INTRODUCTION
Allergic diseases are a broad spectrum of inflammatory hypersensitive reactions, and their prevalence has significantly risen, representing one of the most important challenges worldwide today. Aspergillus fumigatus is a ubiquitous fungus, can cause from invasive disease to allergic disease allergic bronchopulmonary aspergillosis (ABPA),1 its allergens are important sensitizers and trigger symptoms in allergic individuals with other allergic conditions.2–4 For example, this fungi have been frequently found in nasal and skin microbiota of allergic patients5 and is an important inducer of allergic and auto reactive responses in patients suffering atopic dermatitis.6,7 Fungal sensitization in allergic subjects with respiratory diseases range from 2.3% to even 80%, according to Mirabi et al,8 in particular for A. fumigatus various allergens have been characterized, such as manganese superoxide dismutase (MnSOD),1 thioredoxin9 and Cyclophilin B,10 making the study of its allergens of pivotal importance to understand fungal allergy and the designing of specific allergen recombinants for immunotherapy and component resolved diagnosis. In the tropics mite’s sensitization is the main risk factor for asthma and rhinitis and co-exposure with Aspergillus is common, both are colonizers of human skin and abundant in the tropical environment, thus potential of cross sensitization and cross reactivity for these two allergenic sources needs to be explored.
Cross-reactivity among allergenic sources is an important immunological mechanism involved in cross sensitization and exacerbation of allergic responses.11 This is a phenomenon described for allergy to foods and mites.11,12 For A. fumigatus, cross-reactivity with other fungus such as: Malassezia simpodialis, Penicillium notatum, Malassezia furfur, Alternaria alternata and Cladosporium herbarum, have been studied.13 For these fungus species, allergens related to MnSOD, cyclophilins, thioredoxins, serin proteases, enolases, P1 and P2 ribosomal proteins, heat shock protein and peroxisomal proteins are involved in cross reactivity,13 being these allergens highly conserved, which is of relevance when studying cross-reactivity in the related A. fumigatus specie. Nowadays, eighty fungus genomes are available,14 constituting a powerful tool to study cross reactivity using in silico approaches. However, cross-reactivity among fungus and other allergenic sources has been poorly explored. Here, we examined potential cross reactivity among A. fumigatus and mites, both important allergenic sources in tropical regions, using mainly a bioinformatic approach.15
METHODS
Selection of allergens and homologous search
Aminoacid sequence from allergens derived of Aspergillus fumigatus and Dermatophagoides pteronyssinus were retrieved from database Allergome.16 Allergens used for this study are listed in Tables 1 and 2. Aminoacid sequences from A. fumigatus were used as input in BLASP (https://blast.ncbi.nlm.nih.gov/) to search similar sequences in mite’s proteome using the term and taxid “house-dust mites (taxid:6952)”. Aminoacid sequences with similarity upper 25% were selected for furthers analysis. A similar approach to extend analysis of cross reactivity, but in this situation, aminoacid sequences from allergen characterized in allergenic source D. pteronyssinus was used as input to perform BLASTP against A. fumigatus, using term and taxid “Aspergillus fumigatus (taxid:746128)”. Upon sequences were selected, binary alignment was performed and identity level was determined with IBIVU PRALINE tool.17
Table 1. Allergens characterized from A. fumigatus with homology and identity shared with D. pteronyssinus.
