Introduction
Subulo gouazoubira (Bernegossi et al. 2022) is the only native deer that inhabits Uruguayan wooden environments (Black-Décima et al. 2010). Like other deer, it plays a relevant role in forest species composition, stand and canopy structures, and complexity (Vavra et al. 2007; Reed et al. 2022). The diet of gray brocket deer mainly involves fruits, leaves, fibers, and flowers (Prado 2013), and shows significant changes in composition associated with different ecosystems (Gayot et al. 2004; Kufner et al. 2008, Serbent et al. 2011).
Different complementary techniques have been used to evaluate the diet composition for large herbivores (Rayé et al. 2010; Nakahara et al. 2015). Diet studies of Neotropical cervids have been conducted mainly by microhistological analysis of plant fragment recovery from feces and stomach contents (Bodmer 1991; Pinder 1997; Gayot et al. 2004; Kufner et al. 2008; Cosse et al. 2009; Serbent et al. 2011). Although this is a non-invasive method that provide information in terms of species composition, the microhistological analysis is complex and often leads to the ambiguous identification of species in many plant groups (Garnick et al. 2018). The use of molecular techniques for species identification (DNA metabarcoding) reduces taxonomic identification errors (Taberlet et al. 1999; Miller and Waits 2003; Waits and Paetkau 2005; Deagle et al. 2013; Alberdi et al. 2019; Deagle et al. 2019), allowing for a more precise taxonomic resolution than with classic microscopic analysis. For example, the DNA contained in the feces can be amplified by PCR, allowing the fast production of large amounts of genetic information and, by contrasting with DNA barcodes in reference databases, the species present in the samples can be correctly identified (Davison et al. 2002). The power of the technique is based on the chosen genetic marker, which must have sufficient sequence-level differences between the species to be identified (barcode gap), and the level of completeness of the reference database. A marker widely used in DNA metabarcoding of plants is the chloroplast trnL (UAA) intron (Taberlet et al. 2007), a small DNA fragment, variable at sequence level in different species and amplified with universal primers.
Our aim was to evaluate the applicability of DNA barcoding for gray brocket deer dietary analysis related to: i) the resolution level at which the DNA-based method can describe the diet, using a global database enriched with a local acquired database, and ii) the diet composition focusing on native and alien species. To achieve these goals, we conducted next-generation sequencing of the chloroplast trnL (UAA) region from fecal samples obtained by non-invasive sampling to assess the diet of gray brocket deer in a patch of xeric, hillside forest of Uruguay.
Materials and methods
Study site. The study was carried out in the “Reserva Natural Salus (RNS)” in Lavalleja Department, Uruguay (34º 23’ 36.79’’ S; -55º 18’ 58.54’’ W) covering 1,500 ha. The altitudes vary around 497 masl, with the dominant vegetation types being grasslands, hillside shrublands, and native forests limited in extension to hillsides (see Bonifacino and Rossado 2016) for a more detailed description of the study area). In addition, eucalyptus and pine plantations also occur. Gray brocket deer (Subulo gouazoubira) coexist in the area with two exotic ungulates: wild boar (Sus scrofa) and axis deer (Axis axis).
Completion of the local reference database of the trnL (UAA) intron sequences. A local species list was created based on the technical report “Reserva Natural Salus Flora” (Bonifacino and Rossado 2016) and records of observations made by local park rangers (pers. comm. 2022) of highly palatable shrubby species cultivated in residential gardens in the vicinity of the study area and accessed by deer. A local database of trnL (UAA) intron reference sequences for chloroplast DNA (Taberlet et al. 2007) was retrieved from GenBank (Benson et al. 2013) based on matching records with the local species list. Also, trnL (UAA) sequences were generated to cover some missing taxa and local variants of the native species. Native plant species were gathered from various locations, including the botanical garden "Museo y Jardín Botánico de Montevideo", the gardens of the "Facultad de Agronomía-Udelar" the greenhouse of the "Instituto Nacional de Semillas" and the IIBCE Mimosoideae collection at the "Instituto de Investigaciones Biológicas Clemente Estable". Plant vouchers were herborized following standard protocols (Bridson and Forman 1992) and deposited at MVFA herbarium at Facultad de Agronomía-Udelar (see supplementary data). Plant tissue samples were preserved in silica gel until they were dried.
