Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Accesos
Accesos
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO
Compartir
Revista mexicana de ciencias pecuarias
versión On-line ISSN 2448-6698versión impresa ISSN 2007-1124
Rev. mex. de cienc. pecuarias vol.11 no.2 Mérida abr./jun. 2020 Epub 23-Oct-2020
https://doi.org/10.22319/rmcp.v11i2.5686
Articles
Characterization of Aspergillus flavus and quantification of aflatoxins in feed and raw milk of cows in Aguascalientes, Mexico
a Universidad Autónoma de Aguascalientes, Centro de Ciencias Agropecuarias, Av. Universidad 940, Col Cd. Universitaria, 20131, Aguascalientes, México.
b Universidad Autónoma de Aguascalientes. Centro de Ciencias Básicas. Aguascalientes, Aguascalientes, México.
c Instituto Politécnico Nacional. Centro de Biotecnología Genómica. Reynosa, Tamaulipas, México.
d Instituto Tecnológico El Llano, Municipio de El Llano, Aguascalientes, México.
e Universidad México Americana del Norte A. C. Coordinación de Investigación. Reynosa, Tamaulipas, México.
Contamination of agricultural and livestock products with aflatoxins (AF) is distributed worldwide. AFs are toxic, carcinogenic, and immunosuppressive; however, in Mexico, there is little information about Aspergillus flavus, the main fungus that produces them. The objective was to characterize the molecular and morphological, aflatoxigenic isolates of A. flavus and quantify the AFs in the feed and in the milk of Holstein cows in Aguascalientes (Mexico). A dairy production unit (2,749 cows) was selected for reasons of convenience, and monthly samples of food ingredients and total mixed ration (n= 267), raw milk (n= 288), and agricultural soil (n =40) were collected during 24 months and were cultivated (in PDA) using the pour plate technique with serial dilutions. The fungi were characterized using SEM, TLC and vapors of ammonium in coconut agar; the genes of calmodulin and a regulator of the biosynthetic pathway of AF, as well as the region of the internal spacer of the transcript, were sequenced. AFs were quantified in feed with HPLC and in milk, using ELISA. A total of 283 fungal isolates were characterized molecularly; of which 88 proved to be Aspergillus spp. Five of these were A. flavus with an aflatoxigenic capacity, and one was non-aflatoxigenic. 99.3 % of the samples of feed and 39.9 % of the milk samples exhibited detectable levels of AF (14.8 and 0,021 µg/kg). The cows ate daily 621 µg of AF and eliminated 0.09 % as AFM1 in milk. This suggests that the occurrence of aflatoxigenic A. flavus in the feed of dairy cows leads to a widespread contamination of the diets and food chain with AF.
Key words Aflatoxins; A. flavus; Dairy foods; Calmodulin gene; Regulatory gene of aflatoxin
La contaminación de productos agrícolas y pecuarios con aflatoxinas (AF) está distribuida mundialmente. Las AF son tóxicas, inmunodepresoras y carcinogénicas, pero en México es escasa la información sobre Aspergillus flavus, principal hongo que las produce. El objetivo fue caracterizar morfológica, molecular y aflatoxigénicamente aislados de A. flavus y cuantificar AF en pienso y en leche de vacas Holstein en Aguascalientes (México). Se seleccionó por conveniencia una unidad de producción lechera (2,749 vacas) y se recolectaron muestras mensuales (24 meses) de ingredientes alimenticios y ración total mezclada (n= 267), leche cruda (n= 288) y suelo agrícola (n= 40), las cuales se cultivaron (PDA) mediante vaciado en placa con diluciones seriadas. Los hongos se caracterizaron mediante MEB, TLC y vapores de amonio en agar coco; se secuenciaron los genes de calmodulina y regulador de la ruta biosintética de AF, así como la región de los espaciadores internos de la transcripción. Se cuantificaron AF en pienso con HPLC y en leche con ELISA. Se caracterizaron molecularmente 283 aislados fúngicos; 88 resultaron ser Aspergillus spp, de los que 5 fueron A. flavus con capacidad aflatoxigénica y uno no aflatoxigénico. El 99.3 % de las muestras de alimento y 39.9 % de las muestras de leche presentaron niveles detectables de AF (14.8 y 0.021 µg/kg). Las vacas ingirieron diariamente 621 µg de AF y eliminaron el 0.09 % como aflatoxina M1 en leche. Lo anterior sugiere que la ocurrencia A. flavus aflatoxigénico en el alimento de vacas lecheras conduce a una amplia contaminación por AF de las dietas y de la cadena alimenticia.
Palabras clave Aflatoxinas; A. flavus; Alimentos lácteos; Gen de calmodulina; Gen regulador de aflatoxinas
Introduction
Mexican bovine dairy is developed especially in a biogeographic province known as the Mexican Central High Plateau1. Within this region, dairy production units (DPU) face insufficiency issues in the production, economic profitability and safety of the products2. Among the food safety problems, the presence of aflatoxins (AF) has been highlighted because they cause a strong economic impact associated with the contamination of agricultural crops, a deterioration in the health of animals, a decline in productivity, and contamination of food of animal origin3. AF have hepatotoxic, nephrotoxic and immunosuppressive properties and are considered as the most powerful known natural carcinogens4. AF have been quantified in feed, milk and milk products intended for food for the Mexican human population5-7. This suggests that the presence of AFs in the feed of cows is a common problem in the DPUs that leads to contamination of the milk with the metabolites of AF.
When dairy cows consume agricultural products contaminated with AF, the enzymatic mechanisms bioactivate the mycotoxin through the formation of an epoxide that reacts with the cellular structures and nucleic acids, damaging their integrity4). The reactive epoxide can also be neutralized by conjugation with glutathione or its excretion as a metabolite that is eliminated in the milk mainly as aflatoxin M1 (AFM1); furthermore, AFM1 is considered as a toxic and carcinogenic agent for humans4.
