Ipomoea L. is the largest genus of Convolvulaceae and is one of the largest genera of angiosperms, with approximately 800 species (Wood et al. 2020). The distribution of the genus is pantropical, and even reaches temperate regions, with 425 species native to the New World (Wood et al. 2020, Muñoz-Rodríguez et al. 2023). Diverse life forms such as herbs, shrubs, vines, lianas, and trees have been documented in the genus (Muñoz-Rodríguez et al. 2023). Although most of its species have cordate or pandurate leaves, sagittate, reniform, retuse, pinnate, oblong, and palmate leaf forms have also been recorded (Inamdar & Shenoy 1982, Simões & Staples 2017, Aye & Lwin 2018, Wood et al. 2020, Paramesh & Reddy 2021). The flowers are trumpet- or funnel-shaped with variable colors. They can be lavender, purple, or whitish, frequently darker inside the floral tube, stamens unequal, ovary 3-loculate, with trilobed stigma (Inamdar & Shenoy 1982, Simões & Staples 2017, Aye & Lwin 2018, Wood et al. 2020, Paramesh & Reddy 2021).
Ipomoea species are frequently known as “Morning Glory” because the flowers of many cultivated species such as I. indica (Burm.) Merr. open at dawn and close by midday (Wood et al. 2020). In addition to sweet potato (Ipomoea batatas (L.) Poir.), several species of the genus are edible. The leaves of some Ipomoeas are eaten, e.g., the water spinach I. aquatica Forssk, native to Southeast Asia. Here, we focus on I. dumosa (Benth.) L.O.Williams, which is another edible species and is commonly known as “Xonequi”. The leaves of this species are added as a flavoring to dishes in several regions of Mexico (Piedra-Malagón et al. 2022). It is a perennial herbaceous vine with petiolate cordate leaves, fibrous roots, and pink or purplish funnel-shaped flowers with exserted stamens and styles (Austin & Huáman 1996, Wood et al. 2020). It has a widespread distribution occurring in both the Sierra Madre Occidental and the Sierra Oriental in Mexico, reaching Central America and the north of South America (Wood et al. 2020). Leaves of I. dumosa have been utilized by different ethnic communities in Mexico as a condiment and for their antioxidant and medicinal properties for humans and animals during pregnancy, birth, and puerperium ailments (Heindorf et al. 2020, De Jesús et al. 2024, Espinoza-Pérez et al. 2024).
Several molecular phylogenies have been implemented in Ipomoea, the majority based on the nuclear marker ITS with distinct sampling (Miller et al. 1999, Manos et al. 2001, Miller et al. 2004, Zhao et al. 2019, Muñoz-Rodríguez et al. 2024). In the ITS phylogeny of New World Ipomoea, I. dumosa was retrieved in the “Quamoclit clade”, in a small group with I. seducta House and I. uhdeana D.F.Austin, and with a sister clade comprising I. aristolochiifolia G.Don, I. variabilis (Schltdl. & Cham) Choisy, I. meyeri (Spreng.) G.Don, and I. expansa J.A.McDonald (Wood et al. 2020). However, these phylogenies were poorly resolved and, considering the status of Ipomoea as one of the largest genera of the angiosperms, it has been indicated that diagnostic characters along with the use of novel molecular data would help clarify the recognition of monophyletic groups within the genus (Muñoz-Rodríguez et al. 2023).
The objectives of this study are to perform phylogenetic analyses based on the accessible plastid genomes of Ipomoea to determine the position of I. dumosa and to assemble and annotate its plastome as the first step to further elucidate genetic differences related to the management of this species by different ethnic groups. Genomic approaches are a valuable resource for understanding the domestication and management of underutilized edible plants (Li et al. 2021, Teshome et al. 2024, Chai & Sun 2025). Notwithstanding the enormous number of members of this genus, only 27 plastid genomes of Ipomoea species have been fully sequenced (e.g., Ipomoea alba L., I. batatas (L.) Lam., I. nil (L.) Roth, I. purpurea (L.) Roth, I. quamoclit L., I. trifida (Kunth) G.Don, I. triloba L., etc.) and utilized to determine phylogenetic relationships (Park et al. 2018, Sun et al. 2019, Xiao et al. 2021, Yang et al. 2022, Wang et al. 2021, 2023, Laux et al. 2022, Li et al. 2024, Sudmoon et al. 2024). We hypothesize that I. dumosa will prove to be related to species previously identified in the “Quamoclit clade” by Wood et al. (2020).
