filtrate from PDB inhibited Fusarium spp. and R. solani but not Pythium sp. The difference in activity of the culture filtrates displayed by PDB and Czapek's liquid media on the pathogens indicates the importance of substrate in fungal production of secondary metabolites.
The presence of inhibition zones in dual cultures between F. oxysporum f. sp Lycopersici and T. harzianum suggested secretion of diffusible nonvolatile inhibitory substance by the T. harzianum isolates, which has also been documented by Grodona etal, (1997)and Behzade et al., (2008).
The DNA analysis of T. harzianum isolates from Embu showed existence of intraspecific variation. The genetic variation shown by the four random primers 203, 230, 220, Op 13 resulted in bands ranging from 350bp to 2000bp. This agrees with Moller et al. (1995) who detected intraspecific diversity not only between isolates of Chaunopycnis alba from different geographic regions or hosts, but also between isolates from a single location.
Based on RAPD data isolates from the same land use appeared in different groups due to intraspecific variation, for example isolate 044E and isolate 01 IE collected from camphor plantation were clustered in different groups and were 54.4% dissimilar. Goes et al., (2002) also found intraspecific genetic variation among Trichoderma isolates. In his work the isolate Tm isolated from corn seed was grouped in the Dendrogram with the isolate 2820 isolated from sugar cane.
CONCLUSION
T. harzianum is a good candidate for biological control due to the different modes of action the fungus employs in inhibiting the growth of other fungi. Through RAPD technique intraspecific genetic variation between T. harzianum isolates was observed. The results presented here support the existence of cryptic species with similar morphology.
ACKNOWLEDGEMENTS
We would like to express our special thanks to the University of Nairobi for laboratory equipment and enabling environment to conduct research and the Conservation and sustainable management of below ground biodiversity (CSM-BGBD) project. A project executed by TSBF/CIAT with cofinancing from the Global Environment Facility (GEF) and implementation support from the United Nations Environment Programme (UNEP) for financial support of the research.
REFERENCES
Basim, H., Ozturk, S.B. and Yegen, O. 1999. Efficacy of a biological fungicide (Planter Box Trichoderma harzianum Rifai T-22) against seedling root rot pathogens (Rhizoctonia solani, Fusarum sp) of cotton. GAP-Environmental Symposium. Sanliurfa. Turkey, 137-144. [ Links ]
Behzad, H., Mousa, T., Mohammad, R.M. and Mahdi, D. 2008. Biological potential of some Iranian Trichoderma isolates in the control of soilborne plant pathogenic fungi. African Journal of Biotechnology. 7: 967-972. [ Links ]
Bills, G. F. and Polishook, J. D. 1994. Abundance and diversity of microfungi in leaf litter of a lowland rain forest in Costa Rica. Mycologia. 86: 187-198. [ Links ]
Carlie, M. J. and Watkinson, S. C. 1994. The fungi. Academic press,Ltd., London, United Kingdom. [ Links ]
Chet, I. and Inbar, J. 1994. Biological control of fungal pathogens. Applied Biochemistry and Biotechnology 48: 37-43. [ Links ]
Claydon, N., Allan, M, Hanso, J. R. and Avent, A. G. 1987. Antifungal alkyl pyrones of Trichoderma harzianum. Transactions British Mycological Society. 88: 503-513. [ Links ]
Cook, R. J. and Baker, K. F. 1983. The Nature and Practise of Biological Control of Plant Pathogens. American Phytopathological Society, St Paul M N, 539. [ Links ]
Domsch, K.H., Gams, W. and Anderson, T.H. 1980. Compendium of soil fungi. Academic Press London. 809. [ Links ]
Dubey, S.C. and Suresh, M. 2006. Randomly Amplified Polymorphic DNA Markers for Trichoderma species and Antagonism against Fusarium oxysporum f. sp. ciceris Causing Chickpea Wilt. Journal Phytopathology. 154: 663-669. [ Links ]
Edington, L.L., Khew, K.L. and Barron, G.I. 1971. Fungitoxic spectrum of benzimidazole compounds. Phytopathology. 61: 42-44. [ Links ]
Elad, Y., Chet, I. and Katan, Y. 1980. Trichoderma harzianum a biocontrol agent effective against Sclerotium rolfsii and Rhizoctonia solani. Phytopathology. 70: 119-121. [ Links ]
Freeman, S., Minz, D., Kolesnik, I., Barbul, O., Zreibil, A., Maymon, M, Nitzani, Y., Kirshner, B., Rav-David, D., Bilu, A., Dag, A., Shafir, S. and Elad, Y. 2004. Trichoderma biocontrol of Colletotricum acutatum and Botrytis cinérea, and survival in strawberry. [ Links ]
Gams, W., Aa, H.A. van der; Plaats-Niterink, A.J. van der; Samson, R. A. and Stalpers, J. A. 1987. CBS course of mycology. Baarn, Netherlands; Centraalbureau voor Schimmelcultures. 3, 136 [ Links ]