| Allergen N° | Alergenos de Aspergillus Fumigatus | Uniprot acceso | PDB template | Homologous | Access code | Identity with D. pteronyssunis |
|---|---|---|---|---|---|---|
| 1 | Ribosomal protein L3 | Q8NKF4 | 6y2l.2.A | 60S ribosomal protein L3-like | XP_027199289.1 | 69 |
| 2 | Molecular chaperone Mod-E/Hsp90 | B0Y324 | 4z1f.1.A | Heat shock protein 83-like | XP_027197940.1 | 66 |
| 3 | Enolase | Q96X30 | 4g7f.1.A | Enolase-like | XP_027197319.1 | 66 |
| 4 | Acidic ribosomal protein P2 | Q9UUZ6 | 4v6i.72.A | Ribosomal protein P2-like protein | AUX14772.1 | 65 |
| 5 | peptidyl-propyl cis-trans isomerase | Q9Y7F6 | 2cfe.1.A | Cis-trans isomerase F (Der f 29) | XP_027204080.1 | 64 |
| 6 | Superoxide dismutase Mn | Q92450 | 2cdy.1.A | Superoxide dismutase [Mn] | XP_027200320.1 | 52 |
| 7 | Thioredoxin Asp | Q1RQJ1 | 2ypm.1.A | Thioredoxin-like protein 1 | XP_027200765.1 | 43 |
| 8 | Aspergillopepsin-1 | P41748 | 1ibq.1.A | Lysosomal aspartic protease-like | XP_027204642.1 | 33 |
| 9 | Alkaline protease 1 | P28296 | 3f7m.1.A | Membrane-bound transcription factor site-1 protease-like | XP_027194529.1 | 33 |
| 10 | Alkaline protease 2 | P87184 | 3f7o.1.A | Membrane-bound transcription factor site-1 protease-like | XP_027194529.1 | 29 |
| 11 | Major allergen Asp f 2 | P79017 | 1eb6.1.A | Uncharacterized protein LOC113788378 | XP_027193641.1 | 28 |
| 12 | Peroxiredoxin Asp f3 | O43099 | 1h4o.8.A | Peroxiredoxin 1-like | XP_027200335.1 | 28 |
| 13 | Allergen Asp f 7 | O42799 | 5ntb.1.A | Sister chromatid cohesion protein PDS5 homolog B-like | XP_027205714.1 | 24 |
| 14 | Ribonuclease mitogillin | P67875 | 1jbr.1.C | N/A | ||
| 15 | Allergen Asp f 15 | O60022 | 3m3g.1.A | N/A | ||
| 16 | Probable glycosidase crf1 | Q8J0P4 | 6ibu.1.A | N/A | ||
| 17 | Cell wall protein phiA | A4FSH5 | N/A | |||
| 18 | Allergen Asp f 4 | O60024 | 3qis.1.A | N/A | ||
| 19 | Extracellular metalloproteinase mep | P46075 | 4m65.1.A | N/A | ||
| 20 | IgE-binding protein | O60025 | 5csd.3.A | N/A |
Table 2. Allergens characterized D. pteronyssinus with homology and identity shared with A. fumigatus.
| Mites | Allergen | Uniprot code | Identidad | A. fumigatus | Homologous |
|---|---|---|---|---|---|
| Der p 28 | Heat Shock Proteins | A0A291KZD8 | 77.89% | XP_750490.1 | molecular chaperone Hsp70 |
| Der f 26 | Myosins | A0A291KYZ8 | 33.54% | XP_751821.2 | Calmodulina |
| Der p 25 | Triosephosphate isomerase | A0A291KYZ7 | 54.81% | XP_753309.1 | triosephosphate isomerase |
| Der p 24 | Ubiquinol-Cytochrome C Reductase Binding Protein | A0A0K2GUJ4 | 29.89% | XP_752147.2 | ubiquinol-cytochrome c reductase complex 14 kDa |
| Der p 18 | Chitinases | Q4JK71 | 27.03% | EDP48562.1 | class V chitinase |
Modeling 3D models of allergens from A. fumigatus
All allergens without experimental structures resolved and reported in protein data bank were modeled using Swiss-model server. For this, aminoacid sequences for each allergen without PDB reported were used as input in server for modeling. Templates were selected based on highest identity level share with aminoacid sequence used for it, and Angstrom value. Quality model was assessed by Prosa web.18 Visualization of models was performed with Pymol software.19
Epitope prediction
B cell epitope prediction was done using Ellipro v 3.0 server. 3D allergen models were used as input. Minimum score and maximum distance (Angstrom) were set to 0.5 and 6, respectively. Epitopes with high conserved rates were visualized on 3D model.
Evolutionary analysis of Aspergillus allergen
Evolutionary or conservation analysis of each aminoacid for each allergen from A. fumigatus among invertebrates was estimated by using Consurf server.20
RESULTS
Allergens selected
In total, twenty aminoacid sequences reported in allergome database were retrieved for analysis (Table 1). Thirteen proteins shared conservation and identities with some sequence in proteome from D. pteronyssinus. In this, we found that Ribosomal protein L3, from A. fumigatus exhibited the highest identity level (69%) with a homologous from D. pteronyssinus (60S ribosomal protein L3-like).
