Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Accesos
Accesos
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO
Compartir
Veterinaria México OA
versión On-line ISSN 2448-6760
Veterinaria México OA vol.10 Ciudad de México 2023 Epub 22-Oct-2024
https://doi.org/10.22201/fmvz.24486760e.2023.1215
Original Research Articles
Strong antibiotic resistance profiles in Salmonella spp. isolated from ground beef in Central Mexico
1 Universidad Nacional Autónoma de México, Facultad de Medicina Veterinaria y Zootecnia, Ciudad de México, México.
2 Centro Nacional de Referencia de Inocuidad y Bioseguridad Agroalimentaria, Estado de México, México.
3 Instituto Nacional de Salud Pública, Dirección de Enfermedades Crónicas, Centro de Investigación en Salud Poblacional, Cuernavaca, Morelos, México.
We determined the prevalence of Salmonella enterica (SE) in retail ground beef sold across eight state capital cities from Central Mexico (n = 115) as well as the antimicrobial resistance (AMR) phenotype and genotype of the isolates obtained. SE was detected in 48/115 samples, with variable prevalence (10-80 %) across geographical regions (χ2 = 24.2, P = 0.0021). We collected 116 isolates and observed circulation of serovars implicated in human salmonellosis in Mexico (Agona, Anatum, Infantis, Newport, Derby, Give y Typhimurium). Resistance was more frequently observed for tetracycline (39.7 %), chloramphenicol (37.9 %), streptomycin (37.1 %), trimeto-prim-sulfamethoxazole (31.0 %), and ampicillin (28.4 %). Resistance against azithromycin was moderate (14.7 %), few isolates resisted cephalosporins (2.6-3.4 %), all were susceptible to carbapenems, and 38.8 % of the isolates were multidrug resistant (MDR). The sequenced genomes carried AMR alleles against aminoglycosides (aadA, aac, aph), beta-lactams (bla-CARB, bla-PSE, bla-TEM, bla-CTX-M, bla-CMY), phenicols (floR), folate pathway inhibitors (sul, dfrA), fluoroquinolones (qnrAB, oqxAB), tetracyclines (tetABM), and macrolides: mph(A) y lnu(F). In conclusion, the beef under study works as a reservoir of SE with worrisome MDR phenotypes. The pathogen has acquired AMR genes against antibiotics used in human and veterinary medicine. The emergence of resistance to azithromycin is particularly alarming and has not been reported to date. Further studies are needed to better characterize AMR in SE populations associated with cattle.
Keywords: Salmonella spp.; Ground beef; Antibiotics; Phenotype; Genotype; Multi-drug resistance
Se determinó la frecuencia de contaminación con Salmonella enterica (SE) en la carne molida de res que se comercializa en ocho capitales estatales del centro de México (n = 115), así como el fenotipo y genotipo de resistencia a antimicrobianos (RAM) de los aislamientos obtenidos. Se detectó SE en 48/115 muestras, con tasas variables de contaminación (10-80 %) por zona geográfica (χ2 = 24.2, P = 0.0021). Se obtuvieron 116 aislamientos y se observó circulación de serotipos implicados en salmonelosis humana en México (Agona, Anatum, Infantis, Newport, Derby, Give y Typhimurium). La resistencia fue más frecuente en tetraciclina (39.7 %), cloranfenicol (37.9 %), estreptomicina (37.1 %), trimetoprima-sulfametoxazol (31.0 %) y ampicilina (28.4 %), moderada en azitromicina (14.7 %), escasa en cefalos-porinas (2.6-3.4 %) y nula en carbapenémicos, mientras que 38.8 % de los aislamientos fue multi-drogo resistente (MDR). Los genomas secuenciados portaban alelos de resistencia contra aminoglucósidos (aadA, aac, aph), beta-lactámicos (bla-CARB, bla-PSE, bla-TEM, bla-CTX-M, bla-CMY), fenicoles (floR), inhibidores de la vía del folato (sul, dfrA), fluoroquinolonas (qnrAB, oqxAB), tetraciclinas (tetABM) y macrólidos: mph(A) y lnu(F). Se concluye que la carne bajo estudio es reservorio de SE con preocupantes fenotipos de MDR y que el patógeno ha adquirido genes de RAM contra antibióticos usados en medicina humana y veterinaria. El auge de resistencia contra azitromicina es particularmente alarmante y no se había reportado hasta el momento. Se requieren estudios más amplios para una mejor caracterización de la RAM en las poblaciones de SE asociadas con bovinos.
Palabras clave: Salmonella spp.; Carne de res molida; Antibióticos; Fenotipo; Genotipo; Multirresistencia
Study contribution
This research documents the role of ground beef as a relevant reservoir of Salmonella enterica, one of the major foodborne pathogens worldwide. We demonstrated that this bacterium has acquired a wide repertoire of genes that allow it to resist multiple antibiotics of critical importance in human medicine, and that are also used in veterinary medicine. These findings confirm the need to strengthen food-borne pathogen surveillance by considering their resistance profiles as an additional risk factor for food safety and public health.
Introduction
In its 2015 report on foodborne diseases, the World Health Organization (WHO) reported that non-typhoidal salmonellosis occurs at an annual rate of 150 million cases and causes 60 thousand deaths.1 In Mexico, the incidence of the disease in the last decade (> 60 cases/100 thousand inhabitants)2 is at least three times higher than that of developed countries, highlighting its epidemiological importance in our country.
It has been established that meat from different species, including beef, is a relevant reservoir of SE, the etiological agent of salmonellosis. Studies conducted in Mexico report moderate to high (15 a > 50 %) prevalence of this pathogen in retail beef across different states.3
The latter findings are more relevant in the context of the uprising AMR observed in SE strains, with increasing proportions of MDR phenotypes (26-70 %), both in Mexico and other countries.4-7 This evidence suggests that at least part of the salmonellosis outbreaks linked to ground beef could involve MDR strains. Such reasoning is plausible considering that it occurs in developed countries with far lower SE ground beef contamination rates compared to those observed in Mexico.8
Despite the importance of salmonellosis, and particularly that caused by MDR strains, few studies deal with AMR in SE associated with beef. Most of them are local surveys,9-11 while very few use an integral approach involving both AMR phenotypes and genotypes,4,12 which may contribute to identify the factors involved in the emergence and dissemination of AMR, as well as to outline containment measures.
In this research, we conducted a regional survey to determine SE prevalence in retail ground beef sold in the capital cities of eight states in Central Mexico. Moreover, we characterized the AMR profile of the obtained isolates by determining both AMR phenotypes and genotypes. This approach allowed identifying the antimicrobial classes that are losing effectiveness against this pathogen as well as the genetic factors sustaining these phenotypes and their potential dissemination within SE populations circulating in ground beef.
Materials and methods
Ethical statement
This research did not involve animals. Hence, it did not require the approval of the Institutional Subcommittee for the Care and Use of Experimental Animals (SICUAE) of the Faculty of Veterinary Medicine, National Autonomous University of Mexico. However, it was reviewed and approved by the Research Ethics Committee of the National Institute of Public Health, Registration No. 17CEI00420160708, FWA:00015605.
Study design and sampling
In this project, we analyzed a subsample of retail stores (n = 115) considered in a wider survey. This wider survey consists of a cross-sectional epidemiological study, with multi-level analysis. The unit of grouping was the capital citiy of each state and the sample units were the ground beef retail stores (meat stores, street vendors, wet markets, supermarkets, n = 400) from the capital cities of all Mexican states during the 2021-2023 period.
The global sample size cited above (400) was determined with the formula to estimate a proportion in a population with a specified absolute precision.13 Based on previous studies, SE prevalence was estimated at 30 %3 and precision was set to 5 %. In this way, the estimated global sample size was 384, which was rounded to 400 to compensate for possible sample losses. The number of samples per city was proportional to the number of retail stores in each city: the higher the number of retail stores in a city, the larger the sample size.
