Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Similars in
SciELO
Similars in
SciELO
Share
Revista mexicana de fitopatología
On-line version ISSN 2007-8080Print version ISSN 0185-3309
Rev. mex. fitopatol vol.34 n.2 Texcoco 2016
https://doi.org/10.18781/R.MEX.FIT.1601-1
Scientific articles
Detection of Pineapple mealybug wilt-associated virus 1 and 3 in Mexico
1 Postgrado en Fitosanidad-Fitopatología, Colegio de Postgraduados-Campus Montecillo, km 36.5 Carr. México-Texcoco. Montecillo, Estado de México, CP 56230
2 INIFAP-Campo Experimental Cotaxtla, km 34.5 Carr. Federal Veracruz-Córdoba, Medellín de Bravo, Ver., CP 94270
3 Postgrado en Fitosanidad-Fitopatología, Colegio de Postgraduados-Campus Montecillo, km 36.5 Carr. México-Texcoco. Montecillo, Estado de México, CP 56230
In El Bajo Papaloapan, the main producing area of pineapple of Mexico, leaves with typical symptoms of viral infection consisting in chlorosis, flaccidity, reduced growth and reddening were collected. By RT-PCR with specific primers for the hsp70 gene and subsequent sequencing were detected Pineapple mealybug wilt virus associated-virus 1 (PMWaV-1) and Pineapple mealybug wilt virus associated-3 (PMWaV-3). From the sequences obtained a tree was done with sequences from different regions of the world available in GenBank in order to know their similarity. The sequence obtained from the Mexican isolate PMWaV-1 was genetically related to the sequences of isolates from Cuba, Taiwan, Thailand and Hawaii and more distant from the Australian isolate. The sequence obtained for the Mexican isolate PMWaV-3 was more related to isolates from Hawaii, Cuba, Australia and Taiwan and more distant from the Thailand isolate. This is the first report of the presence of these two viruses in Mexico.
Key words: ampeloviruses; mealybug; pineapple red wilt; phylogeny
En El Bajo Papaloapan, principal zona productora de piña, se colectaron hojas con síntomas de clorosis, flacidez, reducción del crecimiento y enrojecimiento foliar típicos de infecciones virales en este cultivo. Mediante RT-PCR con indicadores específicos para el gen hsp70 y posterior secuenciación, fueron detectados los virus Pineapple mealybug wilt associated-virus 1 (PMWaV-1) y Pineapple mealybug wilt associated-virus 3 (PMWaV-3). A partir de las secuencias obtenidas se construyó un árbol con secuencias procedentes de diferentes regiones del mundo disponibles en el GenBank para determinar su similitud. La secuencia obtenida del aislamiento mexicano del PMWaV-1 resultó ser más próxima genéticamente a secuencias de aislamientos procedentes de Cuba, Taiwán, Tailandia y Hawai y más distante del aislamiento australiano. La secuencia obtenida para el aislamiento mexicano del PMWaV-3 se encontró más relacionada con los aislamientos de Hawai, Cuba, Australia y Taiwán y más distante del aislamiento tailandés. Este es el primer reporte de la presencia de estos dos virus en México.