Modelling allergen
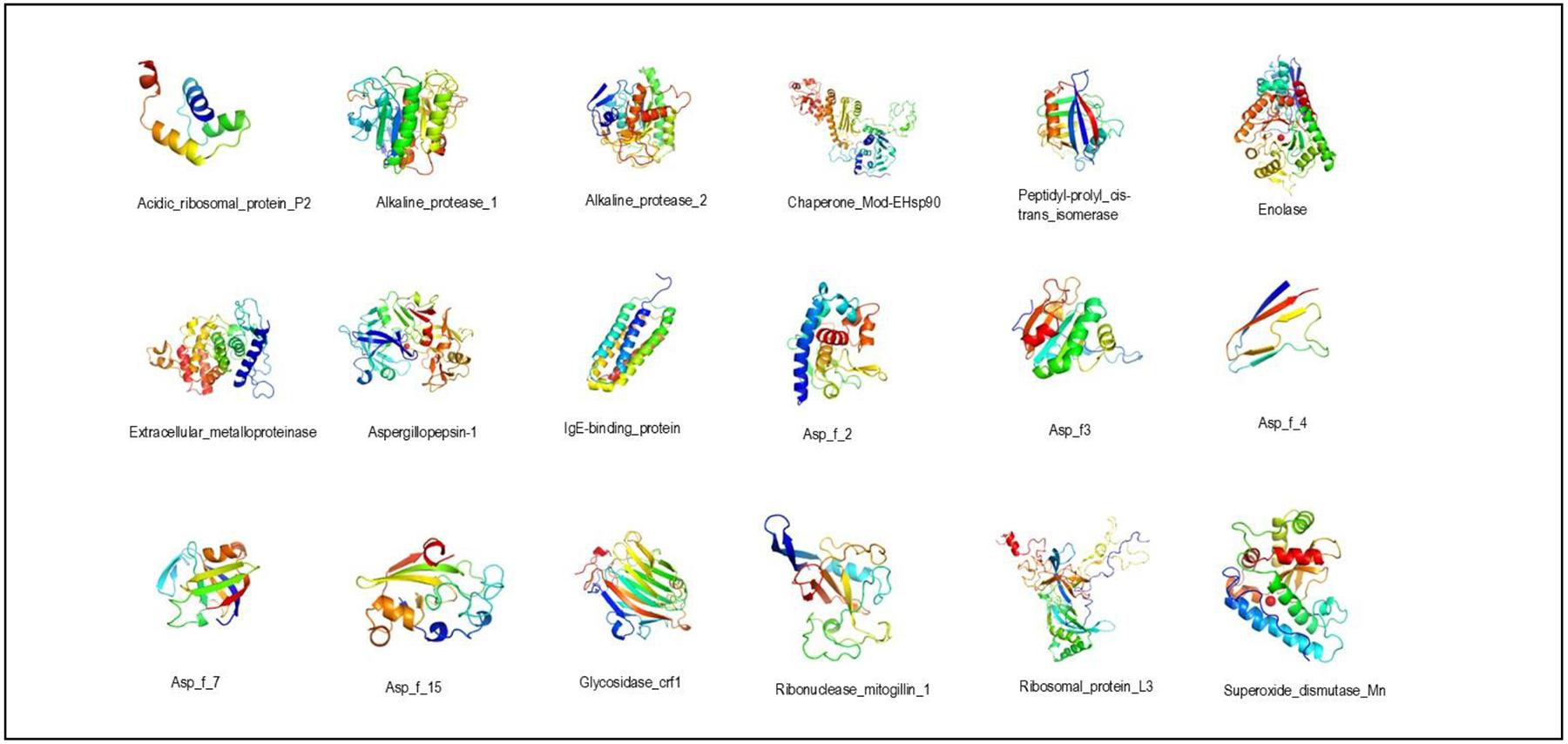
Allergens from A. fumigatus were modeled to predict B cell epitopes. All models showed typical folding related to the family protein they belonged. Seven allergens didn’t exhibit any identity with proteome from D. pteronyssinus but were modeled to further comparative analysis (Table 1 and Figure 1).