Materials and methods
Plant material. Young leaves of Ipomoea dumosa were collected from the living collections of the Francisco Javier Clavijero Botanic Garden, placed in liquid nitrogen, and processed. The source plants were originally wild-collected in Veracruz and the voucher was deposited at XAL (Piedra-Malagón, López & Sosa 767). For DNA extraction, the DNeasy Plant Maxi Kit of QIAGEN©, Aarhus, Denmark, was utilized with a sample of 0.85-0.90 gr of leaves, yielding a DNA concentration between 10.00 to 11.34 μg.
Genome sequencing, assembly, and annotation. Sequencing was performed by Macrogen (www.macrogen.com) using the PacBio HiFi technology. Assembly was performed with Organelle_PBA following default parameters in the Instituto de Ecología, AC. bioinformatic cluster (Soorni et al. 2017). The published Ipomoea nil plastome (Park et al. 2018) was used in Organelle_PBA to search for reads of organelle origin and then perform a de novo assembly. Annotation and image construction of the plastome genome were performed with CPGAVAS2, using the GeneBank annotation of Ipomoea nil as a reference (Shi et al. 2019). The resulting plastome genome of Ipomoea dumosa can be reviewed through GenBank accession PV762244.
Phylogenetic analysis. Based on previous Ipomoea plastome genome research (e.g.,Park et al. 2018, Wang et al. 2023, Sudmoon et al. 2024), twenty-seven Ipomoea plastomes, plus Merremia hederacea, the closest outgroup genome to Ipomoea (Sudmoon et al. 2024), were downloaded from NCBI (www.ncbi.nlm.nih.gov) (Supplementary material, Table S1) and aligned with ClustalW (Thompson et al. 2003) alongside the newly-sequenced I. dumosa plastome genome. The alignment obtained with ClustalW was used to reconstruct the plastome phylogeny of Ipomoea species. The alignment was manually edited using the UGENE v. 42.0 software (Okonechnikov et al. 2012). The alignment flanks that contained more than 50 % gaps were manually removed. The program Trimal v. 1.4.1 (Capella-Gutiérrez et al. 2009) was then applied to the alignment to eliminate any poorly aligned regions. ModelTest-NG v. 0.1.7 (Darriba et al. 2020) was implemented on the non-partitioned alignment (both inversions and inverted repeats were kept) to find the best-fitting evolutionary model.
The Maximum Likelihood tree reconstruction was conducted using the program RAxMLHPC-PTHREADS-SSE3 v. 8.2.12 (Stamatakis 2006) under the best-fit evolutionary model produced by the ModelTest-NG test (GTR+I+G4), with 1,000 bootstrap replicates. The resulting bootstrap output was used as the maximum likelihood tree. Visual representation of the maximum likelihood tree was edited with customized templates on the online platform iTOL v. 6 (Letunic & Bork 2019).
Results
Ipomoea dumosa plastid. The plastome of Ipomoea dumosa has a typical quadripartite structure containing a large single-copy (LSC) region, a small single-copy (SSC) region, and a pair of inverted repeats (IR) regions (Figure 1). The plastome size of I. dumosa was 161,464 bp, containing 88,457 bp in the LSC, 29,771 bp in the IR, and 13,465 in the SSC (Supplementary material, Table S2). The plastome of I. dumosa is composed of 128 genes, including 83 protein-coding sequences (CDS), 37 tRNA, and eight rRNA genes (Supplementary material, Table S3). Total GC content was 37.56 %, while a GC content of 36 % was found in the LSC, 39.34 % in the IR, and 32.35 % in the SSC.