AF are secondary metabolites of several filamentous fungi of the genus Aspergillus, which are distributed worldwide and contaminate a large variety of agricultural products, particularly cereals8. Also A. flavus has been identified in Mexico, both in agricultural soil and in corn grains9,10. Asexual reproduction of Aspergillus species has been described previously11,12, and so has its phylogeny, morphology and biological cycle8,13. Although Aspergillus flavus is considered to be the species with the greatest capacity for production of aflatoxins14, other strains of A. flavus without aflatoxygenic capacity, as well as other species of Aspergillus, produce AFs (A. parasiticus, A. nomius, A. pseudonomiu, A.arachidicola, A. bombycis, A. minisclerotigenes, A. pseudotamarii, and A. Togoensis)15. Aflatoxygenic capacity is expressed mainly under conditions of environmental stress16,17, as long as its genotype includes the information involved in the metabolic production chain of AF18,19. A strategy for the molecular identification of the different species of the genus Aspergillus is the use of the gene of calmodulin (CaM), the fragments corresponding to the region of the internal transcribed spacers (ITS) and the regulatory gene of the aflatoxin (AfIR) biosynthetic pathway8,20.
The objective of this work was to characterize the molecular and morphological, aflatoxigenic isolates of Aspergillus flavus obtained from dairy production units of the Central High Plateau of Mexico.
Material and methods
Study area
The present study was performed with a descriptive, longitudinal and non-experimental design. A dairy production unit was selected for convenience, using the non-probabilistic method, and was followed for 24 mo. The DPU was located in the Central High Plateau of Mexico (21°48'N, 102°03' W; 1986-2008 m asl), with a dry temperate and a semi-warm semi-dry climate, with summer rains, average temperature of 18.4 º C, an annual rainfall of 518.4 mm, and a maximum altitude of 2,300 m asl.
During the observation period, the DPU had an average of 2,749 Holstein cows housed in confinement within open-air pens enclosed with metal fences, shaded areas and freely accessible feeders. The cows were milked with automated equipment, obtaining an average daily production of 28.1 L per cow. The milk production is exported to agro-industrial plants in the region. Diets were developed as a total mixed ration (TMR) in a feed mill, where the corn silage and the concentrate were blended in mixer wagons. The TMR release is formulated to meet the nutritional requirements of dairy cows. The corn for silage is obtained directly from the agricultural areas of the UPL, while the protein-energy concentrate was purchased from the Local Association of Dairy Farmers of Aguascalientes, using as main ingredients canola, soybean, corn, rolled, sorghum, soybean meal, corn distilled dry grains, cottonseed, alfalfa and oats hay, as well as premixed vitamins and minerals.
Sample collection and handling
A total of 288 samples (1.0 kg) were obtained: total mixed ration (TMR), concentrated feed and corn silage were dried up in a forced air circulation oven (OF-22G-TECH, JEOI Lab Companion, Corea). The samples were pulverized (500-800 µm) in a continuously operating universal mill (MF series 10 Basic, IKA®-Werke, Germany) and stored in cooling system (4-5 °C) until processing (<7 d). During the (morning and evening) milking, a total of 288 samples of raw milk were obtained directly from the collector tank (500 ml) for each production batch (high, medium and low). The samples were transported in refrigeration and were preserved in freezing (-20 °C) until processing (<7 d).
Five agricultural plots were sampled with the help of a soil punch, choosing four sampling points on the surface of each plot of land and taking five 100 g sub-samples at each point, at a depth of 3-30 cm. The five sub-samples were gathered in a zip lock bag. Finally, they were screened (500-800 µ) and kept in refrigeration until processing.
Quantification of aflatoxins
Food samples were analyzed according to the official method 990.33 of the AOAC21, using solid phase columns (SPE; SupelcleanTM LC-18 SPE tube, Sigma-Aldrich, USA). The eluate extracted from the samples derivatized with trifluoroacetic acid was analyzed by HPLC (detection limit 2 µg/kg) with a fluorescence detector (binary Pump Vary Pro Star; FP 2020 detector, Varian Associates Inc., Victoria, Australia), a C18 column, and a column guard (LC-18 LC-18; Thermo Fisher Scientific, Massachusetts, USA). The quantification data were obtained using the Galaxie software (version 1.9.302.530), and AF concentrations were calculated using standard curves of purified AFs (Sigma-Aldrich, St. Louis, MO, USA). The AFM1 was quantified in raw milk with the competitive ELISA technique using a commercial kit (Ridascreen fast® aflatoxin M1 R-1121, R-Biopharm, Germany; detection range 0.005-0.08 µg/kg). The samples were homogenized and centrifuged according to the manufacturer's instructions. The absorbance was measured at 450 nm in an ELISA microplate reader (BioTek Instruments, Inc., USA). The results were interpreted upon the basis of the calibration curve made with purified 1 AFM (Sigma-Aldrich, St. Louis, MO, USA).
Fungal Isolation and characterization
The fungi were isolated using the pour plate technique with four serial dilutions in sterile peptone water22; the samples were seeded in a potato, dextrose and agar (PDA), malt extract with rose Bengal and Czapeck medium, and incubated (28 °C) in the darkness during 7 days. The colonies were isolated and purified, and the microscopic morphological characteristics consistent with the description of the genus were identified13. Isolates consistent with A. flavus8,13 were submitted to the process of fixating with glutaraldehyde (2%), gradual alcohol dehydration, and conditioning with a critical point dryer (Samdri Tousimis Research 795, Rockville, Meryland) and a metalizer (Desk II, Denton Vacuum, USA) were digitized in a scanning electron microscope (JEOL JSM-5900 LV,JEOL, USA) and conducted 10 measurements of each structure (stipe, gallbladder, spore, esclerocio and fialide), and using a JEOL software (Scanning Electron Microscope). The morphological structures of the isolates identified were compared against known strains of A. flavus, known as AF-36, AF-Cuatitlan (AF-C) and AF-Tamaulipas, (AF-T)(14.23). The strains obtained in this study were recorded at the NCBI (National Center for Biotechnology Information) with the respective access codes.