Initially, we planned to conduct a random sampling within each city based on the retail stores registered in the National Statistical Inventory of Economic Units (DNUE)14 with an active location in Google Maps (https://www.google.com/maps). However, many retail stores closed down during the COVID-19 pandemic. Therefore, it was necessary to conduct a convenience sampling instead.
The sole inclusion criterium required that the store be open to the public when we conducted the sampling. Otherwise, we took the sample from the nearest open store. To increase the probability of collecting isolates, we decided to take at least 10 samples in cities where the estimated sample size was below 10.
Considering the elements described above, we analyzed 115 samples in this research (Table 1), corresponding to eight capital cities from Central Mexico (Mexico City, Cuernavaca, Tlaxcala, Querétaro, Pachuca, Toluca, Chilpancingo, and Puebla).
Table 1 Distribution of the 115 samples across capital cities and states participating in the survey
| City | State | Meat retail stores | Sample size |
|---|---|---|---|
| Ciudad de México | Ciudad de México | 6 420 | 25 |
| Toluca | Estado de México | 9 548 | 20 |
| Pachuca | Hidalgo | 1 514 | 10 |
| Cuernavaca | Morelos | 1 536 | 10 |
| Puebla | Puebla | 4 153 | 20 |
| Querétaro | Querétaro | 997 | 10 |
| Chilpancingo | Guerrero | 2 016 | 10 |
| Tlaxcala | Tlaxcala | 1 097 | 10 |
Source: National Statistical Inventory of Economic Units.14
In each retail store, we bought a 250-g ground beef package, which was placed in a previously identified and sterile plastic bag. The samples were transported to the laboratory in insulated containers and kept in refrigeration (approximately 4 ºC) with the aid of cooling gel pads. Sample analysis was conducted not later than 24 h post collection.
Salmonella spp. detection, isolation, and confirmation
Samples were analyzed following the methods described in Mexican regulation NOM-210-SSA-2014.15 The analysis include non-selective pre-enrichment in buffered peptone water, selective enrichment in Rappaport-Vassiliadis and tetrationate broths, as well as selective agar plating on Hektoen enteric, xylose-lysine-desoxycholate, and bismuth sulfite media.
Afterwards, isolates with typical Salmonella spp. morphology were biochemically confirmed: triple sugar iron agar, lysine iron agar, and urea tests. Salmonella identity was also confirmed through polymerase chain reaction (end point PCR) by amplifying a fragment of the invA gene (500 bp), which is constitutive of this genus.16 The full description of all these procedures is available from the open access platform protocols.io.17
Antibiotic susceptibility testing (AST)
The AMR phenotype of isolates was determined by the Kirby-Bauer disk diffusion technique.18 We used a panel of 12 antibiotics included in the WHO list of critically important antimicrobials for human medicine.19 This panel included antibiotics that are intensively used in human or veterinary medicine as well as drugs that are used to treat invasive salmonellosis or serious infections caused by enterobacteria (Table 2).
Table 2 Antibiotic panel used in the AST tests and criteria used to determine the resistance phenotype of isolates
| Antibiotics | Concentration (μg) | Phenotype according to the inhibition zone diameter (mm) 1 | |
|---|---|---|---|
| I | R | ||
| Penicillins | |||
| Ampicillin (AMP) | 10 | 14-16 | ≤13 |
| Amoxicillin/clavulanic acid (AMC) | 20/10 | 14-17 | ≤13 |
| Third-generation cephalosporins | |||
| Ceftriaxone (CRO) | 30 | 20-22 | ≤19 |
| Fourth-generation cephalosporins | |||
| Cefepime (FEP) | 30 | 19-24 | ≤18 |
| Carbapenems | |||
| Meropenem (MEM) | 10 | 20-22 | ≤19 |
| Aminoglycosides | |||
| Amikacin (AMK) | 30 | 15-16 | ≤14 |
| Streptomycin (STR) | 10 | 13-14 | ≤12 |
| Fluoroquinolones | |||
| Ciprofloxacin (CIP) | 5 | 21-30 | ≤20 |
| Macrolides | |||
| Azithromycin (AZM) | 15 | - | ≤12 |
| Tetracyclines | |||
| Tetracyclin (TET) | 30 | 12-14 | ≤11 |
| Phenicols | |||
| Chloramphenicol (CHL) | 30 | 11-17 | ≤10 |
| Folate pathway inhibitors | |||
| Trimetoprim-sulfamethoxazole (SXT) | 1.25/23.75 | 11-15 | ≤10 |
1 Criteria used to classify isolates as clinically resistant (R) or intermediate (I).20
Results were interpreted following the criteria (clinical cutoff values) of the Clinical and Laboratory Standards Institute (CLSI).20 As quality control organisms, we used two strains: an Escherichia coli ATCC 8739 (as an alternative to E. coli ATCC 25922, which stopped yielding results within the specified intervals of CLSI’s M100 supplement) and a Pseudomonas aeruginosa ATCC 27853 strain. Isolates that resisted three or more antibiotic classes were classified as MDR.21 The full description of AST procedures is available from the open access platform protocols.io.22
Whole genome sequencing and serovar prediction
Genomic DNA (gDNA) was extracted with Qiagen’s DNeasy Blood & Tissue Kit as per the manufacturer’s instructions. The gDNA was quantified using a Qubit 3.0 fluorometer. The DNA libraries were prepared from 1 ng of gDNA using the Nextera XT kit v.2 (Illumina) and sequenced in an Illumina MiSeq system (paired-end mode and 2 × 250 bp). In silico serovar prediction was performed by analyzing assembled genomes with the Salmonella In Silico Typing Resource (SISTR) program, version 1.1.2.23
Quality control and trimming of raw reads
The initial quality of raw reads was assessed with the FastQC program.24 Next, we used Trimmomatic to remove Illumina adapters and reads with poor quality scores.25 Trimmed sequences were analyzed again with FastQC to ensure that only high-quality reads (Q>30) were used for bioinformatic analyses.
Genome assembly
Genome assembly was performed at the Bacterial and Viral Bioinformatics Resource Center26 using the de novo algorithm in the SPAdes program, version 3.131.27 The assemblies were evaluated with QUAST program (Quality Assessment Tool for Genome Assemblies) version 5.02,28 to ensure they met the minimal requirements to be used in food safety, public health, and epidemiology applications: not more than 300 contigs, the lowest possible fragmentation index (L50), depth of coverage ≥ 30x, and a percentage of guanine + cytosine (GC) and genome size within the typical range of Salmonella enterica (48 ≤ GC ≤ 56 and 4.5-5.1 Mbp, respectively).29 Assembly quality data are provided as supplementary information (Table S1).
Determination of AMR genotypes
The identification of genomic AMR profiles was determined in silico by running assembled genomes in the AMRFinderPlus program, version 3.8,30 configured to detect both AMR genes and mutations.
Statistical analysis
We determined Salmonella spp. prevalence in ground beef from each capital city as well as the proportion of isolates of each serovar exhibiting susceptible, intermediate, or resistant phenotypes to each antibiotic analyzed. Moreover, we conducted chi-square tests and odds ratio calculations to assess if there was an association between SE prevalence and resistance phenotypes with the city where isolates originated and the serovar. We also performed a Pearson correlation analysis to determine whether AMR phenotypes and genotypes were correlated. These analyses were conducted with IBM SPSS Statistics 25.
Results
Ground beef as a reservoir of Salmonella spp.
The overall prevalence was 41.7 % (48/115) and we collected 116 Salmonella spp. isolates. Ground beef contamination rates were moderate to high (30-80 %) in most cities (Figure 1), except Tlaxcala and Querétaro (10 %).