DNA extraction from reference plant specimens was conducted using dried young leaves following Fazekas et al. (2012). FastPrep®- 24 (MP Biomedicals) and lysing matrix beads were used for tissue homogenization and cell disruption. DNA extracts were amplified with the universal c-d primer pair (Taberlet et al. 2007), amplifying the entire trnL (UAA) intron (c: 5’-CGAAATCG GTAGACGCTACG-3’; d: 5’-GGGGATAGAGGGACTTGAAC-3’). Each PCR reaction contained 1X Buffer (Invitrogen), 1.5 mM MgCl (Invitrogen), 0.04 U Taq polymerase (Invitrogen), 0.05 mM each dNTP, 0.2 mg/mL BSA and 0.5 µM forward and reverse primers in a final volume of 20µl. The thermal profile consisted of an initial denaturation step at 94 °C for 3 min, followed by 35 cycles at 94 °C for 30 s, 50 °C for 30 s, and 72 °C for 1 min, and a final extension step at 72 °C for 15 min. Amplification success was confirmed by 1.5 % agarose electrophoresis using GoodViewTM nucleic acid stain (SBS Genetech Co., Ltd., Beijing). PCR products were purified using an enzymatic method with 0.8 U/µl of Exonuclease I and 0.2 U/µl of Thermosensitive Alkaline Phosphatase (FastAP; Thermo Fisher Scientific), following the manufacturer's instructions. Forward and reverse sequencing was conducted using the Macrogen Korea sequencing server. For each reference plant, a consensus sequence was obtained and used to build a local reference database of trnL (UAA) intron sequences.
Feces sampling. Samples were collected in two seasons: summer and winter. We performed six 100-meter transects across three different environments: native forests, pine and eucalyptus plantations, and grassland to reach 600 meters (Figure 1). Each transect was visited only one time per sample season to reduce the probability of resampling the same individual. Fecal sampling was performed opportunistically on a visual basis along the transects. We collected fecal pellets randomly chosen from each fresh fecal pile and preserved them in 70 % alcohol. Each sample was treated independently.
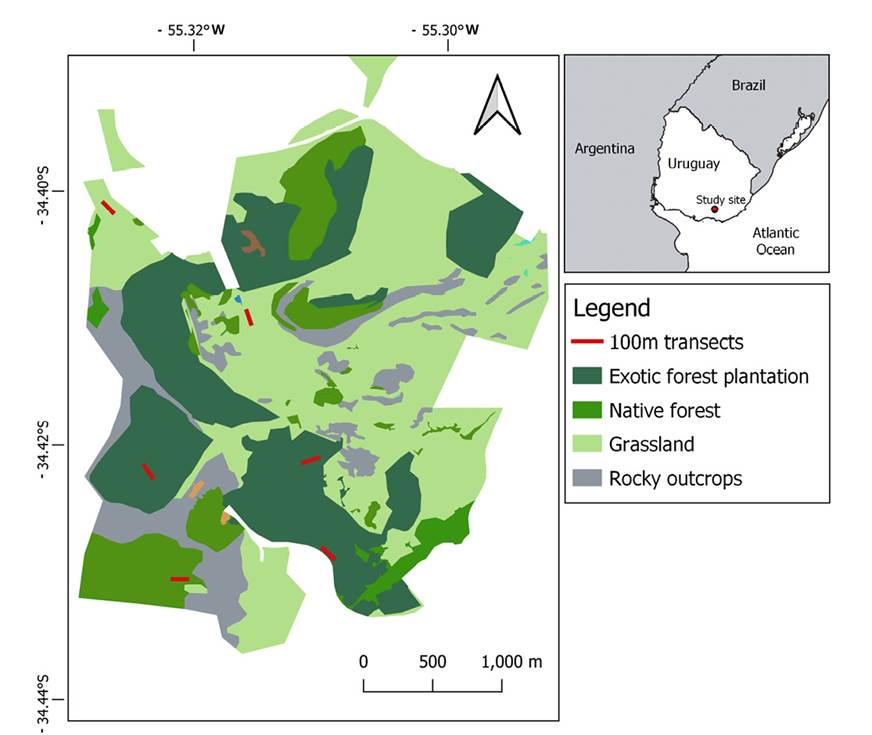
Figure 1 Habitat mosaics inside the 'Reserva Natural Salus' (RNS)'s. Red lines represent the location of each survey transect where gray brocket deer samples were collected. Right: Location of the RNS (red point) in Uruguay.