Palabras clave: ampelovirus; piojo harinoso; marchitez roja de la piña; filogenia
El Bajo Papaloapan is the main producing area of pineapple (Ananas comosus (L.) Merr.) of Mexico which includes the regions of Loma Bonita (Oaxaca), Linda Vista, Villa Azueta, El Zopilote, Isla, Juan Rodríguez Clara and Los Tigres (Veracruz) where this crop is an important economic activity. At global level a viral complex associated with pineapple is known (Sether et al., 2010), consisting of Pineapple mealybug wilt associated virus-1 (PMWaV-1), Pineapple mealybug wilt associated virus-2 (PMWaV-2) and Pineapple mealybug wilt associated virus-3 (PMWaV-3) transmitted by mealybugs Dysmicoccus neobrevipes and Dysmicoccus brevipes (Sether et al., 1998). Mealybug wilt of pineapple (MWP) is one of the most destructive diseases of this species in many parts of the world and it is associated with the infection of PMWaV-2 combined with mealybug feeding (Sether et al., 2005). In Hawaii, it was found that mealybug feeding alone or in combination with PMWaV-1 or PMWaV-3 does not cause MWP requiring the presence of PMWaV-2 for this to occur (Sether and Hu, 2002). Symptoms of the disease consist of a severe dieback, reddish coloration of leaves and wilting which together cause the collapse of mature plants (Carter, 1945). It has also been reported a brown coloration of the affected plants, flaccidity, downward curling of the leaf tips, reduction of the root system and fruits with fibrous pulp and tasting sour (Borroto et al., 1998). Detection of these viruses has been done by tissue blot immunoassay and by RT-PCR with primers specific to the hsp70 gene (Sether et al., 2001; Sether et al., 2005). In the producing area of pineapple of El Bajo Papaloapan producers and technicians have been observed plants with symptoms of flaccidity, chlorosis, light reddening or wilt without root rot or necrosis of vascular bundles randomly distributed within the plots. Since no damage of the root system or the vascular bundles has been observed in the affected plants it is possible that these symptoms are associated with PMWaV-1, or PMWaV-2 or PMWaV-3, causal agents of the MWP one of the most important diseases of this crop in the world. So, the objective of this study was to determine the presence of these viruses in the region and to know their relationships with several isolates of the world.
Materials and methods
Sample collection
In plots with different production system located in the region of Loma Bonita (Oaxaca), Linda Vista, Sabaneta, Ejido, Isla and Los Tigres (Veracruz), a directed sampling was conducted in July 2011 to collect leaf tissue of pineapple plants var. Cayena or the MD2 hybrid with symptoms of flaccidity, wilting, reddish or tan coloration of leaves, general reduction of growth associated with pineapple mealybug wilt as well as asymptomatic plants. All sampled plants were revised in search of mealybugs and in those showing wilting, roots were dug up to observe them and discard that this flaccidity was associated with root rot or necrosis of vascular bundles. The plant tissue was placed in polyethylene bags and labeled for its transportation to the laboratory.
RNA extraction and RT-PCR
Total RNA extraction from collected leaves was performed with Trizol® method (AFGC, 2002), according to the manufacturer's instructions. The RNA obtained was employed to generate cDNA and use it to realize PCR (MMLV enzyme and Taq polymerase from Promega® were used) with specific primersofasegmentofth ehsp70genfrom PMWaV-1 (5'-ACAGGAAGGACAACACTCAC3'/5'-CGCACAAACTTCAAGCAATC-3'), PMWaV-2 (5'-CATACGAACTAGACTCATACG3'/5'-TCATTCCACTCACTTATCGTTG-3') and PMWaV-3 (5'-AGTTCACTGTAGATTTCGGA3'/5'-ATTCATGGATGTGTATCG-3'), that amplify 589, 609 and 495 bp, respectively (Sether et al., 2005; Sether et al., 2009). PCR products were analyzed by electrophoresis on a 2 % agarose gel containing ethidium bromide. As reference a molecular weight marker of 100 bp was used, the gels were visualized on a UV transilluminator Syngene© mod. Gene Snap. They were then purified using the Wizard SV gel and PCR clean-up system© and sequenced at the Institute of Biotechnology, UNAM.
Sequence analysis
Sequences were used to obtain Contigs with DNA Baser Sequence Assembler v2 (www.dnabaser.com). Contigs were after analyzed using the Basic Local Alignment Search Tool (BLAST) of NCBI (National Centre for Biotechnology Information, http://www.ncbi.nlm.nih.gov/BLAST).
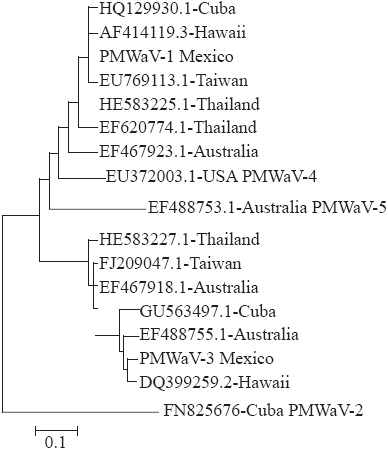
The resulting sequences were compared with the 15 existing sequences found in GenBank (NCBI), with the following accession of PMWaV-1: Cuba (HQ129930.1), Taiwan (EU769113.1), Thailand (I583225.1 & EF620774.1), Hawaii (AF414119.3) and Australia (EF467923.1); PMWaV-2: Cuba (FN825676); PMWaV-3: Cuba (GU563497.1), Hawaii (DQ399259.2), Australia (EF488755.1 & EF467918.1), Taiwan (FJ209047.1) and Thailand (HE583227.1); PMWaV-4: USA (EU372003.1); and PMWaV5: Australia (EF488753.1).