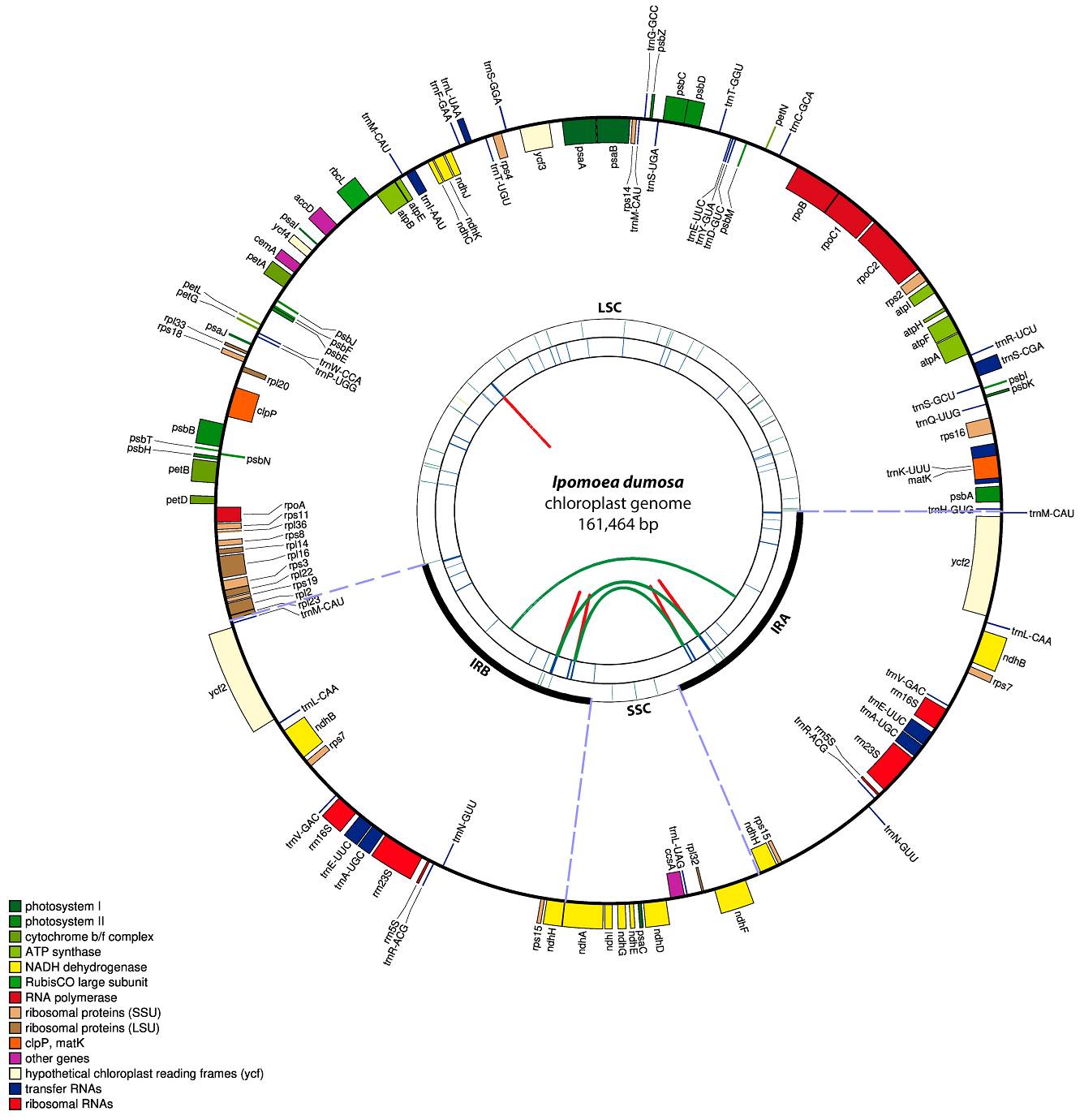
Figure 1 Ipomoea dumosa chloroplast genome structure. Genes placed at the outside and inside of the circle are transcribed counterclockwise and clockwise, respectively. Genes are color-coded based on their functional classification. Functional gene names are shown at the bottom-left part of the figure. GenBank accession PV762244.
Phylogenetic analysis. Based on complete plastomes, the phylogenetic tree of Ipomoea species, including I. dumosa, depicted the outgroup Merremia hederacea (Burm f.) Hallier as closest to Ipomoea biflora (L.) Pers, I. dumosa as sister to I. quamoclit L. (bootstrap bst 100), followed by two clades, the first comprising I. maurandioides Meisn. and I. asarifolia Roem. & Schult (bts 100), and the second comprising I. goyazensis Gardner and I. cavalcantei D.F.Austin (bts 94) (Figure 2). Two major clades were retrieved (Figure 2). Clade 1 had I. pes-caprae (L.) Roth as a sister lineage and comprised the following species: I. alba L., I. purpurea (L) Roth, I. indica (Burm) Merr, I. nil, I. imperati (Vahl) Griseb., I. aquatica, I. cairica (L.) Sweet, I. tiliifolia (Desr.) Roem. & Schult., and I. obscura (L.) Kew Gawl (bts 89). Between these two major clades, there is a small clade comprised of I. carnea Jacq. and I. marabensis D.F.Austin & Secco (bts 100) (Figure 2). Clade 2 included I. splendor-sylvae House as a sister lineage and comprised I. trifida (Kunth) G.Don, I. tabascana J.A.McDonald & D.FAustin, I. batatas, I. lacunosa L., I. cordatotriloba Dennst., I. ramosissima (Poir). Choisy, I. triloba L., and I. cynanchifolia Meisn. (bts 100) (Figure 2). In addition, Clade 1 was further divided into two subclades: Clade 1A, comprising I. alba L., I. purpurea (L.) Roth, I. indica (Burm) Merr., and I. nil (bts 100) and Clade 1B, comprising I. imperati (Vahl) Griseb., I. aquatica, I. cairica, I. tiliifolia (Desr.) Roem. & Schult., and I. obscura (bts 97) (Figure 2). High bootstrap values (Hillis & Bull 1993) supported the clades in the phylogenetic tree (Figure 2).