DNA extraction and PCR of the feces samples. DNA extraction was performed using the Stool Kit QIAGEN®. Sterile materials were used, and extraction was performed in an isolated room. Each sample was confirmed for species assignment (Aristimuño 2013) using a 159 pb fragment of the mitochondrial DNA D-loop region.
We performed PCR experiments to amplify a fragment of approximately 141 bp corresponding to a region of the intron of the gene trnL (UAA) of the chloroplast using the primers c (5’-CGAAATCGGTAGACGCTACG-3’) and h (5’-CCATTGAGTCTCTGCACCTATC-3’), as described by Taberlet et al. (2007). These primers had non-barcode and barcode adapter sequences for Ion Torrent-PGM® (Life Technologies). We used two barcode adapters (BC54 and BC57) on forward primers to distinguish two pooled samples based on the season sampling time. The PCR was done in a final volume of 10 µl, containing 1x buffer Invitrogen, 1.5 mM MgCl2, 0.2 of BSA, 0.05 mM of each dNTP, 0.5 pM of each primer, 0.25 U of DNA Taq Polymerase (Invitrogen), and 1 µl of DNA (Taberlet et al. 2007). The thermocycling conditions consisted of an initial denaturation at 94 °C for 3 min; 35 cycles of initial denaturation at 94 °C for 45 s, followed by 54.5 °C for 30 s and 72 °C for 60 s; and a final extension at 72 °C for 20 min.
PCR products were purified using the same protocol as the reference sequences and pooled for each season according to the protocols for NGS by Moreno et al. (2016). The mixture was quantified using a QubitR 2.0 Fluorometer (Life Technologies, USA) and diluted to obtain almost the same number of molecules per pooled PCR product. The run was performed using the Ion Torrent-PGM® (Life Technologies) platform from IIBCE facilities with a 316 chip. A 5000x coverage was achieved. The metadata are available in the NCBI BioSample database (http://www.ncbi.nlm.nih.gov/biosample/) under accession number: SAMN39224164 and SAMN39224165.
Bioinformatic Analyses. The software SEED2 (Větrovský et al. 2018) was used for amplicon high-throughput sequencing data analysis from FastQ files. The pipeline excluded short sequences (< 50 bp), low-quality sequences (mean quality value < 27), and chimeras. In addition, we performed clustering of similar sequences (97 %) into molecular taxa or Operational Taxonomic Units (OTUs) using the algorithm implemented in Vsearch (Rognes et al. 2016). We then used the G-Ins algorithm from SEED2 (Větrovský et al. 2018) to align sequences representing each cluster. The aligned sequences were cut to 148 bp, including gaps. Taxonomic assignment of the OTUs was carried out using the BLAST alignment software (Altschul et al. 1997). Clusters with fewer than 10 reads were not included in the analyses to prevent misidentification of plants owing to sequencing errors and to increase the reliability of species composition (Nikodemova et al. 2023). BLAST in SEED2 was executed twice, using: i) the local reference database constructed with trnL (UAA) intron sequences from the local species list, and ii) the global reference database from GenBank (Benson et al. 2013). Species assignment was based on the “Parque Salus” species list (Bonifacino and Rossado 2016) and records of local park rangers (pers. comm. 2022) and the best score on similarity and cover for both references used (local or global GenBank). Once the clusters were taxonomically assigned, the relative abundance was calculated by dividing reads for each taxon count by the total number of reads classified as species, genus, or family (frequency of each OTU). The importance of each item was evaluated by calculating the relative read abundance (RRA) of the items identified through DNA metabarcoding (Deagle et al. 2019; Tosa et al. 2023).