The aflatoxigenic capacity of the isolates was characterized by thin-layer chromatography (TLC)24; silica gel plates without fluorescence indicator were utilized (Z265829, Sigma-Aldrich, USA) activated in a high temperature oven (OF-22G-TECH, JEOI Lab Companion, Corea). The plates with purified standards of AFs (6636-50MG, Sigma Aldrich, USA) were placed inside of a chromatographic chamber with mobile phase chloroform - acetone - isopropanol (85:10:5, v/v/v) for an hour and a half. Eventually, the dry plates were visualized in a transilluminator. The technique ammonium vapors in coconut agar technique was also used according to the methodology described above25. Monosporic isolates were inoculated in sterile coconut agar and were left to incubate in the dark (30 °C, 5-7 d). Subsequently, ammonium hydroxide (200 µl; J.T. Baker, Mexico) at 25 % was added to the cover of the Petri dishes, and the distribution and the intensity of the color was observed. The presence of color was taken as indicative of AF production.
Molecular analysis
The genomic DNA of monosporic cultures of Aspergillus spp. was extracted according to previously standardized methods26. The quality of the obtained DNA was displayed with electrophoresis (45 min to 85 volts) in agarose gel (1%) with a TAE 1X damper and was quantified by comparing against known concentrations of DNA of phage λ (Thermo Fisher Scientific, MA USA). The DNA visualization was performed using a device for photoanalysis (Bio-Rad Gel Image®- Doctm XR, CA USA) with the Quantity One software version 4.6.7.
An amplification of a fragment corresponding to the region of internal spacers of the transcript (ITS; ITS1-5.8S-ITS2 RNAr) and ITS4 (5´-TCCTCCGCTTATTGATATG -3´) was carried out in accordance with the previously described protocols27,28; the gene of calmodulin (CaM) was amplified with the primers CMDA7F (5'-GCCAAAATCTTCATCCGTAG-3') and CDMA8R (5'-ATTTCGTTCAGAATGCCAGG-3') and the regulatory gene of the aflatoxin biosynthetic pathway (aflR) was amplified using the primers aflR-F (5'-GGGATAGCTGTACGAGTTGTGCCAG aflR-3') and aflR-R (5'-TGGKGCCGACTCGAGGAAYGGGT-3') of Eurofins Genomics, Louisville KY, USA. The polymerase enzyme Go-Taq (Promega, Madison, WI USA) and a model 9700 thermal cycler (Applied Biosystems) were utilized in the amplification. The PCR products obtained (ITS, calmodulin and aflR) were separated by electrophoresis in (1%) agarose gel and they were observed using as intercalating agents SYBR® Gold and Orange DNA Loading Dye (Thermo Fisher Scientific, MA USA); the resulting bands were observed in an image analysis device (BIO-RAD IMAGE® GEL Molecular- DOCTM XR CA, USA) with the Quantity One software (version 4.6.7). Molecular weight marker ladders (Axygen Biosciences, CA, USA) were included. PCR products were purified with the reagent ExoSAP-IT® PCR Product Cleanup (Afflymetrix, Thermo Fisher Scientific Inc. Santa Clara, California, USA). The purified PCR products were sequenced in forward and reverse chains with the dideoxy method29. The samples were injected into a sequencer (ABI 3730XL Genetic Analyzers), and the resulting sequences were recorded in an electropherogram. The electropherograms were visualized with the Chromas Lite software and were compared with the records of the NCBI using BLAST (Basic Local Alignment Search Tool).
Statistical analysis
The quantitative data of the total mixed ration, milk production, AF concentration in feed and milk, and of the measurements of the structure of each isolate were subjected to a one-way variance analysis (ANOVA), considering as separate factors the time of year and the level of milk production (high, medium or low) in which the cows were classified in batches. The differences between the means and the 95 % confidence intervals were determined using the honestly significant difference (HSD) Tukey’s test; P<0.05 was considered significant for all statistical analyses.
Results
Frequency of Aspergillus spp
It was identified a total of 283 fungal isolates; 56.8 % came from samples of feed for dairy cows, and the rest, from the agricultural land (Table 1). A total of 88 isolates (31.1 %) exhibited morphological features corresponding to those described for the genus Aspergillus. Figure 1 shows the conidiophores and conidia, and the characteristic metulae. Isolates were also found with a morphology compatible with the genera Penicillium, Fusarium, Rhizopus, Mucor, Cladosporium, Richoderma, Alternaria, Curvularia and Bipolaris. The proportions of each gender are shown in Table 1.
Table 1 Fungal genera identified in monthly samples * of a dairy production unit in the Central High Plateau of Mexico
| Gender | Soil | Corn silage | Feed concentrate | Total mixed ration | Total | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No. | % | No. | % | No. | % | No. | % | No. | % | |||||
| Samples | 40 | -- | 96 | -- | 96 | -- | 96 | -- | 328 | -- | ||||
| Aspergillus | 30 | 24.6 | 16 | 42.1 | 19 | 32.8 | 23 | 35.4 | 88 | 31.1 | ||||
| Penicillium | 23 | 18.9 | 3 | 7.9 | 4 | 6.9 | 9 | 13.8 | 39 | 13.8 | ||||
| Fusarium | 20 | 16.4 | 3 | 7.9 | 11 | 19.0 | 2 | 3.1 | 36 | 12.7 | ||||
| Rhizopus | 15 | 12.3 | 4 | 10.5 | 7 | 12.1 | 8 | 12.3 | 34 | 12.0 | ||||
| Mucor | 15 | 12.3 | 6 | 15.8 | 4 | 6.9 | 9 | 13.8 | 34 | 12.0 | ||||
| Cladosporium | 5 | 4.1 | 4 | 10.5 | 10 | 17.2 | 10 | 15.4 | 29 | 10.2 | ||||
| Trichoderma | 10 | 8.2 | 0 | 0.0 | 0 | 0.0 | 0 | 0.0 | 10 | 3.5 | ||||
| Alternaria | 4 | 3.3 | 2 | 5.3 | 1 | 1.7 | 2 | 3.1 | 9 | 3.2 | ||||
| Curvularia | 0 | 0.0 | 0 | 0.0 | 2 | 3.4 | 1 | 1.5 | 3 | 1.1 | ||||
| Bipolaris | 0 | 0.0 | 0 | 0.0 | 0 | 0.0 | 1 | 1.5 | 1 | 0.4 | ||||
| Total | 122 | 100 | 38 | 100 | 58 | 100 | 65 | 100 | 283 | 100 | ||||
*Sample of food ingredients: 24 months, in quadruplicate. Soil samples: 5 plots, in quadruplicate, 2 seasons.