In order to obtain a visual representation of sequence variability between all isolates, we determinate identity percentage; Neighbor-joining cluster analysis was used to generate rootless trees of the relationships between the 15 sequences. The data set of each of these trees was bootstrapped 1000 times.
Results
During the sampling no plants with typical symptoms of pineapple mealybug wilt described or shown in photographs in different publications were observed. The most frequently observed alterations were flaccidity, chlorosis, and reddish coloration in the center of leaves (Figure 1). No mealybugs were observed in any of the collected plants as well as no visible damage or necrosis in the roots were recorded in those showing wilting.
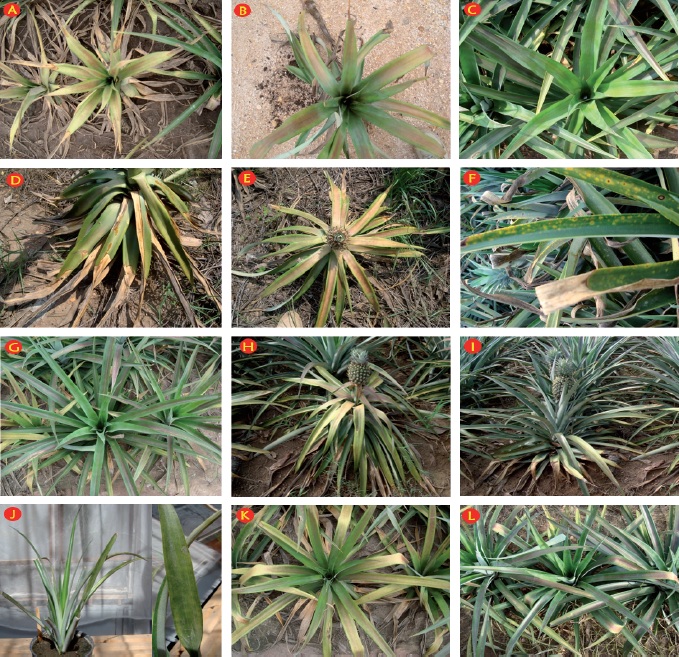

Figure 1 Pineapple plants grown in the region of El Bajo Papaloapan with putative symptoms of mealybug wilt: A: apex leaf necrosis, chlorosis and reddish coloration; B: reddish coloration in the central part of the leaves and downward curvature of the apex; C: Asymptomatic plant; D: flaccidity and apical necrosis of leaves; E: flaccidity, severe chlorosis and reddening of the central part of the leaves; F: yellow chlorotic spots, some with necrotic center; G: Asymptomatic plant; H: wilting, chlorosis in leaves of the middle part and reddening of upper leaves; I: Asymptomatic plant; J: reduction of growth and irregular chlorotic spots; K: chlorosis and reddening of lower leaves; L: reduction of growth, chlorosis of upper leaves, reddening and flaccidity of lower leaves. Plants D and E were of MD2, the rest of Cayena.
PMWaV-1 was found in one sample (called as PMWaV-1Mexico) showing reduction of growth and irregular chlorotic spots (Figure 1J) and PMWaV-3 was present in one sample too (called as PMWaV-3Mexico) with chlorosis and reddening of lower leaves (Figure 1K), both collected at Isla, Veracruz. PMWaV-2 was not detected in any of the samples analyzed in this study. Both genomic fragments of PMWaV-1Mexico (GenBank access KC800714.1) and PMWaV-3Mexico (GenBank access KC800715.1) sequenced in this work are located within the hsp70 gene, which encodes the heat shock 70-like protein into the ORF 3. PMWaV-1 sequence (589pb) corresponded to the 8.386 8.974 positions of the genome of a Hawaiian PMWaV-1 isolate (GenBank accession AF414119), and PMWaV-3 sequence (324pb) corresponded to the 7.004 7.327 positions of the genome of a Hawaiian PMWaV-3 isolate (GenBank accession DQ399259.2).