Results
Local reference database of trnL (UAA) intron sequences. The list of flora species from the “Reserva Natural Salus” (Bonifacino and Rossado 2016) included 488 species, 268 genera, and 80 families. In the global base of sequences GenBank (Benson et al. 2013), trnL (UAA) intron sequences were found for 226 (51.6 %) of the species listed for the Reserve. In addition, 211 (48.2 %) representative species of the same genera were retrieved from this reference base. The local reference database generated includes the recovered sequences from GenBank (Benson et al. 2013) and 138 sequences of native species or local varieties of exotic species, comprising 71 genera and 32 families obtained in this study and from the “CENUR-Rivera, Udelar” trnL (UAA) sequence collection. In total, 496 species from 302 genera and 85 families were included. The local base of reference was created with 1,050 sequences that represented all genera in the area and more than one specimen for some species.
Diet Analyses. Forty-eight feces were collected in summer and 51 in winter, however more than half did not provided good enough DNA quality required for PCR. From the DNA extraction of these samples, it was possible to confirm the identity of the deer species in nine and five samples collected in summer and winter respectively. For the summer sampling, three PCR products of the TrnL (UAA) region were obtained and pooled, generating 43,906 reads. After filtering for quality and size, 28,229 reads were assigned to 21 OTUs. For winter sampling (June), three samples were pooled, which generated 52,136 reads, reducing to 33,588 after filtering to obtain 16 OTUs. With the two pooled samples, we obtained a total of 25 OTUs belonging to 17 families. At the family level, five families were represented in one of the two samples by more than 3 % of the reads. The dominant families were Anacardiaceae and Rosaceae, representing 77 % of the reads obtained in each sampling season (Figure 2).
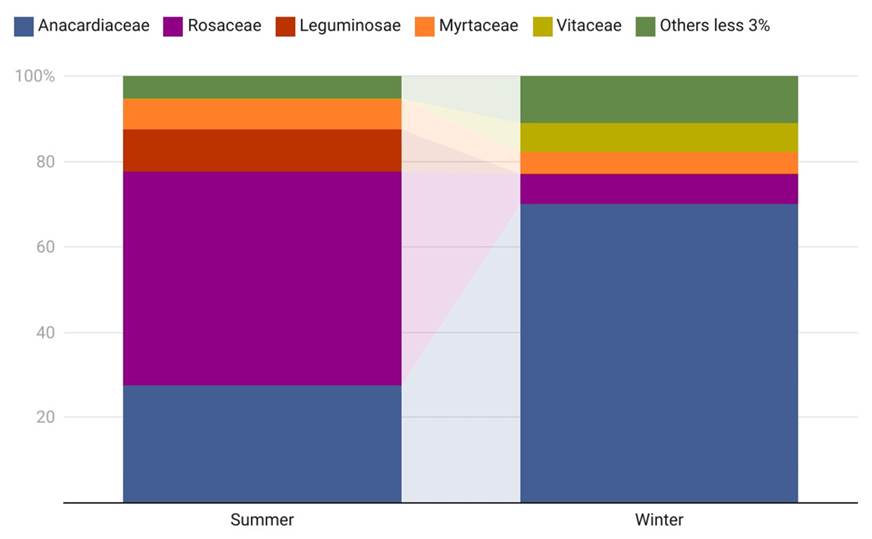
Figure 2 Frequency of reads for the most abundant families in feaces on summer and winter sampling dates.
We found that the amplified trnL (UAA) region does not exhibit sequence differences within certain species from the Anacardiaceae, Myrtaceae, Leguminosae, and Vitaceae families. Consequently, in these cases, taxonomic assignment was only possible at the genus or family level. To address this limitation, we conducted a broad comparison of all potential species assignable to an OTU with the list of flora species documented in the 'Reserva Natural Salus'. In certain instances, the flora of Parque Salus featured only a single species within a specific genus, allowing for the precise determination of the consumed species (e. g., Schinus engleri, Anacardiaceae). However, in other cases, particularly within the Myrtaceae and Leguminosae families, the study area harbored several species, preventing us from assigning OTUs to a particular genus or species within these families.