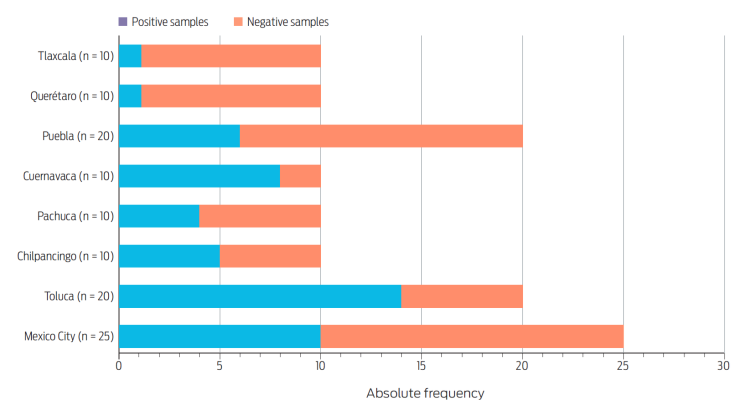
Figure 1 Salmonella spp. prevalence in retail ground beef sold in eight state capital cities of Central Mexico (May-December 2021).
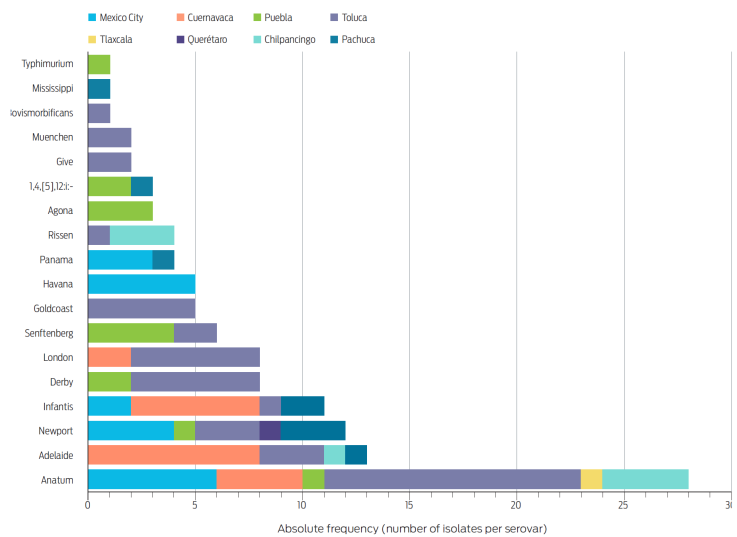
We detected a significant association (χ2 = 24.2, P = 0.0021) between the proportion of contaminated meat and the city, being higher in Cuernavaca and Toluca: odds ratio (OR) 25.5, 95 % confidence interval (95CI) 2.2-299.3, P = 0.0016. Although there was considerable serovar diversity in the sample under study (18 in total), Anatum (n = 28), Adelaide (n = 13), Newport (n = 12), and Infantis (n = 11) collectively represented 55 % of the isolates and were also the most widely distributed (Figure 2).
Strong AMR profiles in Salmonella spp. associated with ground beef
We observed phenotypic resistance to ten of the 12 antibiotics analyzed. The most frequent resistance phenotypes involved tetracycline (39.7 %), chloramphenicol (37.9 %), streptomycin (37.1 %), trimetoprim-sulfamethoxazole (31.0 %), and ampicillin (28.4 %) (Figure 3). Moreover, nearly 15 % of the isolates resisted azythromycin, one of the few alternatives to treat invasive salmonellosis.31
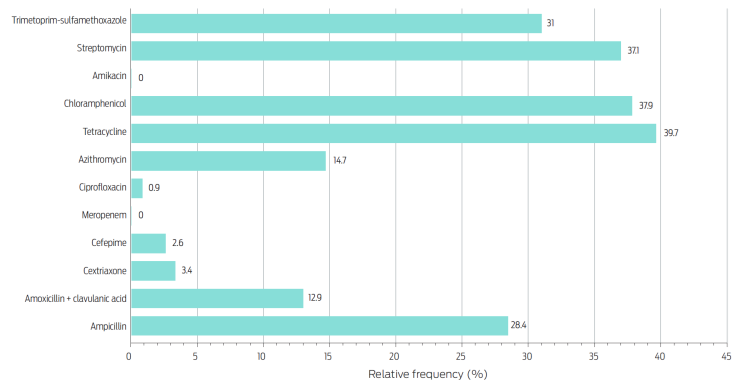
Figure 3 Relative frequency of Salmonella spp. isolates showing resistance to each antibiotic (n = 116).
Fortunately, there was a low proportion of isolates (< 4 %) that resisted antibiotics included in the WHO list as critically important and highest priority,19 such as third- and fourth-generation cephalosporins (ceftriaxone and cefepime) and ciprofloxacin. Likewise, all isolates were susceptible to meropenem and amikacin. The distribution of the observed phenotypes was nearly binary since 54.3 % of isolates were pan-susceptible and 38.8 % were MDR (Figure 4). Furthermore, we found a strong association between serovar and resistance profile (χ2 = 38.9, P < 0.0001), with a higher proportion of MDR phenotypes in isolates of serovars Newport, Senftenberg, Agona, 1,4,[5],12:i-, and Typhimurium compared with the others: OR = 69.7, 95CI 8.9-547.5, P < 0.0001.
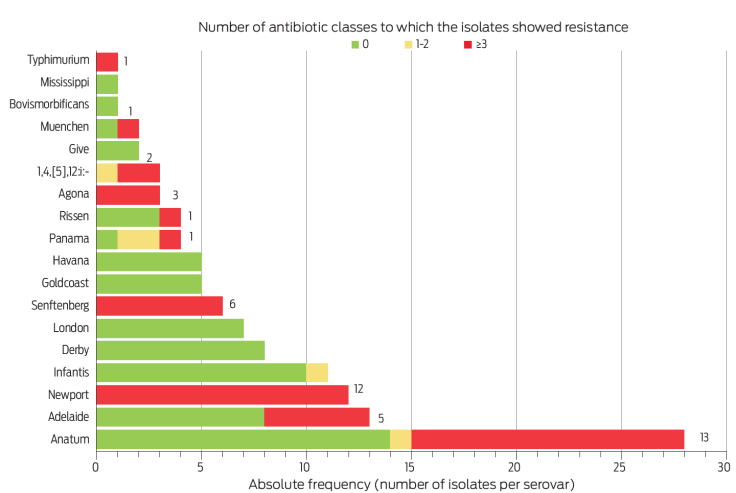
Figure 4 Number of isolates of each serovar that were pan-susceptible (0), mono- o bi-resistant (1-2), or MDR (≥ 3).21 Numbers next to the red bars indicate the absolute frequency of MDR phenotypes.
Genetic determinants associated with AMR phenotypes
Resistance to streptomycin was strongly associated (χ2 = 87.8, P < 0.0001) with the presence of allelic variants encoding aminoglycoside-modifying enzymes, such as adenilil-transferases (aadA1, aadA2, aadA5), acetil-transferases [aac(3)-Iva, aac(3)-IId] and phospho-transferases [aph(3’)-Ia, aph(3”)-Ib, aph(6)-Id y aph(4)-Ia] (Figure 5).