The isolate PMWaV-1Mexico showed a 100 % of identity with Cuban isolate and 98 % higher than the rest of isolates, except with Australian isolate (88.6 % of identity). By the other hand, the isolate PMWaV-3Mexico showed a 95.5 % identity higher with the rest of isolates in its group, except with Thailand isolate (88.2 % of identity). Figure 2 shows the similarity of Mexican isolates in relation with those reported in GenBank used for comparison. PMWaV-1Mexico and PMWaV3Mexico grouped with groups 1 and 3, respectively, but no with groups 2, 4 and 5.
Discussion
Mealybug wilt of pineapple (MWP) is a disease characterized by a severe dieback, reddening of leaves, wilt and collapse of mature plants associated with the viruses Pineapple mealybug wilt associated virus-1 (PMWaV-1), and Pineapple mealybug wilt associated virus-3 (PMWaV-3) (Sether and Hu, 2000). In this study it was not observed in any of the plants analyzed the typical symptoms described in literature for MWP. PMWaV-1 was detected in a plant showing reduction of growth and irregular chlorotic spots on all leaves, especially the mature ones (Figure 1), these symptoms are different from those observed in field by Hu et al., (1997) who also mentioned that a high percentage of plants with MWP are infected with PMWaV-1 while others are asymptomatic. PMWaV-1 is limited to phloem; it is not transmitted mechanically and in field is spread by the mealybug Dysmicoccus brevipes and D. neobrevipes (Sether et al., 1998). In this study did not find mealybugs in any of the sampled plants including the one that tested positive for PMWaV-1, however, this insect is a common pest in the area of study and this form of transmission in field is not discarded; it is possible that its spread may also occur in a vegetative form too (Sether et al., 2005) since this virus causes asymptomatic infections (Hu et al., 1997). In the area of study a reduction of the crop yield has not been observed; however, the presence of PMWaV-1 in the region can eventually affect the production if vector management and water supply is neglected since it has been shown that infection by this virus and limited irrigation produce a negative effect (6.7 and 4.2 %, respectively) and an additive effect (13.4 %) on size and weight of the pineapple fruit (Sether and Hu, 2001). The isolate of PMWaV-1 detected in this study is similar to that reported in Cuba (Figure 2) due perhaps to the highest trade exchange of propagative material between the two countries and least similar to isolates from Australia and Hawaii related to differences in systems production, cultivated plant material and prevailing environmental conditions in all three cases (Sether et al., 2010).
Like PMWaV-1, PMWaV-3 does not induce MWP by itself or together with the mealybug feeding in Hawaii (Sether et al., 2005) and it is unknown what is its effect on the production of pineapple (Sether et al., 2009). Only one plant from the same location that showed different symptoms resulted positive for PMWaV-3. This plant had a reddish coloration in the central part of the mature leaves while young ones had a dark green, and in general did not show a decrease in growth nor flaccidity. As already mentioned, mealybug wilt of pineapple (MWP) is one of the most devastating diseases of this crop worldwide and has been associated with five different ampeloviruses (designated as Pineapple mealybug wilt associated virus-1 to 5) (Gambley et al., 2008). However, it has been shown that the typical symptoms of this disease are only produced when infection simultaneously occurs of Pineapple mealybug wilt associated virus-2 and feeding of Dysmicoccus brevipes and/or D. neobrevipes (Sether et al., 1998). In this study mealybugs were not observed and PMWaV-2 was no detected in the samples analyzed, which is similar to that reported in Hawaii where the incidence of this virus ranges from 0 to 20 % depending on the production system and the hybrid used (Sether et al., 2001) and can cause up to 100 % losses of the fruits when it is present (Sether and Hu, 2001). However, it is necessary to analyze a larger number of samples to know the situation of PMWaV-2 in the area of study. Since PMWaV-1Mexico and PMWaV-3Mexico are more distant and less similar to the Australian and Thailand isolates, respectively, it could suggest that these isolates had different geographical origins. However, it would be necessary to study other genes and largest number of isolates to confirm this hypothesis.