Conservation of Epitopes among A. fumigatus and D. pteronyssinus
To determine cross-reactivity, B cell epitope prediction was performed. We informed only epitopes conserved between allergens from A. fumigatus and D. pteronyssinus. Of thirteen allergens modeled from Aspergillus, seven were used for cross-reactivity exploration, these were selected based on identity level shared with homologous in mite’s proteome (> 40%). Allergens used were: Ribosomal protein L3, Molecular chaperone Mod-E/Hsp90, Enolase, Acidic ribosomal protein P2, peptidyl-propyl cis-trans isomerase, Superoxide dismutase Mn and Thioredoxin (Table 1 and Figure 2).
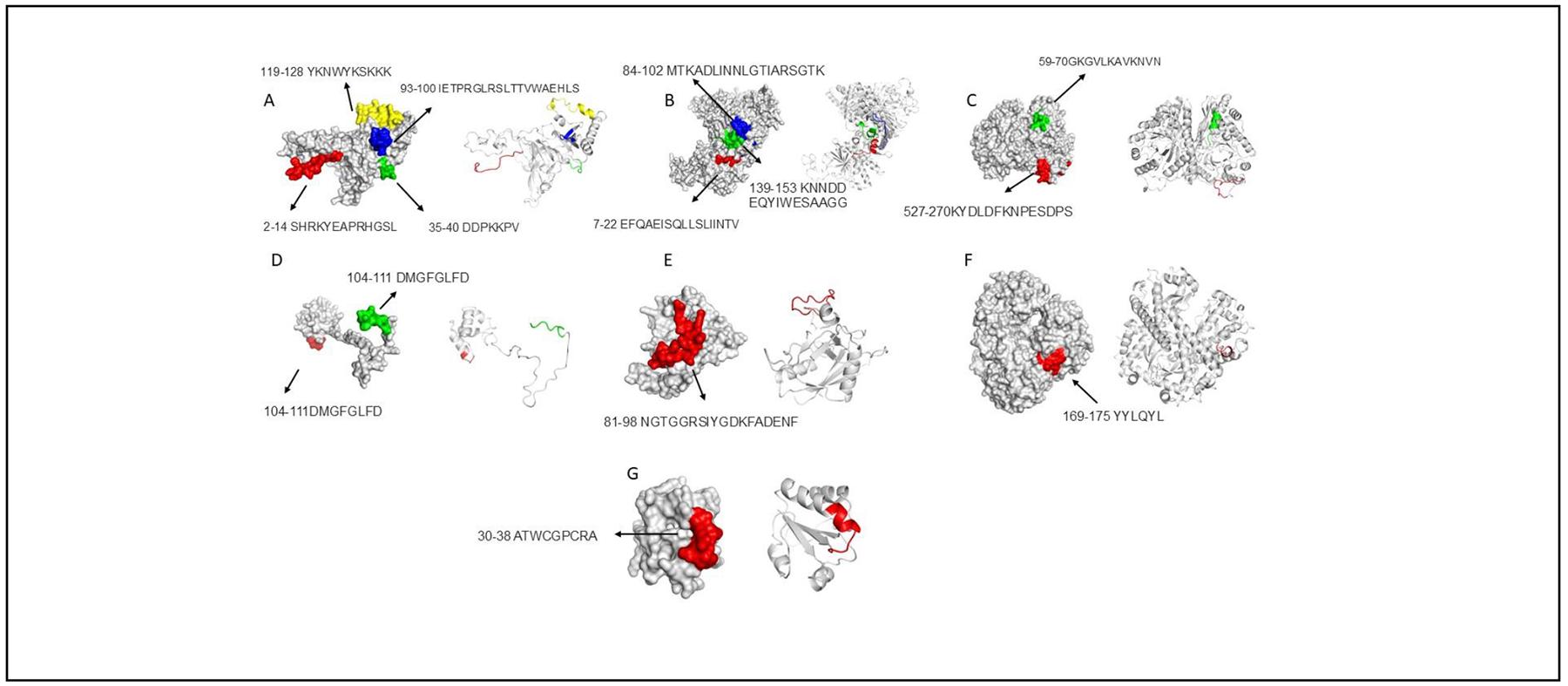
Figure 2. Surface and cartoons models showing predicted epitopes between A. fumigatus and D. pteronyssinus. A: Ribosomal protein L3, B: HSP90, C: Enolase, D: Acidic ribosomal protein P2, E: peptidyl-propyl cis-trans isomerase, F: MnSOD and G: Thioredoxins.