Taxonomic assignment showed two species as the more frequent items in the deer diet. For summer samples, Rubus ulmifolius (Rosaceae) represented 42.4 % of reads, followed by Schinus engleri, with 27.2 % of reads. In addition, Schinus engleri was the most abundant in winter, representing more than 68.7 % of the reads obtained. Other species present in the diet had a minor representation of less than 8 % in both samples (Table 1).
Table 1 List of species present on Subulus gouazoubira diet for summer and winter samples from “Reserva Natural Salus (RNS).” The genera with asterisks refer to the species not mentioned on the list but observed in home gardens from the area. Number of reads (N), relative percentage of reads (%).
| Summer | Winter | ||||||
|---|---|---|---|---|---|---|---|
| Order | Family | Gender | N | % | N | % | |
| Sapindales | Anacardiaceae | Schinus engleri | 7,677 | 27.20 | 23,073 | 68.69 | |
| Rosales | Rosaceae | Rubus ulmifolius | 11,964 | 42.38 | 2,259 | 6.73 | |
| Rosales | Rosaceae | Pyracantha coccinea | 2,159 | 7.65 | 83 | 0.25 | |
| Myrtales | Myrtaceae | Psidium sp./Myrcianthes sp./Eugenia sp.* | 1,914 | 6.78 | 1,706 | 5.08 | |
| Vitales | Vitaceae | Cissus striata | 38 | 0.13 | 2,179 | 6.49 | |
| Fabales | Leguminosae | Leguminosae Genera | 1,619 | 5.74 | 0.00 | 0.00 | |
| Species representing 1 - 3% | 1,867 | 6.60 | 1,973 | 5.90 | |||
| Species representing less than 0.99% | 991 | 3.50 | 2,315 | 6.90 | |||
| Total | 28,229 | 100 | 33,588 | 100 | |||
Discussion
Several studies have been conducted on the trophic aspects of Subulo gouazoubira, which vary in their methods and scope. They also focused on different aspects of feeding behavior, such as food selection, preferences, and dietary seasonality, among others. Most studies have focused on determining whether a species is frugivorous or consumes a higher proportion of leaves and grass (browsers). In Argentina, Paraguay, and the Bolivian Chaco, it has been observed that this species of deer feed on a wide variety of species, ranging between 25 and 100 species present in their diet, primarily consisting of leaves and fruits (Richard et al. 1995; Juliá and Richard 2001; Kufner et al. 2008, Serbent et al. 2011). The incidence of fruits and seeds varies in these regions, ranging from 11 % to 40 % of their diet; however, occasional consumption of specific fungi (Richard et al. 1995) and cacti, which can account for up to 30 % of their consumption, has also been observed (Cartes 1998; Caballero 2001). In Paraguay, the type of fruit (hard and dry or soft and fresh) depends on the season, which can be dry or wet (Stallings 1984). In contrast, in the Brazilian Pantanal, Pinder (1997) found a low incidence of fruits in the diet (with a peak of 19.2 % and annual average of 4.3 %). In the Peruvian Amazon rainforest, fruits constitute 80 % of the diet of the gray brocket deer (Bodmer 1989).