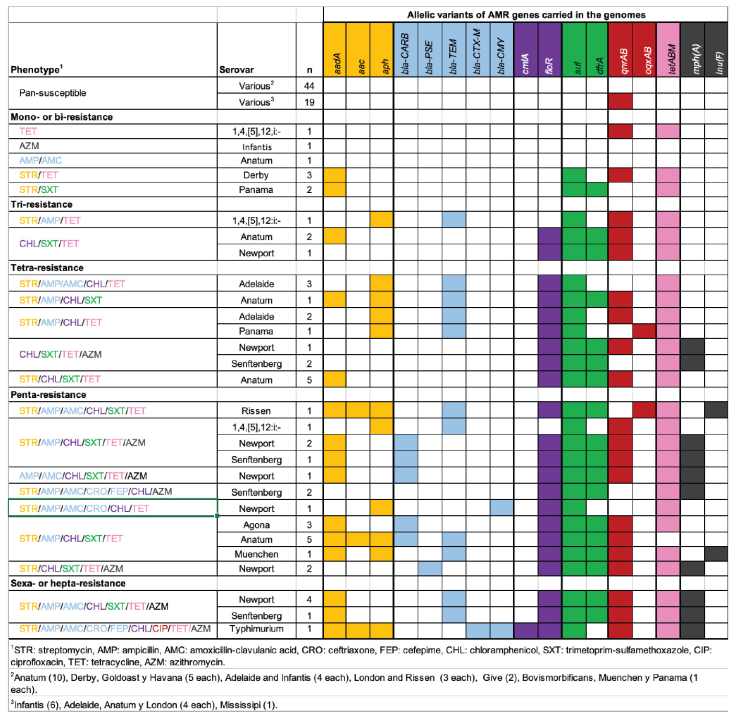
Figure 5 AMR phenotypic and genotypic profiles of isolates under study. Antibiotic classes and AMR genes are color coded. Within AMR genotypes, cells filled with the corresponding antibiotic class color indicate that the gene is present, while blank cells denote its absence.
In betalactams, resistance occurred almost exclusively in isolates carrying genes that encode class-A (bla-CARB, bla-TEM, bla-CTX-M, bla-PSE) or class-C (bla-CMY) betalactamases (χ2 = 97.0, P < 0.0001). As a remarkable exception, two isolates of serovar Senftenberg resisted all tested betalactams except carbapenems and did not carry betalactamase-encoding genes in their genomes. The same occurred in one isolate of serovar Anatum that resisted penicillins (Figure 5).
Phenicol resistance was mainly mediated by the active efflux mechanism encoded by the floR gene, which was present in 100 % of chloramphenicol-resistant isolates (Figure 5). Only one isolate of serovar Typhimurium carried a different phenicol resistance gene (cmlA1), but it also encodes an active efflux mechanism. Regarding folate pathway inhibitors, the resistance phenotypes were associated (χ2 = 64.7, P < 0.0001) with the presence of allelic variants encoding enzymes that are resistant to these antibiotics: dihydropteroate synthase (sul1, sul2) and dihydrofolate reductase (dfrA1, dfrA12 y dfrA17).
Interestingly, although fluoroquinolone resistance was rare (it was only observed in one isolate of serovar Typhimurium), over 40 % of the isolates (49/116) exhibited intermediate resistance to ciprofloxacin (supplementary information, Table S2). This phenotype was significantly associated (χ2 = 53.7, P < 0.0001) with the presence of plasmid-mediated quinolone resistance (PMQR) genes. Nearly 90 % of the isolates that showed intermediate resistance to ciprofloxacin carried allelic variants encoding DNA mimicry proteins (qnrA1, qnrB19) that inhibit the effects of fluoroquinolones on their target molecules (DNA gyrase and topoisomerase IV) (Figure 5).32
Tetracycline resistance was associated (χ2 = 100.1, P < 0.0001) with the presence of alleles encoding active efflux factors tet(A) and tet(B), and ribosomal protection tet(M).33 These genes were present in 97.8 % (45/46) of the isolates showing this phenotype. Finally, macrolide resistance was associated (χ2 = 83.1, P < 0.0001) with the presence of alleles encoding the enzymatic modification factors mph(A) and lnu(F) (Figure 5). We identified point mutations in several genes that are associated with resistance to different antibiotic classes (Table 3). However, these mutations were not associated with the resistance phenotypes observed in our study (χ2 = 1.1, P = 0.58). All the same, there was a strong correlation between phenotypic and genotypic resistance (r = 0.98, P < 0.001, Figure 6).
Table 3 Point mutations in genes associated with resistance phenotypes in the study Salmonella spp. isolates
| Gene | Mutations | Relative frequency (%) | Associated resistance phenotype |
|---|---|---|---|
| ramR | M1V, L115I, M83T | 100.0 | Multi-drug resistance35 |
| acrB | M964T | 100.0 | Macrolides36 |
| gyrA | D795E, T661N | 3.7 | Quinolones37 |
| gyrB | T717N, Q624K | 12.8 | Quinolones37 |
| parC | T255S, T57S, S469A, H747P, N395S, A620T, E6A, R365L | 100.0 | Quinolones37 |
| parE | T599I, P231L | 5.5 | Quinolones37 |
| pmrA | T89S, P102L | 22.0 | Colistin38 |
| pmrB | M15T, A111T, GV73SI, I83V | 21.1 | Colistin38 |
| 16s_rrsD | A1296T, A406G, AAG538TTC, C1074T, C523A, T412C, T409G, A508G, A525T, G536A, G540C, T544C, G1453A | 100.0 | Aminoglycosides39 |
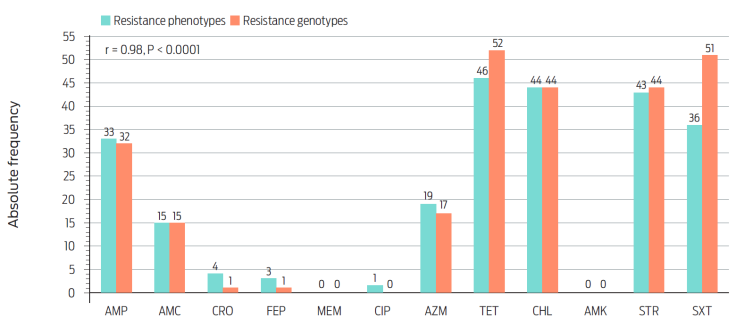
Figure 6 Correlation between AMR phenotypes and genotypes of the sequenced isolates (n = 116). STR: streptomycin, AMP: ampicillin, AMC: amoxicillin-clavulanic acid, CRO: ceftriaxone, FEP: cefepime, CHL: chloramphenicol, SXT: trimetoprim-sulfamethoxazole, CIP: ciprofloxacin, TET: tetracycline, AZM: azithromycin.
It is also important to highlight the high AMR gene diversity (ARGd) observed in this study. The ARGd defines the number of genes encoding different resistance factors (protein families) against the same antibiotic class as well as the total number of AMR genes present at population scale.34 Overall, the isolates carried 15 AMR genes against the seven antibiotic classes under study, and at least two genes encoding different resistance factors per class, except for aminoglycosides, with three (aac, aadA y aph) (Figure 5).
Discussion
In this research, we observed a high prevalence of SE (41.7 %) in retail ground beef sold in the capital cities of eight Central Mexico states. However, this phenomenon did not occur uniformly as there were moderately low contamination rates in Tlaxcala and Querétaro (10 %) and very high in Toluca and Cuernavaca (70 and 80 %, respectively). These findings are in line with those previously reported in other Mexican cities (5-71 %),6,11,40,41 a variability that has been linked to differences in terms of hygienic practices predominating in the formal market (i. e. supermarkets) and informal commercialization channels (eg. street vendors and wet markets).42 Regardless of this variability, our results confirm that in contrast with other nations with a low SE prevalence in ground beef (eg. 0-5 %),43,44 this type of meat works as an important reservoir of this pathogen in Mexico. Moreover, we documented the circulation of SE serovars that are frequently implicated in human salmonellosis (eg. Agona, Anatum, Infantis, Newport, Derby, Give, and Typhimurium)3 and nearly four out of 10 isolates also exhibited strong AMR profiles.