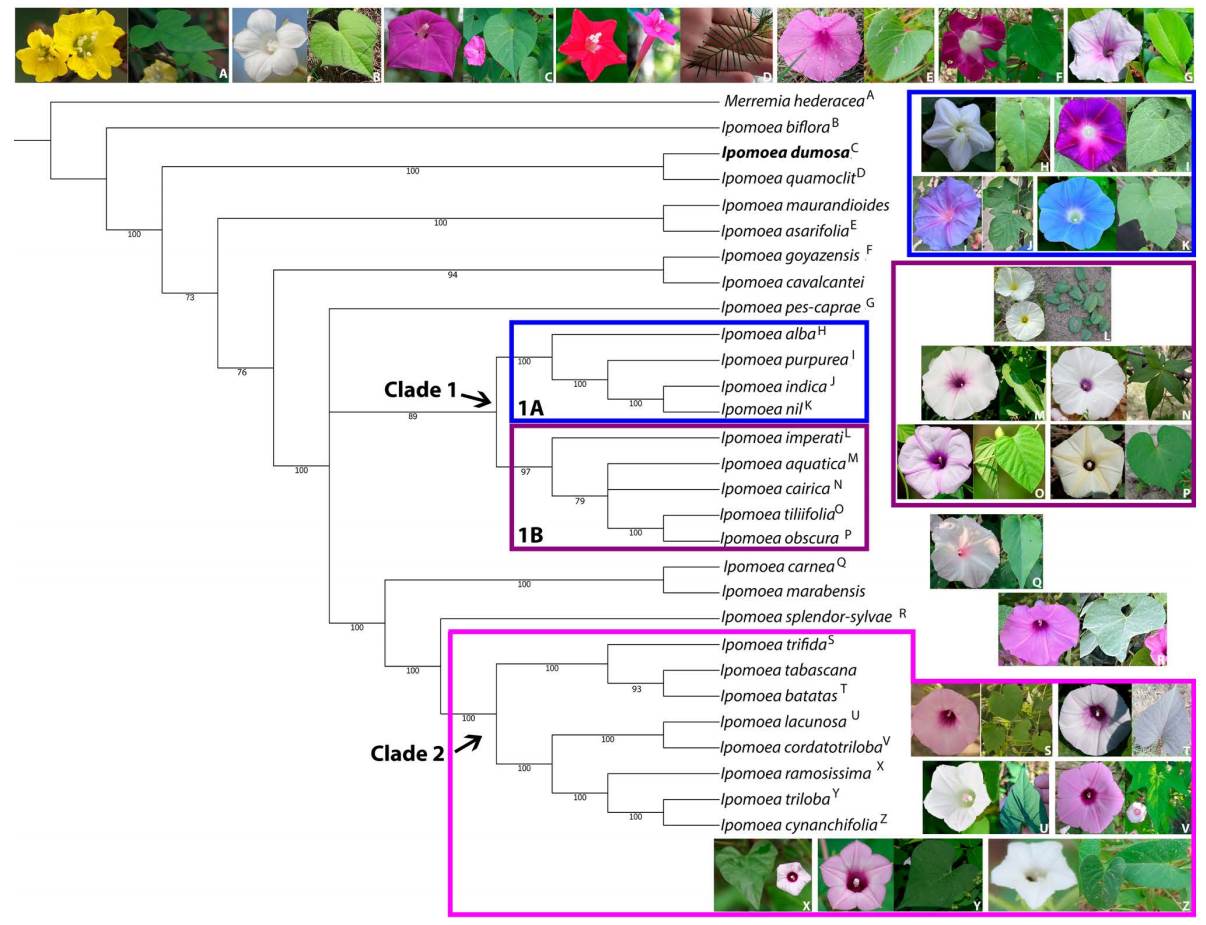
Figure 2 RAxML Ipomoea’s chloroplast genome species phylogenetic consensus tree. A-Z letters: species name referenced to its flower and leaf inset picture. Blue frame: clade 1A. Purple frame: clade 1B. Pink frame: Clade 2. Tree rooted to outgroup species. Ipomoea dumosa name in bold. Bootstrap values shown under tree branches. All images are under the CC0 copyright license.