Ribosomal protein L3 showed 69% in identity level with a homologous in D. pteronyssinus identified as 60S ribosomal protein L3-like. According to epitope prediction, these proteins shared four conserved epitopes. This was the highest number of predicted epitopes shared for allergens used. Next, Hsp90 was the second allergen with an identity level above 60%, and bioinformatic approaches predicted three epitopes conserved with homologous in D. pteronyssinus. For enolase and Acidic ribosomal protein P2, two epitopes were predicted. Both allergens exhibited 66 and 65%, correspondly, with their Homologous in mites. Finally, for allergens peptidyl-propyl cis-trans isomerase, MnSOD and, thioredoxins, just one epitope was conserved. These allergens shared different identity levels, for peptidyl propyl was 64% with the allergen Der f 29 from D. pteronyssinus, MnSOD is 52% and 43% for the thioredoxin allergen in A. fumigatus.
Conservation of Epitopes among allergens of D. pteronyssinus and homologous in A. fumigatus
We found five allergens of D. pteronyssinus with identity in their aminoacid sequences with homologous in A. fumigatus (Table 2). In Figure 4, two representative allergens (Der f 28 and Der f 25) that shared 77.8 and 54.8% of identity are shown. For first allergen, Der p 28, three conserved epitopes with molecular chaperone Hsp70 of A. fumigatus were predicted. For Der p 25, two epitopes are informed (Figure 4 C-D).
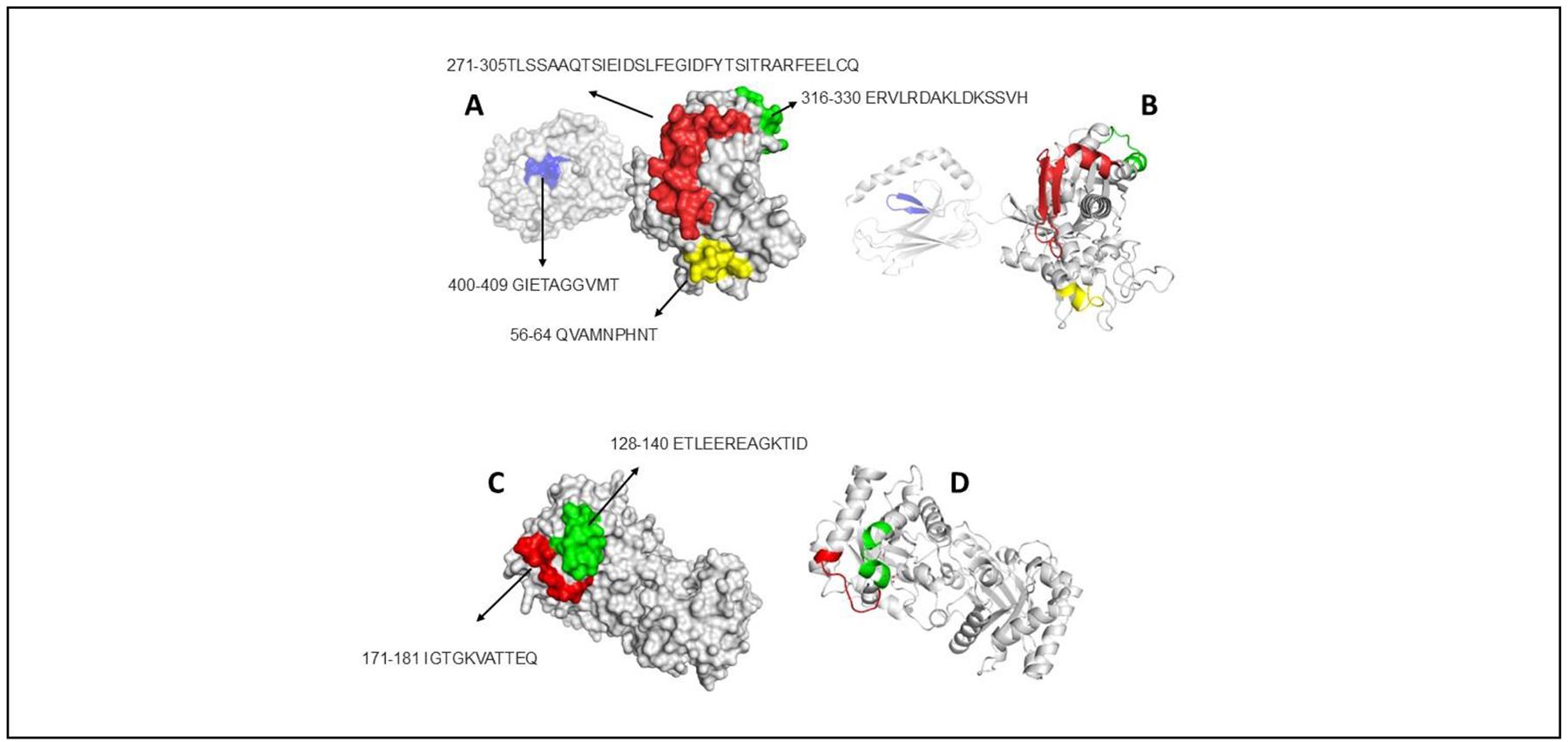
Figure 4. Surface and cartoon models showing conserved epitopes between allergen characterized in D. pteronyssinus and Homologous of A. fumigatus. A: Surface model showing epitopes on Der p 28, B: Cartoon model to show epitopes on model of Der p 28, C and D: Surface and cartoon models of allergen Der p 25.