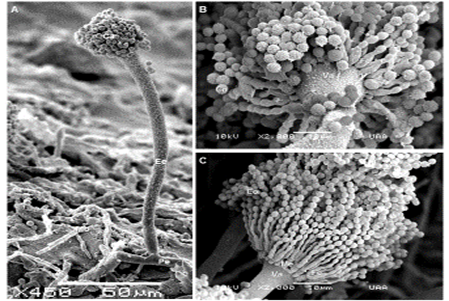

Panels: A) Conidiophore: Cc = conidial head, Ee = Stipe, Pe = foot cell, Mo = mycelium. B) conidial head uniseriate portions: Co = Conidium or spore; Fe = Phyalid, Va = Vesicle C) Biseriate conidial head: Co = Conidium, Fe = Phyalid , Va = Vesicle, i = Metula.
Figure 1 Aspergillus morphological structures
Only six isolates exhibited a morphology consistent with A. flavus and came from the feed concentrate (AC1, AC2 and AC3) and corn silage (EM1, EM2, EM3); when cultivated in PDA, they exhibited an olive green hue with a whitish periphery. Most of the isolates (5/6) exhibited the presence of sclerotia of dark brown color; only one did not manifest these structures. At a microscopic level, conidiophores of A. flavus exhibited irradiate and uniseriate conidial heads, the stipe with rough walls, the spherical vesicle, globe-shaped conidia with irregularities in their surface, and a whitish, septate mycelium (Figure 2).
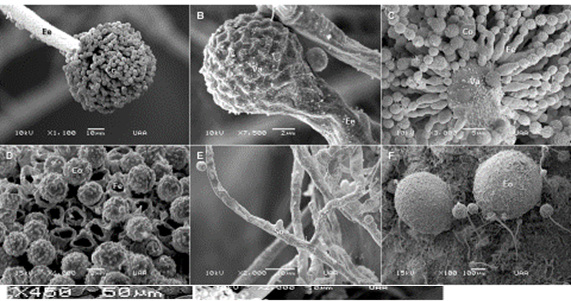
Panels: A) Cc = conidial head, Ee = stipe. B) Va = vesicle. C) Conidiophore. Fe = phyalids, Co = conidium. D) Cc = conidium, Fe = phyalid. E) Mo = mycelium, So = septum. F) Eo = Sclerotium.
Figure 2: Morphological structures of Aspergillus flavus
The six isolates of A. flavus from feed exhibited significant differences in the morphometry of their structures when compared against the control strains (AF-C and AF-T; Table 2). These strains presented conidial heads, with rough radiated stipe and spherical vesicles, while AF-C only exhibited (uniseriate) phyalids and (biseriate) AF-T phyalids metulae; the conidia were globe-shaped with irregularities on the surface. Four strains isolated were classified as L (long sclerotium, >400 µm), one as S (short sclerotium, <400 µm), and one as without sclerotium.
Table 2: Comparison of the dimensions of the morphological structures of Aspergillus flavus isolates obtained from corn silage (CS) and whole feed (WF) for dairy cows, as well as of the Cuautitlan (C) and Tamaulipas (T) strains
| Isolate | Stipe | Vesicle | Spore | Mean | Sclerotium | Mean | Fialide | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Mean | LL | UL | Mean | LL | UL | Mean | LL | UL | LL | UL | LL | UL | ||||||||||||
| AF-C | 237 | c | 183 | 291 | 33.6 | c | 27.5 | 39.7 | 3.3 | b | 3.1 | 3.4 | 418 | cd | 353 | 483 | 5.9 | ab | 4.9 | 6.9 | ||||
| AF-T | 646 | a | 564 | 727 | 76.6 | a | 67.7 | 85.5 | 2.8 | c | 2.7 | 2.9 | 453 | bc | 361 | 545 | 6.4 | ab | 4.9 | 7.8 | ||||
| AF-AC1 | 313 | ab | 241 | 386 | 54.3 | b | 46.3 | 62.2 | 3.1 | b | 3.0 | 3.3 | 592 | ab | 516 | 667 | 5.1 | b | 4.4 | 5.8 | ||||
| AF-AC2 | 373 | ab | 292 | 454 | 58.2 | b | 49.3 | 67.0 | 3.0 | bc | 2.8 | 3.1 | 428 | cd | 352 | 503 | 5.1 | b | 4.3 | 5.8 | ||||
| AF-AC3 | 441 | ab | 347 | 534 | 47.8 | ab | 37.5 | 58.0 | 3.1 | b | 2.9 | 3.3 | -- | -- | -- | 6.6 | ab | 5.5 | 7.7 | |||||
| AF-EM1 | 331 | ab | 255 | 407 | 50.5 | b | 42.1 | 58.8 | 3.5 | a | 3.4 | 3.7 | 268 | d | 193 | 344 | 4.8 | b | 4.1 | 5.5 | ||||
| AF-EM2 | 218 | c | 115 | 320 | 38.8 | ab | 27.6 | 50.1 | 3.2 | b | 3.1 | 3.4 | 672 | a | 597 | 748 | 5.4 | ab | 4.1 | 6.6 | ||||
| AF-EM3 | 320 | b | 239 | 401 | 47.3 | ab | 38.4 | 56.2 | 3.6 | a | 3.5 | 3.8 | 561 | ab | 505 | 617 | 6.9 | a | 6.0 | 7.9 | ||||
-- Without sclerotium- LL, UL, lower and upper limit of the confidence intervals.
abcd Means of columns with different letters show significant statistical differences (P<0.05).
The isolates identified as A. flavus were analyzed using TLC and ammonium vapors in coconut agar techniques in order to identify the presence of aflatoxin (Figure 3). Five isolates (AF-AC1, AF-AC2, AF-AC3, AF-EM2 and AF-M3) had a positive reaction to the presence of AF, while the other (AF-EM1) produced no aflatoxins, with both techniques. These results coincided with the amplified aflR gene.