The above-described evidence highlights the risk of human exposure to MDR SE strains through ground beef in the central region of the country. Although food-borne disease attribution is barely performed in Mexico, ground beef has been linked to salmonellosis outbreaks in countries with lower SE prevalence.8,45 Regarding AMR profiles, the antibiotic classes that were less effective against SE were those that have been used in animal production for decades (tetracyclines, phenicols, aminoglycosides and sulfonamides) and are currently registered for use in bovines in Mexico.46
These results are consistent with those from previous studies on SE isolated from bovines4,6,10,47-49 that reported resistance rates of up to 90 % against the antibiotic classes referred to before. They also show that the pathogen has massively acquired AMR genes against these drugs, a phenomenon that has been documented in developed countries as well.50 Many of the mentioned AMR genes are usually carried in mobile genetic elements such as plasmids33 and transposons.51 This set up allows horizontal gene transfer by conjugation and favors its dissemination both within SE populations as well as to other pathogens associated with cattle such as Brucella spp., for which there are few therapeutic alternatives, including tetracyclines.52
Another relevant factor to consider is the risk of AMR dissemination associated with the use of the same antibiotic classes in animals and humans. In this regard, an illustrative example is that of phenicols. Across the world, chloramphenicol has been exclusively used in human medicine for decades. However, florfenicol is registered for veterinary use.46 Both in previous research and in this study,4,12 chloramphenicol resistance has been linked predominantly to the active efflux factor floR, which affects both phenicols.53
In line with previous studies, betalactam resistance mainly affected penicillins,4,6,54,55 highlighting the role of ground beef as a reservoir of penicillin-resistant SE and as a factor that may contribute to disseminating AMR genes against this antibiotic class.
Fortunately, resistance to third- and fourth-generation cephalosporins (3GC and 4GC) and carbapenems was rare or absent. These findings are consistent with the predominance among study isolates of genes encoding class-A betalactamases (bla-CARB, bla-CTX-M, bla-PSE y bla-TEM), which hydrolize penicillins.56 Conversely, only two isolates (one of serovar Typhimurium and one of serovar Newport) carried the bla-CMY gene, which encodes a class-C betalactamase that hydrolyzes both penicillins and cephalosporins.56
Carbapenems, 3GC, and 4GC are considered by WHO as critically important and high priority antibiotics as they are among the few alternatives to treat serious infections caused by Gram-negative bacteria.19 Fortunately, results from this research and from previous studies3,4,12 suggest that beef cattle, at least to date, are not a relevant source of SE resistant to these antibiotic classes. Despite these findings, the progressive loss of effectiveness of antibiotics that have been widely used in animal production is leading to their substitution with more potent ones, such as 3GC (eg. ceftriaxone),46 which is critically important for human medicine.19
Considering 3GC resistance in foodborne SE in Mexico is low, it is important to revise its authorization for veterinary medicine. In Canada and the United States of America, restricting the use of 3GC reserved for veterinary (ceftiofur) and human medicine (ceftriaxone) in animals has contributed to reduce resistance against this vital antibiotic class in SE associated with food-producing animals.50
Regarding fluoroquinoles, we did not observe an association between point mutations and resistance phenotypes, despite mutations being considered the most important mechanism of resistance against these antibiotics in enterobacteria. 57 Apparently, this can be explained by the fact that the mutations carried by our isolates were outside the quinolone resistance-determining region.58
In line with previous studies,4,12 our results confirm that the presence of PMQR genes (qnrA and qnrB alleles) is an increasingly common feature of SE associated with cattle in Mexico. Their wide dissemination is consistent with its plasmid origin as well as with the decreased susceptibility to ciprofloxacin in the isolates that carried these genes, which is reflected in high rates of intermediate resistance (40-50 %).10,12 Taken together, the experimental evidence accumulated to date documents the rising abundance of PMQR genes in SE of bovine origin. Moreover, it suggests that these genes may be subjected to selective pressure in the production context, which is consistent with the authorization of enrofloxacin and ciprofloxacin to treat infections in this species.46
To date, there is no evidence supporting PMQR genes to confer fluoroquinolone resistance in clinical settings. However, their increasing dissemination among foodborne SE populations, as well as the reported association between the emergence of fluoroquinolone resistance phenotypes and the presence of PMQR genes,59,60 highlights the need for further research in this field.
As recently observed,12 100 % of our isolates carried a mutation in the acrB gene. Although this phenomenon has been linked to resistance to macrolides in typhoidal SE strains,36 the mutation observed here (M964T) was not associated with azithromycin resistance. Moreover, as far as we could investigate, it has not been reported as a relevant mechanism of resistance to macrolides.
Strikingly, however, nearly 15 % of our isolates resisted azithromycin and carried genes encoding macrolides’ enzymatic modification mechanisms: mph(A) and lnu(F). In this particular case, we do not observe association with the use of this antibiotic in animals as it is not registered for that purpose in Mexico.46 Mexican studies conducted in the last decade (2000-2017) do not report azithromycin resistance in foodborne SE. Likewise, our research team recently documented a low prevalence of AMR genes against macrolides in SE associated with avian and bovine species.4,12 Hence, further research is needed in this field to determine if the emergence of macrolide resistance observed here is regional or if it is linked to the use of other macrolides (eg. tylosine) that are registered for veterinary use.46
Our study isolates also carried AMR genes against antibiotics that were not included in the AST panel. For instance, fosfomycin is considered a critically important antimicrobial, and is frequently used to treat urinary tract infections in humans.61 Nevertheless, it is also registered in Mexico for therapeutic purposes in cattle and pigs,46 which might explain the presence of these genes in nearly 15 % of our isolates (supplementary information, Table S2).
Along the same lines, although only one isolate carried the mcr1.1 gene, which confers resistance to colistin,62 the finding is not minor since polymyxins are last-resort antibiotics to treat serious infections caused by Gram negative bacteria. (19 Moreover, this gene is plasmid-borne62 and consequently this property could facilitate its dissemination among bacterial populations associated with cattle.
Another important finding is the widespread distribution among study isolates of resistance genes against disinfectants of the quaternary ammonium compounds group (qacL, qacE, and qacEdelta-1). These genes encode active efflux factors,63 and both in this study (supplementary information, Table S2) and previous research,4 they have been linked to MDR phenotypes since they are usually recruited in integrons with other AMR genes.64 It is yet to be determined, however, if the intensive use of quaternary ammonium compounds along the food chain favors the acquisition and conservation of integrons with multiple resistance gene cassettes.
A moderately high proportion of isolates (~40 %) exhibited worrisome MDR phenotypes involving antibiotic classes considered by WHO as critically important and highest priority (macrolides), critically important and high priority (aminoglycosides, penicillins), and very important (tetracyclines, phenicols, sulfonamides).19 The association observed here between the frequency of MDR phenotypes and serovars of public health significance confirms that the acquisition of AMR genes is a hallmark of epidemiologically relevant SE strains.65 Furthermore, it emphasizes the need to keep AMR surveillance in foodborne SE.
Our results suggest that some AMR genes could be subjected to selective pressure in SE associated with animal production. In that sense, we observed that our isolates carried between two and three genes encoding different AMR factors against the same antibiotic class. Furthermore, although some isolates did not carry any of the known AMR genes, they still exhibited resistant phenotypes. These findings reveal the existence of convergent evolution of SE strains, as shown by the accumulation of diverse AMR mechanisms against the same antibiotic, at the population scale, in apparent response to the selective pressure they face in this ecological niche. However, further studies are needed on AMR evolution dynamics in SE populations circulating in cattle, an area that goes beyond the scope of this research.
In summary, this study demonstrates the role of retail ground beef, sold across the state capitals of Central Mexico, as a reservoir of SE, and particularly of epidemiologically relevant strains with strong MDR phenotypes.