Conclusions
Pineapple mealybug wilt associated virus-1 (PMWaV-1) was detected in a pineapple plant showing reduction of growth and irregular chlorotic leaf spots and Pineapple mealybug wilt associated virus-3 (PMWaV-3) in one plant showed a reddish coloration in the center of mature leaves. The PMWaV-1Mexico isolate is similar to one reported in Cuba and somewhat different from that of Australia and Hawaii, while PMWaV-3Mexico isolate show genetic differences with other isolates of PMWaV-3 reported in the Genbank.
Acknowledgements
We thank for the partial funding granted by the Comité Veracruzano de Productores de Piña A.C. for conducting this investigation.
REFERENCES
AFGC. 2002. Arabidopsis functional genomics consortium. Total RNA isolation. http://www.arabidopsis.org/portals/masc/AFGC/RevisedAFGC/site2RnaL.htm . Revised on Sept. 09 2010. [ Links ]
Borroto EG, Cintra M, González J, Borroto C and Oramas P. 1998. Pineapple plants (Ananas comosus cv. Smooth Cayenne) affected with pineapple mealybug wilt in Cuba. Plant Disease 82:263. http://dx.doi.org/10.1094/PDIS.1998.82.2.263C [ Links ]
Carter W. 1945. Some etiological aspects of mealybug wilt. Phytopathology 35:305-315. [ Links ]
Gambley CF, Steele V, Geering ADW and Thomas JE. 2008. The genetic diversity of ampeloviruses in Australian pineapples and their association with mealybug wilt disease. Australasian Plant Pathology 37:95-105. http://dx.doi.org/10.1071/AP07096 [ Links ]
Hu JS, Sether DM, Liu XP, Wang M, Zee F and Ullman D. 1997. Use of a tissue blotting immunoassay to examine the distribution of pineapple closterovirus in Hawaii. Plant Disease 81:1150-1154. http://dx.doi.org/10.1094/PDIS.1997.81.10.1150 [ Links ]
Sether DM, Ullman DE and Hu JS. 1998. Transmission of pineapple mealybug wilt associated virus by two species of mealybugs (Dysmicoccus spp.). Phytopathology 88:1224-1230. http://dx.doi.org/10.1094/PHYTO.1998.88.11.1224 [ Links ]
Sether DM and Hu JS. 2000. A closterovirus and mealybug exposure are both necessary components for mealybug wilt. Phytopathology 90:S71. http://dx.doi.org/10.1094/PHYTO.2000.90.6.S1 [ Links ]
Sether DM and Hu JS. 2001. The impact of Pineapple mealybug wilt-associated virus-1 and reduced irrigation on pineapple yield. Australasian Plant Pathology 30:31-36. http://dx.doi.org/10.1071/AP00060 [ Links ]
Sether DM, Karasev AV, Okumura C, Arakawa C, Zee F, Kislan MM, Busto JL and Hu JS. 2001. Differentiation, distribution, and elimination of two different Pineapple mealybug wilt-associated viruses found in pineapple. Plant Disease 85:856-864. http://dx.doi.org/10.1094/PDIS.2001.85.8.856 [ Links ]
Sether DM and Hu JS. 2002. Closterovirus infection and mealybug exposure are necessary for the development of mealybug wilt of pineapple disease. Phytopathology 92:928-935. http://dx.doi.org/10.1094/PHYTO.2002.92.9.928. [ Links ]
Sether DM, Melzer MJ, Busto J, Zee F and Hu JS. 2005. Diversity and mealybug transmissibility of ampeloviruses in pineapple. Plant Disease 89:450-456. http://dx.doi.org/10.1094/PD-89-0450 [ Links ]
Sether DM, Melzer MJ, Borth WB and Hu JS. 2009. Genome organization and phylogenetic relationship of Pineapple mealybug wilt associated virus-3 with family Closteroviridae members. Virus Genes 38: 414-420. http://dx.doi.org/10.1007/s11262-009-0334-5 [ Links ]
Sether DM, Borth WB, Melzer MJ and Hu J. 2010. Spatial and temporal incidences of Pineapple mealybug wiltassociated viruses in pineapple planting blocks. Plant Disease 94:196-200. http://dx.doi.org/10.1094/PDIS-942-0196 [ Links ]
Received: January 06, 2016; Accepted: March 28, 2016











 text in
text in