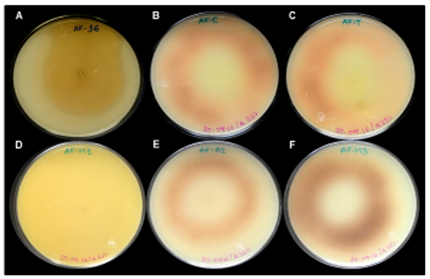
Panels: A) AF-36, non-aflatoxigenic (AF-). B) AF-Cuatitlán, and C) AF-Tamaulipas, aflatoxigenic (AF+). D) AF-EM1, AF- isolate from corn silage. E) AF-AC2, AF+ isolate from concentrated feed. F) AF-EM3, AF+ isolate from corn silage.
Figure 3 Intensity of color in a coconut agar medium in contact with ammonia vapors of Aspergillus flavus isolates after 4 d of growth
Molecular analysis
Of the 88 isolates morphologically identified as Aspergillus spp., 49 % were amplified for the calmodulin gene, and 31 % were amplified for ITS (Figure 4). The analysis of the sequences obtained showed that the species were A. oryzae (45.5 %), A. niger (10.2 %), A. ochraceus (3.4 %), A. pseudodeflectus (4.5 %), A. ustus (10.2 %), A. flavus (6.8 %), A. versicolor (5.6 %), A. nidulans (5.6 %), A. sublatus (4.5 %) and A. Sydowii (3.4 %).
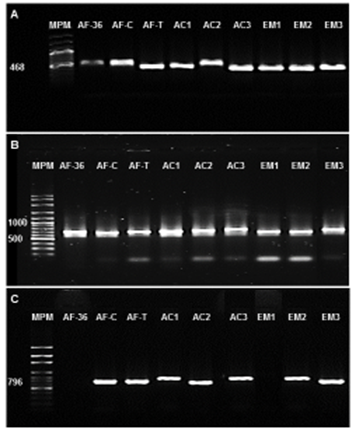
Rails: MWM) Molecular weight marker (100pb). Controls: Non-aflatoxigenic A. flavus (AF-36) and aflatoxigenic fungi (Cuautitlan, AF-C; Tamaulipas, AF-T). A. flavus isolates of corn silage (EM1, EM2 and EM3) and feed concentrate (AC1, AC2 and AC3) intended for dairy cows
Figure 4 Electrophoresis in an agarose gel at 1% of the PCR products amplified from the calmodulin gene (Panel A), internal transcribed spacer of the rDNA region 5.8S-ITS2 (Panel B), and regulatory gene of the aflatoxin biosynthetic pathway, aflR (Panel C)
The five local isolates identified as aflatoxigenic A. flavus amplified a fragment of 796 pb for the aflR gene. The analysis of the sequences of the control strains (AF-36, AF-C and AF-T) and the six isolates of A. flavus (AF-AC1, AF-AC2, AF-AC3, AF-EM1, AF-EM2 and AF-EM3) have been shown to have an identity percentage of over 90% with the isolates of A. flavus registered in the database of the NCBI (Table 3).
Table 3 Identity of the isolates of Aspergillus flavus obtained from diets for dairy cows in Aguascalientes, Mexico, compared to existing scripts at the National Center for Biotechnology Information (NCBI)a
| ID of the Isolate | Origin | Primerb | Access code | Coincidence (%) |
|---|---|---|---|---|
| Control- AF-36 | Control | ITS1 | LN482513.1 | 99 |
| ITS4 | KX550912.1 | 98 | ||
| Control+ C (AHY255) | Corn (Cuatitlan) | CMDA7F | AY974341.1 | 98 |
| CMDA8R | AY974340.1 | 96 | ||
| ITS1 | LC105688.1 | 100 | ||
| ITS4 | KT254587.1 | 99 | ||
| AFLRF | FN398160.1 | 100 | ||
| AFLRR | L32576.1 | 100 | ||
| Control+ T (AHY256) | Corn (Tamaulipas) | ITS1 | KX015990.1 | 100 |
| ITS4 | KF221065.1 | 99 | ||
| AFLRF | XM_002379905.1 | 100 | ||
| AFLRR | AF441435.2 | 99 | ||
| AC1 (AHY257) | Feed concentrate | ITS1 | KF434090.1 | 100 |
| ITS4 | KF434090.1 | 99 | ||
| AFLRF | AF441434.1 | 100 | ||
| AFLRR | XM_002379905.1 | 99 | ||
| AC2 (AHY258) | Feed concentrate | CMDA7F | AY974341.1 | 99 |
| ITS1 | KM285408.1 | 100 | ||
| ITS4 | EF409812.1 | 95 | ||
| AFLRF | XM_002379905.1 | 100 | ||
| AFLRR | FN398160.1 | 93 | ||
| AC3 (AHY259) | Feed concentrate | CMDA7F | XM_002374071.1 | 99 |
| CMDA8R | XM_002379905.1 | 99 | ||
| ITS1 | KF434090.1 | 99 | ||
| ITS4 | KT356196.1 | 97 | ||
| AFLRF | AF441435.2 | 99 | ||
| AFLRR | AF441434.1 | 99 | ||
| EM1 (AHY195) | Corn silage | CMDA7F | AY510451.1 | 95 |
| CMDA8R | XM_002374071.1 | 100 | ||
| ITS1 | HQ010119.1 | 99 | ||
| ITS4 | KX641192.1 | 99 | ||
| EM2 (AHY203) | Corn silage | ITS1 | KT356196.1 | 98 |
| ITS4 | LN482513.1 | 100 | ||
| AFLRF | MH752566.1 | 100 | ||
| AFLRR | KY769955.1 | 100 | ||
| EM3 (AHY204) | Corn silage | CMDA7F | XM_002374071.1 | 100 |
| CMDA8R | XM_002374071.1 | 100 | ||
| ITS1 | KX572367.1 | 99 | ||
| ITS4 | KT356196.1 | 99 | ||
| AFLRF | L32577.1 | 100 | ||
| AFLRR | MH752564.1 | 100 |
a NCBI (National Center for Biotechnology Information): https://blast.ncbi.nlm.nih.gov/
B CMDA = calmodulin; ITS = internal transcribed spacer; AFLR = regulator of the biosynthetic pathway; R = Reverse; F = forward.