Our results suggest that SE populations are massively acquiring AMR genes against antibiotic classes that are used in both human and veterinary medicine. In addition, the wide repertoire of AMR genes against the antibiotic classes considered here suggests that the pathogen is exposed to these drugs in the production setting. Of particular importance is the apparent emergence of azithromycin resistance, which is one of the few alternatives to treat invasive salmonellosis. To date, this phenotype has rarely been found in foodborne SE in Mexico.3
The above analysis highlights the need to revise the authorization of some antibiotics for therapeutic use in livestock species. Moreover, it is important to determine whether there is cross resistance between antibiotics from the same class that are reserved for use in animals and humans. Finally, although this study involved an acceptable number of isolates, the sampling scheme conducted here is not comprehensive. Therefore, wider surveys are required to provide more precise estimations and more thorough characterization of AMR in SE populations circulating in this ecological niche.
Acknowledgments
We appreciate the collaboration of personnel and undergraduate and graduate students from the Faculty of Veterinary Medicine, National Autonomous University of Mexico. We also thank the personnel from the National Reference Center of Food Safety and Agri-Food Biosecurity, and the National Institute of Public Health.
REFERENCES
1. World Health Organization. WHO estimates of the global burden of food-borne diseases. Foodborne disease burden epidemiology reference group 2007-2015 (PDF). 2015. https://apps.who.int/iris/bitstream/handle/10665/199350/9789241565165_eng.pdf [ Links ]
2. Secretaría de Salud. Anuarios de morbilidad (PDF). 2021. https://epidemiologia.salud.gob.mx/anuario/2021/morbilidad/nacional/distribucion_casos_nuevos_enfermedad_fuente_notificacion.pdf [ Links ]
3. Godinez-Oviedo, A., Tamplin, ML., Bowman, JP., Hernandez-Iturriaga, M. Salmonella enterica in Mexico 2000-2017: epidemiology, antimicrobial resistance, and prevalence in food. Foodborne Pathogens and Disease. 2020;17(2):98-118. doi: 10.1089/fpd.2019.2627. [ Links ]
4. Delgado-Suárez, EJ., Ortíz-López, R., Gebreyes, WA., Allard, MW., Barona-Gomez, F., Rubio-Lozano, MS. Genomic surveillance links livestock production with the emergence and spread of multi-drug resistant non-typhoidal Salmonella in Mexico. Journal of Microbiology. 2019;57(4). doi: 10.1007/s12275-019-8421-3. [ Links ]
5. Ma, Y., Li, M., Xu, X., Fu, Y., Xiong, Z., Zhang, L., et al. High-levels of resistance to quinolone and cephalosporin antibiotics in MDR-ACSSuT Salmonella enterica serovar Enteritidis mainly isolated from patients and foods in Shanghai, China. International Journal of Food Microbiology. 2018;286:190-196. doi: 10.1016/j.ijfoodmicro.2018.09.022. [ Links ]
6. Ballesteros-Nova, N., Rubio-Lozano, MS., Delgado-Suárez, EJ., Méndez-Medina, RD., Braña-Varela, D., Rodas Suárez, O. Perfil de resistencia a antibióticos de serotipos Salmonella spp. aislados de carne de res molida en la Ciudad de México. Salud Pública de México. 2016;58(3):1-7. doi: 10.21149/spm.v58i3.7897. [ Links ]
7. Almeida, F., Seribelli, AA., Medeiros, MIC., Rodrigues, DDP., de Mello Varani, A., Luo, Y., et al. Phylogenetic and antimicrobial resistance gene analysis of Salmonella Typhimurium strains isolated in Brazil by whole genome sequencing. PLoS One. 2018;13(8):e0201882. doi: 10.1371/journal.pone.0201882. [ Links ]
8. Laufer, AS., Grass, J., Holt, K., Whichard, JM., Griffin, PM., Gould, LH. Outbreaks of Salmonella infections attributed to beef -United States, 1973-2011. Epidemiology and Infection. 2015;143(9):2003-2013. doi: 10.1017/S0950268814003112. [ Links ]
9. De laGarza-García, JA., Rubio Lozano, MS., Wacher-Rodarte, MDC., Navarro Ocaña, A., Hernández-Castro, R., Xicohtencatl-Cortes, J., et al. Frecuencia de contaminación y de serotipos de Salmonella enterica y Escherichia coli en una operación integrada de matanza y deshuese de bovinos. Revista Mexicana de Ciencias Pecuarias. 2020;11(4):971-990. doi: 10.22319/rmcp.v11i4.5111. [ Links ]
10. Realpe-Quintero, M., Barba-Leon, J., Perez-Montano, JA., Pacheco-Gallardo, C., Gonzalez-Aguilar, D., Dominguez-Arias, RM., et al. Genetic diversity and antimicrobial resistance of Salmonella serotypes recovered throughout the beef production chain and from patients with salmonellosis. PeerJ. 2018;6:e5482. doi: 10.7717/peerj.5482. [ Links ]
11. Cabrera-Diaz, E., Barbosa-Cardenas, CM., Perez-Montano, JA., Gonzalez-Aguilar, D., Pacheco-Gallardo, C., Barba, J. Occurrence, serotype diversity, and antimicrobial resistance of Salmonella in ground beef at retail stores in Jalisco state, Mexico. Journal of Food Protection. 2013;76(12):2004-2010. doi: 10.4315/0362-028X.JFP-13-109. [ Links ]
12. Delgado-Suarez, EJ., Palos-Guiterrez, T., Ruiz-Lopez, FA., Hernandez Perez, CF., Ballesteros-Nova, NE., Soberanis-Ramos, O, et al. Genomic surveillance of antimicrobial resistance shows cattle and poultry are a moderate source of multi-drug resistant non-typhoidal Salmonella in Mexico. PLoS One. 2021;16(5):e0243681. doi: 10.1371/journal.pone.0243681. [ Links ]
13. Aguilar-Barojas, S. Fórmulas para el cálculo de la muestra en investigaciones de salud. Salud en Tabasco. 2005;11(1-2):333-338. [ Links ]
14. Instituto Nacional de Estadística y Geografía. Directorio Estadístico Nacional de Unidades Económicas. 2018. https://www.inegi.org.mx/app/descarga/?ti=6 [ Links ]
15. NORMA Oficial Mexicana NOM-210-SSA1-2014. Productos y servicios. Métodos de prueba microbiológicos. Determinación de microorganismos indicadores. Determinación de microorganismos patógenos. https://dof.gob.mx/nota_detalle_popup.php?codigo=5398468 [ Links ]
16. Rahn, K., De Grandis, SA., Clarke, RC., Curtiss, R., Gyles, CL. Amplification of an invA gene sequence of Salmonella Typhimurium by polymerase chain reaction as a specific method of detection of Salmonella. Molecular and Cellular Probes. 1992;6:271-279. doi: 10.1016/0890-8508(92)90002-F. [ Links ]
17. Delgado Suárez, EJ. Salmonella spp. detection and isolation. Protocols.io repository. 2021. doi: https://dx.doi.org/10.17504/protocols.io.bpybmpsn. [ Links ]
18. Bauer, AW., Kirby, WM., Sherris, JC., Turck, M. Antibiotic susceptibility testing by a standardized single disk method. American Journal of Clinical Pathology. 1966;45(4):493-496. [ Links ]