Discussion
This is the first phylogeny constructed on plastid genomes including Ipomoea dumosa. In our phylogenetic tree, this species is closely related to I. quamoclit. Furthermore, Wood et al. (2020), presented a summarized phylogeny based on the nuclear ITS, the plastid markers matK and trnH-psbA, and 605 nuclear genes, in which different analyses with every set of markers included variable sampling. The summarized tree displays two clades in Ipomoea: The Old World Clade and the New World Clade (Wood et al. 2020). One of the main sub-clades in the New World is a large group, mainly Mexican, including the subgenera proposed by Austin (1979) and Quamoclit as well as some other species. Ipomoea quamoclit forms part of this clade characterized by twining herbs, variable leaves, sub-crateriform corolla, and exserted stamens (Wood et al. 2020) (see Table 1, which includes selected characters for the studied species). Wood et al. (2020) suggested that I. dumosa shares characters with species in the Calonyction clade, such as subcylindrical corolla and exserted stamens, with varying leaf shape, blunt spines, and gemmiform pollen (Table 1; Wood et al. 2020). Contrary to this hypothesis regarding the affinities of I. dumosa, Austin & Huáman (1996) and McDonald et al. (2011) included this species in the subgenus Quamoclit, which presents characters such as a large corolla and homomorphic stamens. In our phylogenetic tree, I. dumosa was determined closely related to I. quamoclit corroborating this hypothesis.
Table 1 General morphological characters of leaves’ shape and flowers stamen/style length of Ipomoea and outgroup species considered in phylogenetic analyses based on Wood et al. (2020)
| Species | Life form | Leaf shape (base/apex) | Corolla | Stamens |
|---|---|---|---|---|
| I. alba L. | Twining | Cordate | Hypocrateriform | Exserted |
| I. aquatica Forssk. | Twining | Sagittate/Cordate | Funnel-shaped | Inserted |
| I. asarifolia Roem. & Schult. | Twining/Prostrate | Reniform | Funnel-shaped | Inserted |
| I. batatas (L.) Lam. | Twining/Erect/Prostrate | Cordate | Funnel-shaped | Inserted |
| I. cairica (L.) Sweet | Twining/Prostrate | Leaflets (Palmate like) | Funnel-shaped | Inserted |
| I. carnea Jacq. | Twining/Erect | Cordate | Campanulate | Inserted |
| I. cavalcantei D.F. Austin | Erect | Oblong | Hypocrateriform | Exserted |
| I. cordatotriloba Dennst. | Twining | 3-5-lobed (Pandurate like) | Inserted | |
| I. cynanchifolia Meisn. | Twining | Cordate | Funnel-shaped | Inserted |
| I. goyazensis Gardmer | Twining | Cordate | Campanulate | Inserted |
| I. imperati (Vahl) Griseb. | Prostrate | Cordate | Campanulate | Inserted |
| I. indica (Burm.) Merr. | Twining | 3-lobed (Pandurate like) | Funnel-shaped | Inserted |
| I. lacunose L. | Twining | Cordate | Funnel-shaped | Inserted |
| I. marabensis D.F.Austin & Secco | Twining | Oblong | Campanulate | Inserted |
| I. maurandioides Meisn. | Twining/Prostrate | Sagittate/Cordate | Campanulate | Inserted |
| I. nil (L.) Roth | Twining | 3-lobed (Pandurate like) | Campanulate | Inserted |
| I. obscura (L.) Ker Gawl. | Twining/Prostrate | Cordate | Funnel-shaped | Inserted |
| I. pes-caprae (L.) R.Br. | Twining/Prostrate | Retuse | Campanulate | Inserted |
| I. purpurea (L.) Roth | Twining | Cordate | Campanulate | Inserted |
| I. quamoclit L. | Twining | Pinnate | Hypocrateriform | Exserted |
| I. ramosissima (Poir.) Choisy | Twining | Cordate | Hypocrateriform | Inserted |
| I. splendor-sylvae House | Twining | Cordate | Funnel-shaped | Inserted |
| I. tabascana J.A.McDonald &D.F Austin | Twining/Prostrate | Sagittate | Funnel-shaped | Inserted |
| I. dumosa Benth. (L.O. Williams) | Twining | Cordate | Hypocrateriform | Exserted |
| I. tiliifolia (Desr.) Roem. & Schult. | Twining | Cordate | Funnel-shaped | Inserted |
| I. trifida (Kunth) G.Don | Twining | Cordate | Funnel-shaped | Inserted |
| I. triloba L. | Twining/Prostrate | Cordate | Campanulate | Inserted |
| Ipomoea biflora (L.) Pers. | Twining | Cordate | Campanulate | Inserted |