Aflatoxins detected in feed and in milk
Of the 288 samples obtained from food, 286 (99.3 %) exhibited detectable levels of AFs (18.5 ± 3.7 µg/kg); of which 10.4 % exceeded the maximum limits allowed by the Mexican law (20 µg/kg). The presence of AFM1 (0.021 µg/kg) was detected in 39.9% of the 183 samples of raw milk, and 12.0% exceeded the maximum permissible limit used as a standard by the agro-industry which acquired the raw milk (0,050 µg/kg). The highest incidence of AF in feed occurred in summer and autumn (P<0.01), compared to winter and spring (17.4a ± 3.2 and 14.8ab ± 1.6 vs 8.1 bc ± 0.5 and 5.9c ± 0.5 µg/kg, respectively). In turn, the concentration of AFM1 was directly correlated with the concentration of AFs in the TMR in a double squared model (P<0.01; R2= 30.5 %). Also the presence of AFM1 in milk was significantly correlated with the level of milk production and the consumption of TMR; on average, the cows consumed daily 621 µg of aflatoxins in 42 kg of TMR; had a daily production of 26.2 L of milk with a total load of 0.603 µg of AFM1 ―which represented a great capacity of dairy cows to metabolize the AF―, and eliminated only a fraction (0.09 %; Table 4); in general, cows with a high production were exposed to the ingestion of larger amounts of AFs (658 µg/cow/day), and the presence and elimination of AFM1 in the milk of highly productive cows was greater than in cows with a medium and low production (0.22 µg/cow/d).
Table 4 Average exposure (± SE) of dairy cows to the natural contamination by aflatoxins in the total mixed ration (TMR) and average elimination of AFM1 per productive batch
| Batch |
Cows (No) |
Milk production (Kg/cow/day) |
TRM (Kg/cow/day) |
AF ingestion (µg/cow/day) |
AFM1 elimination | ||||
|---|---|---|---|---|---|---|---|---|---|
| (µg/kg | (µg/cow/day) | (%) | |||||||
| High | 1760 | 32.0a | 44.7a | 658a | ± 97.0 | 0.024a | ± 0.005 | 0.72 | 0.11 |
| Means | 606 | 20.2b | 36.4c | 546a | ± 83.4 | 0.022a | ± 0.005 | 0.44 | 0.08 |
| Low | 388 | 14.8c | 38.4b | 568a | ± 82.6 | 0.015b | ± 0.004 | 0.22 | 0.04 |
| Total | 2754 | 28.0 | 42.0 | 621 | ± 92.0 | 0.021 | ± 0.005 | 0.58 | 0.09 |
ab Different letters indicate significant differences between production batches P<0.05.
Discussion
In this two-year longitudinal study, the presence of Aspergillus flavus in the feed for dairy cows in the Central High Plateau of Mexico was proven. These isolates exhibited morphological, toxicological and molecular features which are consistent with the strains that have the capacity to produce aflatoxins. Furthermore, the accumulation of AFs in the feed and in the milk produced by cows was corroborated. Although information about the existence in Mexico of indigenous A. flavus populations with and without the ability to produce aflatoxins was already available, in general terms, this study adds for the first time the molecular, toxicological and morphometric characterization of A. flavus to the quantification of aflatoxins in the dairy, which is relevant for livestock production and public health.
Frequency of Aspergillus spp. and A. flavus
The fungal genus Aspergilus had the highest occurrence at the DPU (31 %), followed by Penicillium (13.8 %) and Fusarium (12.7 %). These genera have already been identified in Mexico in stored yellow corn9 and in corn hybrids intended for human and animal consumption30. It has been suggested that the frequent presence of micotoxigenic fungi and their toxins in corn is due to improper handling of the crop and to ineffective implementation of strategies for the control of infections5,31.
In this study, A. flavus was isolated from the feed for dairy cows at a ratio of 6.8% of the total identified Aspergillus spp. (6/88). This species was not identified in agricultural soil intended for the alternating production of fodder maize and oats. In Mexico, A. flavus has been identified in commercial corn kernels32 from soil for crops of grain corn10. This suggests that fungal populations of this species are distributed across the agricultural areas of Mexico.
In this study, the morphometric analysis showed that one-third of the isolates of A. flavus contained abundant short sclerotia (< 400 µm) and were therefore classified as S strains, while the rest were classified as L strains, as they had few but larger sclerotia (>400 µm). The importance of this morphology and its association with the aflatoxigenicity of S strains has already been pointed out; on the other hand, L strains contain both toxigenic and atoxigenic isolates33,34.
Molecular analysis
All of the 25 sequences obtained from the isolates identified morphologically as A. flavus exhibited a large percentage of identity (> 90%) with registered sequences of A. flavus strains. Amplified fragments were also obtained for the 468 pb CaM gene, for the 796 pb aflP gene, and for the ITS with a range of 600-800 pb (ITS1-5.8S-ITS2). It has been pointed out that the rDNA region of the internal transcribed spacers is the official DNA bar code for fungi because it is the most frequently sequenced marker and is a useful tool for the description of A. flavus species8. It has also been said that the CaM gene is able to distinguish between almost all species of Aspergillus8. On the other hand, the aflR gene is necessary for the transcription of most genes involved in the activation of enzymatic reactions that are necessary for the formation of aflatoxins35-37. Based on the above, this work integrates molecular identification to the morphological identification of six isolates of A. flavus obtained from the dairy production chain in the Central High Plateau region of Mexico.
Aflatoxigenic capacity
The six isolates identified as A. flavus were evaluated with TLC and vapors of ammonium in coconut agar in order to identify their capacity to produce aflatoxins; five were classified as aflatoxigenic fungi, and one was classified as a non-producer of aflatoxins. These results evidenced a behavior comparable to those obtained in other studies conducted in Mexico10 in which A. flavus isolates were identified as genetically related to A. flavus AF-3632. This non-aflatoxigenic strain has been employed in the development of biological control strategies to reduce aflatoxin production of aflatoxigenic strains in cotton-producing fields15. The above suggests that the main aflatoxigenic A. flavus populations are distributed worldwide across agricultural ecosystems, and that they coexist with strains that have no capacity to produce aflatoxins.