Goes, L.B., Costa, A.B.L., Freiré, L.L.C. and Oliveria, N.T. 2002. Randomly amplified polymorphic DNA of Trichoderma isolates and antagonism against Rhizoctonia solani. Brazilian Archives Biology Technology. 45: 151-160. [ Links ]
Golenberg, E.M., Giannasi, D.E., Clegg, M.T., Smiley, C.J., Durbin, M, Henderson, D. and Zurawski, G. 1990. Chloroplast DNA sequence from a miocene Magnolia species. Nature 344: 656-658. [ Links ]
Grondona, L, Hermosa, M.R., Tejada, M., Gomis, M. D., Mateos, P.F., Bridge, P., Monte, E. and Garcia-Acha, I. 1997. Physiological and biochemical characterization of Trichoderma harzianum, a biological control agent against soilborne fungal plant pathogens. Appl. Environ. Microbiol. 63: 3189-3198. [ Links ]
Hadrys, H., Balick, M. and Schierwater, B. 1992. Application of random amplified polymorphic DNA (RAPD) in molecular ecology. Molecular Ecology. 1: 55-63. [ Links ]
Herds, C. 1984. Biologicol control. In: Current perspectives in microbial ecology Klug, M.J. Reddy, C. A. (Eds). American Society of Microbiology, Washington. 353-361. [ Links ]
Hjeljord, L.G. and Tronsmo, A. 1998. Trichoderma and Gliocladium in biocontrol: an overwiew,.In Kubicek, C.P. Harman, G.E.(Eds), Trichoderma and Gliocladium. Taylor & Francis, Ltd., London. United Kingdom. 135-151 [ Links ]
Jaccard, P. 1908. Nouvelles recherches surla distribution florale. Bui. Soc. Vaudoise Sci. Nat. 44: 223-270. [ Links ]
Johnson, L.E., Bond, C.J. and Fribourg, H. 1959. Methods for studying soil microflora-plant disease relationships. Minneapolis: Burgess Publishing Company. [ Links ]
Kucuk, C. and Kivanc, M. 2003. Isolation of Trichoderma Spp. and determination of their antifungal, biochemical and physiological features. Turkish Journal Biology. 27: 247-253. [ Links ]
Lorito, M., Woo, S.L., Garcia Fernandez, I., Colucci, G., Harman, G.E., Pintor-Toro, J.A., Filippone, E., Mucciflora, S., Lawrence, C.B., Zoina, A., Tuzun, S. and Scala, F. 1998. Genes from mycoparasitic fungi as a source for improving plant resistance to fungal pathogens. Procceedings National Academy Science, USA. 95: 7860-7865. [ Links ]
Lynch, J.M. 1990. Fungi as antagonists. In: "New Directions in Biological Control: Alternative for suppressing Agricultural Pests and Diseases". Liss, New York. 243-253. [ Links ]
Metcalf, D.D. and Wilson, C.C. 2001.The process of antagonism of Sclerotium cepivorum in white rot affected onion roots by Trichoderma koningii. Plant Pathology. 50: 249-257. [ Links ]
Möller, C, Biihler, T. and Dreyfuss, M.M. 1995. Intraspecific genetic diversity of Chaunopycnis alba detected by random amplified polymorphic DNA assay. Mycology Research 99: 681-688. [ Links ]
Papavizas, G.C. 1985. Trichoderma and Gliocladium: biology, ecology, and potential for biocontrol. Annual Review Phytopathology. 23: 23-54. [ Links ]
Rifai, M.A. 1969. A revision of the genus Trichoderma. Mycology. 116: 1-56. [ Links ]
Samuels, G. J., Chaverri, P., Farr, D.F. and McCray, E.B. 2004. USDA, Beltsville, USA. Trichoderma online systematic Botany and Mycology Laboratory, ARS, USDA. Retrieved September 20, 2004, from http://nt.arsgrin.gov/taxadescriptions/keys/TrichodermaIndex.cfm [ Links ]
Sandhu, G.S., Kline, B.C., Stockman, L. and Roberts, G.D. 1995. Molecular probes for diagnosis of fungal infections. Journal Clinical Microbiology. 35: 2270-2274. [ Links ]
Sherriff, C, Whelan, M.J., Arnold, G.M., Lafai, J.F., Brygoo, Y. and Bailey, J.A. 1994. Ribosomal DNA sequence analysis reveals new species grouping in the genus Colletotricum. Experimental Mycology 18: 121-138. [ Links ]
Sivan, A., Elad, Y. and Chet, I. 1984. Biological control effects of a new isolate of Trichoderma harzianum on Pythium aphanidermatum. Phytopathology 74: 498-501. [ Links ]
Tianhui, Z. and Dexun, Q. 1994. Antagonism of Trichoderma harzianum to Rhizoctonia solani. Journal Sichuan AgriculturalUniversity. 12: 11-11. [ Links ]
Van Belkum, A., Quint, W.G., de Pauw, B.E., Melchers, W.J.G. and Meis, J.F. 1993.Typing of Asprgillus species and Aspergillus fumigatus isolates by inerrepeat polymerase chain reaction. Journal Clinical Microbiology. 31:2502-2505. [ Links ]
Ward, J.H. 1963. Hierarchical grouping to optimize an objective function. Journal American Statistical Association. 58: 236-244. doi: 10.2307/2282967. [ Links ]
Williams, J.G.K., Kubelik, A.R., Livak, K.J., Rafalski, J.A. and Tingey, S.V. 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 18:6531-6535. [ Links ]