19. World Health Organization. Critically important antimicrobials for human medicine, 6th Revision (PDF). Antimicrobial Resistance Division, Global Coordination and Partnership, Nutrition and Food Safety; 2019. https://www.who.int/publications/i/item/9789241515528 [ Links ]
20. Clinical and Laboratory Standards Institute. CLSI M100-ED31:2021 Performance Standards for Antimicrobial Susceptibility Testing. 31st edition. 2021. http://em100.edaptivedocs.net/GetDoc.aspx?doc=CLSI%20M100%20ED31:2021&xormat=SPDF&src=BB [ Links ]
21. Magiorakos, AP., Srinivasan, A., Carey, RB., Carmeli, Y., Falagas, ME., Giske, CG., et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clinical Microbiology and Infection. 2012;18(3):268-281. doi: 10.1111/j.1469-0691.2011.03570.x. [ Links ]
22. Delgado Suárez, EJ. Salmonella spp. antibiotic susceptibility testing by the Kirby-Bauer disk diffusion method. Protocols.io repository. 2021. doi: 10.17504/protocols.io.bpypmpvn. [ Links ]
23. Yoshida, CE., Kruczkiewicz, P., Laing, CR., Lingohr, EJ., Gannon, VP., Nash, JH., et al. The Salmonella In Silico Typing Resource (SISTR): an open web-accessible tool for rapidly typing and subtyping draft Salmonella genome assemblies. PLoS One. 2016;11(1):e0147101. doi: 10.1371/journal.pone.0147101. [ Links ]
24. Andrews, S. FastQC: a quality control tool for high throughput sequence data. 2010. http://www.bioinformatics.babraham.ac.uk/projects/fastqc [ Links ]
25. Bolger, AM., Lohse, M., Usadel, B. Trimmomatic: a flexible trimmer for Illumina Sequence Data. Bioinformatics. 2014;30(15):2114-2120. doi: 10.1093/bioinformatics/btu170. [ Links ]
26. Wattam, AR., Abraham, D., Dalay, O., Disz, TL., Driscoll, T., Gabbard, JL., et al. PATRIC, the bacterial bioinformatics database and analysis resource. Nucleic Acids Research. 2014;42(D1):D581-D591. doi: 10.1093/nar/gkt1099. [ Links ]
27. Bankevich, A., Nurk, S., Antipov, D., Gurevich, AA., Dvorkin, M., Kulikov, AS., et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. Journal of Computational Biology. 2012;19(5):455-477. doi: 10.1089/cmb.2012.0021. [ Links ]
28. Gurevich, A., Saveliev, V., Vyahhi, N., Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics. 2013;29(8):1072-1075. doi: 10.1093/bioinformatics/btt086. [ Links ]
29. Deneke, C., Brendebach, H., Uelze, L., Borowiak, M., Malorny, B., Tausch, SH. Species-specific quality control, assembly and contamination detection in microbial isolate sequences with AQUAMIS. Genes (Basel). 2021;12(5). doi: 10.3390/genes12050644. [ Links ]
30. Feldgarden, M., Brover, V., Haft, DH., Prasad, AB., Slotta, DJ., Tolstoy, I., et al. Validating the AMRFinder tool and resistance gene database by using antimicrobial resistance genotype-phenotype correlations in a collection of isolates. Antimicrobial Agents and Chemotherapy. 2019;63(11):e00483-19. doi: 10.1128/AAC.00483-19. [ Links ]
31. Tack, B., Phoba, MF., Thong, P., Lompo, P., Hupko, C., Desmet, S., et al. Epidemiological cut-off value and antibiotic susceptibility test methods for azithromycin in a collection of multi-country invasive non-typhoidal Salmonella. Clinical Microbiology and Infection. 2022;28(12):1615-1623. doi: 10.1016/j.cmi.2022.06.009. [ Links ]
32. Lee, S., Park, N., Yun, S., Hur, E., Song, J., Lee, H., et al. Presence of plasmid-mediated quinolone resistance (PMQR) genes in non-typhoidal Salmonella strains with reduced susceptibility to fluoroquinolones isolated from human salmonellosis in Gyeonggido, South Korea from 2016 to 2019. Gut Pathogens. 2021;13(1):35. doi: 10.1186/s13099-021-00431-7. [ Links ]
33. Nguyen, F., Starosta, AL., Arenz, S., Sohmen, D., Donhofer, A., Wilson, DN. Tetracycline antibiotics and resistance mechanisms. Biological Chemistry. 2014;395(5):559-575. doi: 10.1515/hsz-2013-0292. [ Links ]
34. Escudeiro, P., Pothier, J., Dionisio, F., Nogueira, T. Antibiotic resistance gene diversity and virulence gene diversity are correlated in human gut and environmental microbiomes. mSphere. 2019;4(3):e00135-19. doi: 10.1128/mSphere.00135-19. [ Links ]
35. Abouzeed, YM., Baucheron, S., Cloeckaert, A. ramR mutations involved in efflux-mediated multidrug resistance in Salmonella enterica serovar Typhimurium. Antimicrobial Agents and Chemotherapy. 2008;52(7):2428-2434. doi: 10.1128/AAC.00084-08. [ Links ]
36. Hooda, Y., Sajib, MSI., Rahman, H., Luby, SP., Bondy-Denomy, J., Santosham, M., et al. Molecular mechanism of azithromycin resistance among typhoidal Salmonella strains in Bangladesh identified through passive pediatric surveillance. PLoS Neglected Tropical Diseases. 2019;13(11):e0007868. doi: 10.1371/journal.pntd.0007868. [ Links ]
37. Hooper, DC., Jacoby, GA. Mechanisms of drug resistance: quinolone resistance. Annals of the New York Academy of Sciences. 2015;1354(1):12-31. doi: 10.1111/nyas.12830. [ Links ]
38. Quesada, A., Porrero, MC., Tellez, S., Palomo, G., Garcia, M., Dominguez, L. Polymorphism of genes encoding PmrAB in colistin-resistant strains of Escherichia coli and Salmonella enterica isolated from poultry and swine. Journal of Antimicrobial Chemotherapy. 2015;70(1):71-74. doi: 10.1093/jac/dku320. [ Links ]
39. Nuncio, ASP., Webber, B., Pottker, ES., Cardoso, B., Esposito, F., Fontana, H., et al. Genomic characterization of multidrug-resistant Salmonella Heidelberg E2 strain isolated from chicken carcass in southern Brazil. International Journal of Food Microbiology. 2022;379:109863. doi: 10.1016/j.ijfoodmicro.2022.109863. [ Links ]
40. Martinez-Chavez, L., Cabrera-Diaz, E., Perez-Montano, JA., Garay-Martinez, LE., Varela-Hernandez, JJ., Castillo, A., et al. Quantitative distribution of Salmonella spp. and Escherichia coli on beef carcasses and raw beef at retail establishments. International Journal of Food Microbiology. 2015;210:149-155. doi: 10.1016/j.ijfoodmicro.2015.06.016. [ Links ]
41. Palós Gutiérrez, T., Rubio Lozano, MS., Delgado Suárez, EJ., Rosi Guzmán, N., Soberanis Ramos, O., Hernández Pérez, CF., et al. Lymph nodes and ground beef as public health importance reservoirs of Salmonella spp. Revista Mexicana de Ciencias Pecuarias. 2020;11(3):795-810. doi: 10.22319/rmcp.v11i3.5516. [ Links ]
42. Pond, A., Miller, M., Echeverry, A., Huerta-Leidenz, N., Rubio-Lozano, MS., Chávez, A., et al. Salmonella and E. coli O157:H7 prevalence and generic E. coli and coliform quantitative baseline in raw pork and beef in retail channels in Mexico. Food Protection Trends. 2016;67(9):2069-2089. [ Links ]
43. Aslam, M., Checkley, S., Avery, B., Chalmers, G., Bohaychuk, V., Gensler, G., et al. Phenotypic and genetic characterization of antimicrobial resistance in Salmonella serovars isolated from retail meats in Alberta, Canada. Food Microbiology. 2012;32(1):110-117. doi: 10.1016/j.fm.2012.04.017. [ Links ]