Aflatoxins detected in feed and in milk
A total of 99.3 % (286/288) of the samples of feed for dairy cows had detectable levels of AFs (18.5 ± 3.7 µg/kg), of which 10.4% exceeded the maximum limits allowed by Mexican law (20 µg/kg); furthermore, 39.9 % (73/183) of the raw milk samples exhibited the presence of AFM1. Comparable proportions of AF contamination have been detected in balanced feeds produced in Mexico5, in the total mixed rations of stables in Mexico38, and in samples of feed of Asian origin39. The incidence of AFM1 has been reported in raw bovine milk in Mexico6,7; also, incidence of AFM1 has been identified in milk for human consumption in various countries3,40-42. This suggests that the aflatoxin contamination of food for human and animal consumption is a global public health issue.
This study showed an association between the level of milk production of cows with the amount and concentration of AFM1 eliminated in the milk, in such a way that highly productive cows consumed a larger amount of feed and were exposed to larger amounts of aflatoxins in their feed; therefore, the total amount was larger, and the concentration of AFM1 in raw milk was higher (P<0.05). It has been pointed out that the AFM1 elimination rate in milk is influenced by the amount of feed consumed per day and the amount of milk produced per day by each animal, as well as by the stage of lactation43.
Conclusions and implications
This study proved morphologically, toxicologically and molecularly the occurrence of A. flavus in corn silage and in the total mixed ration, which led to the aflatoxin contamination of almost all diets of dairy cows, as well as to the residual presence of AFM1 in raw milk. Dairy cows were able to metabolize and eliminate more than 99 % of aflatoxin present in their diet through routes other than milk.
Acknowledgments
Financial support was provided by the National for Science and Technology Council of Mexico (Consejo Nacional de Ciencia y Tecnología; Basic Science Project: 178546 and Grant No. 514818) and the Autonomous University of Aguascalientes (Universidad Autónoma de Aguascalientes) (PIP project / SA 15-1). Also, thanks for the technical support of Araceli Adabache Ortiz and Marcelo Silva Briano.
REFERENCES
1. Domínguez-Domínguez O, Pérez-Ponce de León G. ¿La mesa central de México es una provincia biogeográfica? Análisis descriptivo basado en componentes bióticos dulceacuícolas. Rev Mex Biodivers 2009;80(3):835−52. [ Links ]
2. Romo CE, Valdivia AG, Carranza RG, Camara J, Zavala MP, Flores E, et al. Brechas de rentabilidad económica en pequeñas unidades de producción de leche en el altiplano central mexicano. Rev Mex Cienc Pecu 2014;5(3):273−89. [ Links ]
3. Rahimi E, Bonyadian M, Rafei M, Kazemeini HR. Occurrence of aflatoxin M1 in raw milk of five dairy species in Ahvaz, Iran. Food Chem Toxicol 2010;48(1):129−31. [ Links ]
4. IARC. International Agency for Research on Cancer. Monographs on the evaluation of carcinogenic risks to humans, vol. 82. Working Group on the Evaluation of Carcinogenic Risks to Humans II. Lyon, Press. 2002. [ Links ]
5. Flores CM, Hernández LB, Vázquez J. Contaminación con micotoxinas en alimento balanceado y granos de uso pecuario en México en el año 2003. Rev Mex Cienc Pecu 2006;44(2):247−256. [ Links ]
6. Pérez J, Gutiérrez R, Vega S, Díaz G, Urbán G, Coronado M, et al. Ocurrencia de aflatoxina M1 en leches cruda, ultrapasteurizada y orgánica producidas y comercializadas en el Altiplano Mexicano. Rev Salud Anim 2008;30(2):103−109. [ Links ]
7. Landeros P, Noa M, López Y, González DG, Noa E, Real M, et al. Niveles de aflatoxina M1 en leche cruda y pasteurizada comercializada en la zona metropolitana de Guadalajara, México. Rev Salud Anim 2012;34(1):40−45. [ Links ]
8. Samson RA, Visagie CM, Houbraken J, Hong SB, Hubka V, Klaassen CHW, et al. Phylogeny, identification and nomenclature of the genus Aspergillus. Stud Mycol 2014;78:141−173. [ Links ]
9. Hernández-Delgado S, Reyes-López MÁ, García-Olivares JG, Mayek-Pérez N, Reyes-Méndez CA. Incidencia de hongos potencialmente toxígenos en maíz (Zea mays L.) almacenado y cultivado en el norte de Tamaulipas, México. Rev Mex Fitopatol 2007;25(2):127−133. [ Links ]
10. Ortega-Beltran A, Jaime R, Cotty PJ. Aflatoxin-producing fungi in maize field soils from sea level to over 2000 masl: A three-year study in Sonora, Mexico. Fungal Biol-Uk 2015;119(4):191−200. [ Links ]
11. Frisvad JC, Larsen TO. Chemodiversity in the genus Aspergillus. Appl Microbiol Biot 2015;99(19):7859−7877. [ Links ]
12. Varga J, Szigeti G, Baranyi N, Kocsubé S, O’Gorman CM, Dyer PS. Aspergillus: sex and recombination. Mycopathologia 2014;178(5-6):349−362. [ Links ]
13. Klich MA. Identification of common Aspergillus species. First edition. Lousiana, USA: Centraalbureau voor Schimmelcultures, Utrecht, the Netherlands; 2002. [ Links ]
14. Bayman P, Cotty PJ. Genetic diversity in Aspergillus flavus: association with aflatoxin production and morphology. Can J Bot 1993;71(1):23−31. [ Links ]
15. Ehrlich KC, Cotty PJ . An isolate of Aspergillus flavus used to reduce aflatoxin contamination in cottonseed has a defective polyketide synthase gene. Appl Microbiol Biot 2004;65(4):473−8. [ Links ]
16. Cotty PJ , Jaime-Garcia R. Influences of climate on aflatoxin producing fungi and aflatoxin contamination. Int J Food Microbiol 2007;119(1-2):109−15. [ Links ]