This study presents the first characterization of the diet of gray brocket deer through DNA barcoding, focusing on the trnL (UAA) intron region. This technique provides an approach to understanding the diversity of consumed species but does not provide information about the consumed structures (fruit or leaves). In turn, small sample sizes and pooled samples were employed. For a more accurate characterization of the gray brocket deer's diet, it is desirable to increase the sample size, use of replicates and analyze individual samples independently. This approach allows better estimations of taxonomic richness, to ensure that rare taxa are not overlooked (Alberdi et al. 2019) and an evaluation of individual variability, whereas, in our analysis sample discrimination is limited to season-based distinctions. The sequences obtained from the two samples included 25 species belonging to 17 families. Most of these species represented less than 10 % of the total reads obtained. Estimating abundance from sequence counts involves addressing biases caused by factors such as DNA copy number, sample DNA degradation, and variations in PCR extraction and amplification efficiency (Pompanon et al. 2012). Therefore, while some studies suggest a positive correlation between RRA and the percentage of plant mass consumed (Willerslev et al. 2014), it is essential to recognize that RRA serves as a semi-quantitative indicator of dietary composition (Deagle et al. 2019). However, considering the existence of different biases, the gray brocket deer's diet in the "Reserva Natural Salus (RNS)" was dominated by two species (Rubus ulmifolius and Schinus engleri). These results are consistent with those observed by Serbent et al. (2011), who identified 20 species in the gray brocket deer diet but only five represented 60 % of the diet. These authors found that deer consume a few preferred species, making a strong selection for those that contain higher-quality nutrients. This is supported by our results on the high consumption of blackberry (Rubus ulmifolius) in the summer when a large number of fruits are produced by this species. Studies of the cafeteria in this species under semi-captive conditions have evaluated their preferences among Morus nigra, Psychotria carthagenensis, and Rubus boliviensis (Richard and Fonturbel-Rada 2006). The authors found a clear preference for M. nigra and a rejection of R. boliviensis. These species differ in their nutritional characteristics, M. nigra has lower fiber content and higher crude protein content than R. boliviensis. In our study, the most represented species in summer, Rubus ulmifolius, has fiber values of 1.66 ± 0.06, even lower than M. nigra, and a higher carbohydrate content (Ahmad et al. 2015), which makes Rubus ulmifolius a high nutritional content food.
Small deer, less than 20 kg, such as the gray brocket deer, are generally classified as concentrated selectors or browsers (Hofmann and Stewart 1972; Hofmann 1989). Bodmer (1990) suggested the existence of a continuum between browsers and frugivores in which species can adjust their eating behavior in response to environmental changes. Our results agree with those of Richard and Fontúrbel-Rada (2006) and Serbent et al. (2011), which indicated that S. gouazobira is a selective and opportunistic species (Putman 1988), preferring a few species with high nutritional content. The dietary variations observed in the different habitats occupied by this species could be linked to the need to consume components with high energy content, as well as their plasticity to take advantage of young and highly productive ecosystems (Geist 1998; Serbent et al. 2011). This flexibility in their diet contributes to their success in different habitats and wide geographical distribution (Pinder 1997; Cartes 1998; Caballero 2001; Serbent et al. 2011).
It would be interesting to understand how the gray deer plays a role in the regeneration of native forests, as it is the only native deer species with a wide distribution in Uruguay. Understanding their role as dispersers or predators of invasive and native species seeds is crucial to accomplish this. This role is partly due to the size of the seeds; medium or large seeds appear to be destroyed during rumination (Bodmer 1989), whereas smaller seeds may remain intact when passed through the digestive tract (Gayot et al. 2004). In turn, the rumen bacterial communities (Bodmer 1989) of deer can digest different components of the seeds, softening the cuticles of the palms, but cannot degrade lignified covering present in the seeds of other species. It is necessary to carry out experiments on seed banks from the feces of S. goauzaoubira to determine the effect they can have on the germination rate of the different seeds they consume.
This is the first approximation of a diet determination technique in Subulo gouazoubira by metabarcoding. Our study focuses on evaluating the applicability of DNA barcode-based diet analysis of native herbivores with national capabilities. Although the number of samples was limited and the samples collected during the same sampling season (summer and winter) were processed together, relevant results could be obtained concerning the diet of this species in the "Reserva Natural Salus (RNS)" and the dominance of blackberries reads in the diet during the fruiting period of this species. Regarding the completeness of the reference base for barcode sequences of native species, it is necessary to advance into the generation of sequences of local populations and identify other markers, such as matK, trnH-psbA, rbcL, and ITS2 (Fazekas et al. 2012), which have better species-level discriminatory power for some families, such as Anacardiaceae, Myrtaceae, and Leguminosae. It is important to highlight that progress in this methodology is relevant, as the results of these types of studies are of great importance for species management plans, land management, and diversity conservation.











 nueva página del texto (beta)
nueva página del texto (beta)