44. Yin, M., Yang, B., Wu, Y., Wang, L., Wu, H., Zhang, T., et al. Prevalence and characterization of Salmonella enterica serovar in retail meats in market place in Uighur, Xinjiang, China. Food Control. 2016;64:165-172. doi: 10.1016/j.foodcont.2015.12.029. [ Links ]
45. Centers for Disease Control and Prevention. Outbreak of Salmonella infections linked to ground beef (HTML). 2018. https://www.cdc.gov/salmonella/newport-10-18/index.html [ Links ]
46. Servicio Nacional de Sanidad, Inocuidad y Calidad Agroalimentaria. Productos químico-farmacéuticos vigentes (CSV). Secretaría de Agricultura y Desarrollo Rural, Gobierno de México; 2022. https://datos.gob.mx/busca/dataset/productos-quimicos-farmaceuticos-vigentes [ Links ]
47. Zaidi, MB., McDermott, P., Fedorka-Cray, P., Leon, V., Canche, C., Hubert, S., et al. Nontyphoidal Salmonella from human clinical cases, asymptomatic children and raw retail meats in Yucatan, Mexico. Clinical Infectious Diseases. 2006;42:21-28. doi: 10.1086/498508. [ Links ]
48. Miranda, JM., Mondragón, AC., Martínez, B., Guarddon, M., Rodríguez, JA. Prevalence and antimicrobial resistance patterns of Salmonella from different raw foods in Mexico. Journal of Food Protection. 2009;72(5):966-971. doi: 10.4315/0362-028X-72.5.966. [ Links ]
49. Perez-Montaño, JA., González-Aguilar, D., Barba, J., Pacheco-Gallardo, C., Campos-Bravo, CA., García, S., et al. Frequency and antimicrobial resistance of Salmonella serotypes on beef carcasses at small abattoirs in Jalisco state, Mexico. Journal of Food Protection. 2012;75(5):867-873. doi: 10.4315/0362-028X.JFP-11-423. [ Links ]
50. McDermott, PF., Zhao, S., Tate, H. Antimicrobial resistance in nontyphoidal Salmonella. Microbiology Spectrum. 2018;6(4):ARBA-0014-2017. doi: 10.1128/microbiolspec.ARBA-0014-2017. [ Links ]
51. Akhtar, M., Hirt, H., Zurek, L. Horizontal transfer of the tetracycline resistance gene tetM mediated by pCF10 among Enterococcus faecalis in the house fly (Musca domestica L.) alimentary canal. Microbial Ecology. 2009;58(3):509-518. doi: 10.1007/s00248-009-9533-9. [ Links ]
52. Yousefi-Nooraie, R., Mortaz-Hejri, S., Mehrani, M., Sadeghipour, P. Antibiotics for treating human brucellosis. Cochrane Database of Systematic Reviews. 2012;10:CD007179. doi: 10.1002/14651858.CD007179.pub2. [ Links ]
53. Kim, S., Kim, SH., Chun, SG., Park, MS., Lim, HM., Lee, BK. An additional novel antimicrobial resistance gene cluster in Salmonella Genomic Island 1 of a Salmonella enterica Serovar Typhimurium DT104 human isolate. Foodborne Pathogens and Disease. 2009;6(4):471-479. doi: 10.1089/fpd.2008.0199. [ Links ]
54. Aguilar-Montesde Oca, S., Talavera-Rojas, M., Soriano-Vargas, E., Barba-Leon, J., Vazquez-Navarrete, J., Acosta-Dibarrat, J., et al. Phenotypic and genotypic profile of clinical and animal multidrug-resistant Salmonella enterica isolates from Mexico. Journal of Applied Microbiology. 2018;124(1):67-74. doi: 10.1111/jam.13615. [ Links ]
55. Villalpando-Guzman, S., Vázquez-Quiñones, CR., Natividad-Bonifacio, I., Curiel-Quesada, E., Quiñones-Ramírez, EI., Vázquez-Salinas, C. Frecuencia, susceptibilidad antimicrobiana y patrón de adherencia de Salmonella enterica aislada de carne de pollo, res y cerdo de la Ciudad de México. Revista Chilena de Infectología. 2017;34(5):458-466. doi: 10.4067/S0716-10182017000500458. [ Links ]
56. Mora-Ochomogo, M., Lohans, CT. β-lactam antibiotic targets and resistance mechanisms: from covalent inhibitors to substrates. RSC Medicinal Chemistry. 2021;12(10):1623-1639. doi: 10.1039/d1md00200g. [ Links ]
57. Qiu, H., Gong, J., Butaye, P., Lu, G., Huang, K., Zhu, G., et al. CRISPR/Cas9/sgRNA-mediated targeted gene modification confirms the cause-effect relationship between gyrA mutation and quinolone resistance in Escherichia coli. FEMS Microbiology Letters. 2018;365(13). doi: 10.1093/femsle/fny127. [ Links ]
58. Thong, KL., Ngoi, ST., Chai, LC., The, CS. Quinolone resistance mechanisms among Salmonella enterica in Malaysia. Microbial Drug Resistance. 2016;22(4):259-272. doi: 10.1089/mdr.2015.0158. [ Links ]
59. Pribul, BR., Festivo, ML., Rodrigues, MS., Costa, RG., Rodrigues, EC., de Souza, MM., et al. Characteristics of quinolone resistance in Salmonella spp. isolates from the food chain in Brazil. Frontiers in Microbiology. 2017;8:299. doi: 10.3389/fmicb.2017.00299. [ Links ]
60. Karp, BE., Campbell, D., Chen, JC., Folster, JP., Friedman, CR. Plasmid-mediated quinolone resistance in human non-typhoidal Salmonella infections: an emerging public health problem in the United States. Zoonoses and Public Health. 2018;65(7):838-849. doi: 10.1111/zph.12507. [ Links ]
61. Beharry, Z., Palzkill, T. Functional analysis of active site residues of the fosfomycin resistance enzyme FosA from Pseudomonas aeruginosa. Journal of Biological Chemistry. 2005;280(18):17786-17791. doi: 10.1074/jbc.M501052200. [ Links ]
62. Liu, YY., Wang, Y., Walsh, TR., Yi, LX., Zhang, R., Spencer, J., et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. The Lancet Infectious Diseases. 2016;16(2):161-168. doi: 10.1016/S1473-3099(15)00424-7. [ Links ]
63. Castro, VS., Mutz, YDS., Rosario, DKA., Cunha-Neto, A., Figueiredo, EES., Conte-Junior, CA. Inactivation of multi-drug resistant non-typhoidal Salmonella and wild-type Escherichia coli STEC using organic acids: a potential alternative to the food industry. Pathogens. 2020;9(10). doi: 10.3390/pathogens9100849. [ Links ]
64. Poole, K. Bacterial stress responses as determinants of antimicrobial resistance. Journal of Antimicrobial Chemotherapy. 2012;67(9):2069-2089. doi: 10.1093/jac/dks196. [ Links ]
65. Leekitcharoenphon, P., Hendriksen, RS., Le Hello, S., Weill, FX., Baggesen, DL, Jun, SR., et al. Global genomic epidemiology of Salmonella enterica Serovar Typhimurium DT104. Applied and Environmental Microbiology. 2016;82(8):2516-2526. doi: 10.1128/AEM.03821-15. [ Links ]
Data availability. All relevant data are within the manuscript and its supporting information files. Raw sequences are publicly available at NCBI (https://www.ncbi.nlm.nih.gov/sra) through the accession numbers and metadata reported as supplementary information (Table S1).
Funding statement. This research was funded by the Mexican Council of Science and Technology, “2019 Cutting-Edge Science” funding announcement, grant number CY087198, granted to LM Sánchez-Zamorano y EJ Delgado-Suárez. The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Received: May 09, 2023; Accepted: June 13, 2023; Published: September 05, 2023











 texto en
texto en