17. Cheli F, Campagnoli A, Dell’Orto V. Fungal populations and mycotoxins in silages: From occurrence to analysis. Anim Feed Sci Tech 2013;183(3):1−16. [ Links ]
18. Ehrlich KC, Montalbano BG, Cotty PJ . Sequence comparison of aflR from different Aspergillus species provides evidence for variability in regulation of aflatoxin production. Fungal Genet Biol 2003;38(1):63−74. [ Links ]
19. Reis TA, Baquião AC, Atayde DD, Grabarz F, Corrêa B. Characterization of Aspergillus section Flavi isolated from organic Brazil nuts using a polyphasic approach. Food Microbiol 2014;42:34−39. [ Links ]
20. Yu J. Current understanding on aflatoxin biosynthesis and future perspective in reducing aflatoxin contamination. Toxins 2012;4(11):1024−1057. [ Links ]
21. Scott PM. Natural toxins. Official Methods of Analysis of the Association of Analytical Chemistry. 1995;49:1-30. [ Links ]
22. Iheanocho HE, Njobeh PB, Dutton FM, Steenkamp PA, Steenkamp L, Mthombeni JQ, et al. Morphological and molecular identification of filamentous Aspergillus flavus and Aspergillus parasiticus isolated from compound feeds in South Africa. Food Microbiol 2014;44:180−184. [ Links ]
23. de Luna-López MC, Valdivia-Flores AG, Jaramillo-Juárez F, Reyes JL, Ortiz-Martínez R, Quezada-Tristán T. Association between Aspergillus flavus colonization and aflatoxins production in immature grains of maize genotypes J Food Sci Eng 2013;3(12):688−698. [ Links ]
24. Fani SR, Moradi M, Probst C, Zamanizadeh HR, Mirabolfathy M, Haidukowski M, et al. A critical evaluation of cultural methods for the identification of atoxigenic Aspergillus flavus isolates for aflatoxin mitigation in pistachio orchards of Iran. Eur J Plant Pathol 2014;140(4):631−642. [ Links ]
25. Saito M, Machida S. A rapid identification method for aflatoxin-producing strains of Aspergillus flavus and A. parasiticus by ammonia vapor. Mycoscience 1999;40(2):205−208. [ Links ]
26. Hoffman CS, Winston F. A ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli. Gene 1987;57(2-3):267−272. [ Links ]
27. White TJ, Bruns T, Lee S, Taylor J. Amplification and direct sequencing of fungal Ribosomal RNA genes for phylogenetics. In: Innis MA, et al, editors. PCR protocols: a guide to methods and application New York, USA. Academic Press; 1990:315−322. [ Links ]
28. Shapira R, Paster N, Eyal O, Menasherov M, Mett A, Salomón R. Detection of aflatoxinogenic molds in grains by PCR. Appl Environ Microbiol 1996;62:3270-3273. [ Links ]
29. Sanger F, Nicklen S, Coulson AR. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA. 1977;74(12):5463-5467. [ Links ]
30. Montes GN, Reyes MCA, Montes RN, Cantu AMA. Incidence of potentially toxigenic fungi in maize (Zea mays L.) grain used as food and animal feed. CyTA-J Food 2009;7(2):119−25. [ Links ]
31. Martínez HY, Hernández S, Reyes CA, Vázquez G. El Género Aspergillus y sus Micotoxinas en Maíz en México: Problemática y Perspectivas. Rev Mex Fitopatol 2013;31(2):126−146. [ Links ]
32. Ortega‐Beltran A, Grubisha LC, Callicott KA, Cotty PJ. The vegetative compatibility group to which the US biocontrol agent Aspergillus flavus AF36 belongs is also endemic to Mexico. J Appl Microbiol 2016;120(4):986−98. [ Links ]
33. Cotty PJ. Virulence and cultural characteristics of two Aspergillus flavus strains pathogenic on cotton. Phytopathology 1989;79(7):808−814. [ Links ]
34. Probst C, Bandyopadhyay R, Cotty PJ. Diversity of aflatoxin-producing fungi and their impact on food safety in sub-Saharan Africa. Int J Food Microbiol 2014;174:113−122. [ Links ]
35. Yu J, Chang PK, Ehrlich KC, Cary JW, Bhatnagar D, Cleveland TE, et al. Clustered pathway genes in aflatoxin biosynthesis. Appl Environ Microbiol 2004;70(3):1253−1262. [ Links ]
36. Bhatnagar D, Cary JW, Ehrlich K, Yu J, Cleveland TE. Understanding the genetics of regulation of aflatoxin production and Aspergillus flavus development. Mycopathologia 2006;162(3):155−166. [ Links ]
37. Amare MG, Keller NP. Molecular mechanisms of Aspergillus flavus secondary metabolism and development. Fungal Genet Biol 2014;66(3):11−18. [ Links ]
38. Reyes W, Martínez SP, Isaías VH, Nathal MA, De Lucas E, Rojo F. Aflatoxinas totales en raciones de bovinos y AFM1 en leche cruda obtenida en establos del estado de Jalisco, México. Tec Pecu Mex 2009;47(2):223−230. [ Links ]
39. Ruadrew S, Craft J, Aidoo K. Occurrence of toxigenic Aspergillus spp. and aflatoxins in selected food commodities of Asian origin sourced in the West of Scotland. Food Chem Toxicol 2013;55:653−658. [ Links ]
40. Cano-Sancho G, Marin S, Ramos AJ, Peris-Vicente J, Sanchis V. Occurrence of aflatoxin M1 and exposure assessment in Catalonia (Spain). Rev Iberoam Micol 2010;27:130−135. [ Links ]
41. Shundo L, Navas SA, Lamardo LCA, Ruvieri V, Sabino M. Estimate of aflatoxin M1 exposure in milk and occurrence in Brazil. Food Control 2009;20(7):655−657. [ Links ]
42. Iha MH, Barbosa CB, Okada IA, Trucksess MW. Occurrence of aflatoxin M1 in dairy products in Brazil. Food Control 2011;22(12):1971−1974. [ Links ]
43. Masoero F, Gallo A, Diaz D, Piva G, Moschini M. Effects of the procedure of inclusion of a sequestering agent in the total mixed ration on proportional aflatoxin M1 excretion into milk of lactating dairy cows. Anim Feed Sci Tech 2009;150(1-2):34−45. [ Links ]
Received: September 04, 2018; Accepted: April 29, 2019











 texto en
texto en 