DISCUSSION
Aspergillus fumigatus has wide distribution worldwide. Due to this, it is difficult to approximate a real prevalence. There is estimated prevalence data depending on the context (asthma, cystic fibrosis, immunocompromised) or some regions of the world. In low- and middle-income countries such as Colombia, there is not enough data to calculate its prevalence. It is highlighted that it is not a fungus that is only in the environment, but also on surfaces and is dependent on changes in climate and environmental regulation mechanisms. In humans, cause significant allergic manifestations in sensitized subjects. Fungi and dust mite species cohabit in household dust, yet the precise dynamics of their interaction remain poorly understood. Existing literature offers limited evidence, indicating a potential interplay between fungal presence and Dermatophagoides pteronyssinus, with suggestions of both synergistic and antagonistic relationships. In this study, twelve allergens from A. fumigatus were found sharing homology with proteins reported in D. pteronyssinus. The 3D models obtained showed typical folding to the protein family they belonged. Ribosomal protein L3, Molecular chaperone Mod-E/Hsp90, Acidic ribosomal protein P2, Enolase, and peptidyl-propyl cis-trans isomerase were the allergens with the highest score identity (>60%). At least four B linear epitopes were predicted to be shared between allergens and homologous in D. pteronyssinus.The immune response to the different allergens currently described is variable which compromises the immune response in sensitized individuals.
Research on fungal allergies faces significant challenges due to the complexity of the elicited immune response, which produces a variety of intracellular metabolites, cell wall components and extracellular secretomes, inducing humoral and cellular immune responses. This diversity makes the clinical interpretation of sensitization test results difficult, since sensitization to total fungi extracts does not always reflect precise sensitization to a specific species, due to the variability in the composition and allergenicity of the final products, resulting of different cultivation and extraction processes. Standardization in the production of reliable fungal extracts is presented as a key challenge to advance in the diagnosis and treatment of fungal allergies. This would involve establishing uniform protocols for the cultivation and extraction of fungi, with the aim of minimizing variability between batches of products.
Respiratory diseases caused by Aspergillus are classified according to the immune mechanism: Allergic aspergillosis (IgE-mediated): allergic bronchopulmonary aspergillosis (ABPA), severe asthma with fungal sensitization (SAFS) and severe allergic bronchopulmonary aspergillosis. Aspergillus allergic sinusitis, and hypersensitivity pneumonitis. Diseases due to saprophytic colonization: they include simple or complex aspergilloma (chronic cavitary pulmonary aspergillosis), chronic pulmonary aspergillosis (CPA), fungal sinus aspergillomas. Invasive disease: invasive pulmonary aspergillosis, which can be acute subacute, called chronic necrotizing pulmonary aspergillosis (CNPA), acute fulminant invasive sinusitis, chronic invasive sinusitis, invasive granulomatous sinusitis.21,22 From the perspective of clinical diagnosis traditionally, in the evaluation of ABPA and Aspergillus sensitive asthma (ASA), derivatives of crude extracts of A. fumigatus are used, presenting limitations in their purity and consistency. The differentiation between ASA and ABPA can be complicated and requires multiple approaches, since both can share symptoms between them: difficult-to-manage asthma, high IgE levels and bronchiectasis, the use of antigens could improve the current diagnosis for ABPA.