| Merremia hederacea | Twining/Prostrate | 3-lobed (Pandurate like) | Campanulate | Exserted |
As stated above, Ipomoea is one of the largest genera of angiosperms and presents challenges in terms of performing phylogenetic analyses with comprehensive sampling, and informative molecular markers, or even genomic data (Muñoz-Rodríguez et al. 2023). Therefore, further analyses using novel molecular data and morphological characters will clarify the relationships and phylogenetic position of I. dumosa, which has been considered in the Calonyction clade or the Quamoclit clade. These analyses will also define whether the genus Merremia should be considered within Ipomoea, as suggested by several authors (Wood et al. 2020). However, we have kept the name Merremia hederacea as indicated in GenBank and utilized it as the outgroup. Moreover, Ipomoea species such as I. alba, I. nil, and I. purpurea from Clade I are among the most important ornamental species in the genus. Species in Clade II such as I. cynanchifolia and I. trifolia share morphological characters including small-sized flowers, with a corolla of less than 3 cm. in length. It is also worth mentioning that I. tabascana has been synonymized under I. batatas in the past (Wood et al. 2020), which agrees with our phylogenetic tree.
In general, recent plastid phylogenetic trees comprising the species of Ipomoea described to date (ours included) show some differences in their topology. In our resulting phylogenetic tree, we highlighted major clades such as Clades 1A, 1B, and 2, and a general level of divergence of species since they were retrieved in a very similar manner to that of previous phylogenetic research (e.g.,Muñoz-Rodríguez et al. 2019, Wood et al. 2020, Sudmoon et al. 2024). The topological dissimilarities for reconstructing plastid phylogenetic trees of Ipomoea species might be attributed to methodological differences in each study case, such as the numbers of Ipomoea species and outgroups included and the particular molecular analysis approach employed.
Furthermore, it has been indicated that I. dumosa is the best-known of a complex of species that possess very similar floral morphology, including I. seducta House and I. tubulata Sessé & Mociño. Ipomoea seducta is recognized by its funnel-shaped corolla, with a distribution in southern Mexico and Guatemala while the diagnostic characters of I. tubulata are a lobed corolla with short, ovate-deltoid lobes and a distribution in western Mexico (Wood et al. 2020). However, observations of the distributions of diverse specimens show that these characters are variable (Wood et al. 2020). Therefore, to elucidate whether these can be recognized as different species, further sequencing of different populations throughout their entire distribution is required.
In classifying Ipomoea, Muñoz-Rodríguez et al. (2023) concluded that existing classifications for the genus should be abandoned and that many segregated genera from Ipomoea should be incorporated into this genus to reconcile the properties of monophyly, diagnosability, and completeness. The study of plastid genomes in Ipomoea can therefore contribute to a better understanding of the systematics and evolution of this genus.
The plastid genome of Ipomoea dumosa shows similar size and characteristics to previous plastid genomes of this genus. The phylogenetic reconstruction based on the plastid genomes indicates that I. dumosa is closely related to I. quamoclit, suggesting that it belongs in the Quamoclit clade. Incorporating I. seducta and I. tubulata, which have been identified as presenting a very similar morphology to I. dumosa, in further phylogenetic analyses will determine whether they are closely related to I. dumosa. Studying plastid genomes in Ipomoea can further our understanding of the systematics and evolution of this genus.
Supplementary material
Supplemental data for this article can be accessed here: https://doi.org/10.17129/botsci.3685.











 nueva página del texto (beta)
nueva página del texto (beta)