A. fumigatus is a well allergenic source characterized, several allergens with molecular and immunological different are reported in IUIS. For example, cyclophilins, L3 ribosomal proteins, thioredoxin and MnSOD are important allergens related to trigger allergic reactions.1,23,24 Cross-reactivity of this allergenic source with other fungi or yeast is well reported in the literature,25 Penicillium sp., Alternaria alteria, and Malassezia simpodialis are referred to be cross reactivity with A. fumigatus.26 In this study we demonstrate that another important challenge in the diagnosis of fungal hypersensitivity is distinguishing between genuine sensitization and cross-reactivity. Evaluation of the sensitization profile to specific allergens and panallergens is crucial, since crude allergenic extracts from different fungal sources can have significant cross-reactivity by structural homology, which can complicate the interpretation of sensitization test results.
This is important to understand the cross sensitization that can suffer the patients, and this can lead to exacerbated symptoms. However, cross reactivity of A. fumigatus with other allergenic source different to fungi or yeast is not reported. This is the first study to determine the potential of A. fumigatus in cross reactivity with other allergenic source, for example, mites. Here, using a bioinformatic approach we have identified thirteen allergens with identity (above 20%) in their amino acid sequences between A. fumigatus and D. pteronyssinus. In this study, we focused on bioinformatic characterization of seven allergens (Ribosomal protein L3, HSP90, Enolase, Acidic ribosomal protein P2, peptidyl-propyl cis-trans isomerase, MnSOD and Thioredoxins). Epitope prediction suggests that several antigenic regions would be involved in cross reactivity between allergens from A. fumigatus and their homologous in D. pteronyssinus.
Thioredoxin [Trx] has antioxidant and protein regulation functions of the photosynthetic mechanism of the chloroplast, such as NADP-malate dehydrogenase [NADP-MDH] are identified in humans and in allergenic sources such as Aspergillus spp where the allergens known as ASPID F28, ASPID F29 characterized as thioredoxins, reported in other allergenic sources such as Malassezia sympodialis and Coprinus comatus, also observing cross-reactivity between the human Trx enzyme and those present in Aspergillus spp, Malassezia sympodialis and Coprinus comatus, due to molecular mimicry between the thioredoxin of the allergenic sources and human, presenting cross-reactivity mediated by IgE, thus generating autoreactive responses that exacerbate the symptoms of allergic diseases. Therefore, it is necessary to evaluate this CR in the clinic and if it presents with D. pteronyssinus.
Manganese super oxide dismutase [MnSOD], another cross-reactive allergen, are important enzymes for the physiological response to oxygen toxicity, dismutating toxic superoxide free radicals into oxygen and hydrogen peroxide, oxidative stress and lung parenchymal damage. MnSOD is involved in the IgE-mediated autoreactive immune response. Structural homology has been found with human MnSOD, of Af especially the allergen ASPID F 6, Drosophila melanogaster and Saccharomyces cerevisae, which in previously sensitized individuals favors the crossed and autoreactive immune response due to structural homology.
Is important to define that homologous identified in mites, just Cis-trans isomerase is reported as allergen in D. farinae but not in D. pteronyssinus, registered as Der f 29.27 However, homology and epitopes predicted indicated that homologous in D. pteronyssinus need to be explored in future to determine allergenic potential.
Mites are an important allergenic source in the tropics, and species such as Blomia tropicalis, D. farinae and D. pteronyssinus have been implicated in allergic sensitization.15,28 Cockroach, Anisakis simplex and shrimp are the sources implicated in cross reactivity with mites,29,30 and tropomyosin is considered the major allergen in cross reactivity among invertebrates.31 In Table 2, we reported identity between five allergens from D. pteronyssinus and homologous in A. fumigatus. This increases the repertoire of allergens involved in cross reactivity between these sources.
Methodology used here was based on bioinformatic, we considered this was adequate for initial explorations to determine cross reactivity between the allergenic sources studied. We know that results obtained in our study need to be validated by experimentation. However, bioinformatic tools used in were robust and previously validated for allergen characterization and study.32,33











 nueva página del texto (beta)
nueva página del texto (beta)