1. Introduction
Rock patina or rock varnish is a micrometric layer (50-200 μm thick) that forms on rock surfaces with a slow formation rate of around 1 to 40 µm every 1000 years (Liu and Broecker, 2000). Mineralogically, it is composed of almost 60% clay minerals, ~30% Fe- /Mn-oxides, and ~10% of other elements such as Si, Al, Mg, Ca, Ba, and Ce, among others (Potter and Rossman 1977; 1979; Dorn and Oberlander, 1981; Krinsley et al., 2012; Municchia et al., 2016; Lebedeva et al., 2019). Patinas are common in arid and desert environments (Potter and Rossman, 1977; Jones, 1991; Cremaschi, 1996; Lebedeva et al., 2019), although they can also be present in rocks of other ecosystem types (Krinsley et al., 2012; Lozano and Rossi, 2012; Goossens et al., 2015; Xu et al., 2018).
Desert Varnish can form because of either abiotic or biotic processes. The abiotic one involves varying cycles of deposition, accumulation, and enrichment of minerals with high amounts of manganese (Dickerson, 2011; Otter et al., 2020), weathering and diagenesis of aeolian dust and underlying rock (Municchia et al., 2016; Lebedeva et al., 2019; Andreae et al., 2020), and other physicochemical processes such as photooxidation, humidity, variation in pH and silica gelation (Perry et al., 2006; Dorn, 2007a; 2007b; Xu et al., 2019). On the other hand, biotic processes imply that varnishes are formed by biochemical processes mediated or induced by microorganisms (Krumbein and Jens, 1981; DiGregorio, 2005; Kuhlman et al., 2008), mainly those that mineralize Fe and Mn and deposit these oxidised and/or reduced elements, creating microlaminations.
Finally, some polygenic models imply a complex mixture between both processes (biotic and abiotic) where microorganisms (such as budding bacteria and fungi), physical-chemical and environmental processes all work together as an ecophysiological microsystem (Potter and Rossman, 1979; Dorn and Krinsley, 2011; 2019; Krinsley et al., 2017; Lingappa et al., 2021; Chaddha et al., 2024).
The presence of microorganisms in rock varnishes has been demonstrated in several studies using techniques such as fluorescence microscopy, electron microscopy, DNA sequencing, and culturing (Taylor-George et al., 1983; Lang-Yona et al., 2018; Esposito et al., 2019; Lingappa et al. 2021). These studies have revealed a diverse community of microorganisms, including bacteria, archaea, and fungi associated with the rock varnish (Esposito et al., 2015; Lang-Yona et al., 2018). These microorganisms are thought to play a role in the formation and alteration of the varnish by contributing to the accumulation of minerals and organic matter, such as varnish-associated photosynthetic bacteria, like Chroococcidiopsis sp., which accumulates higher concentrations of Mn in its cytoplasm (Lingappa et al., 2021). Also, Mn-oxidiser bacteria such as Hydrogenophaga sp. and Pedobacter sp., or in consortia, like Nevskia-Rhizobium spp., induce individual extracellular mineralization (Sjöberg et al., 2021).
Today, it is well known that the varnishes of the great deserts of America, such as the Sonoran and Mojave, have associated microorganisms (Taylor-George et al., 1983; Perry et al., 2003; Schelble et al., 2005; Kuhlman et al., 2006; Martínez-Pabello et al., 2021). Because of the extreme conditions in which the varnish is found, the inhabiting microorganisms can be considered extremophiles (exposed to high doses of radiation, extreme aridity, and temperature) (Coleine et al., 2021). Here, we present the first demonstration of morphological micro-biodiversity and the presence of Fe- and Mn-oxidising microorganisms in varnish samples from the Samalayuca Desert.
2. Study area
The Chihuahuan Desert is one of the largest desert areas in the American continent, with a quarter of its size in the US states of Arizona, New Mexico, and Texas, and the rest in Mexico states of Chihuahua, Coahuila, Nuevo León, Durango, and Zacatecas (Figure 1A-B). The municipality of Juárez is in the northern state of Chihuahua, Mexico in the middle of the Chihuahuan Desert (Figure 1A-B), where the locality Samalayuca stands out by its amazing biodiversity and landscape (Hernández et al., 2004). In this area, large amounts of rock varnishes can be found (Figures 1C and 1D), rich in Fe- /Mn-oxides (Martínez-Pabello et al., 2022) and “Los Médanos de Samalayuca”, which has been declared a Protected Natural Area, due to its landscapes, archaeological relevance, and biodiversity (Gatica-Colima et al., 2023). With an annual rainfall of 220 mm and a temperature variation of 23°C (CONANP, 2013), the region has been classified as a cold desert in the BWKx’ (e’) class of the Köppen’s climate classification (Peel et al., 2007). The current climate in the area (Schmidt, 1979) corresponds to an arid highland desert with hot summers, reaching temperatures of up to 41°C, and cold winters, with minimum temperatures dropping to -16°C.
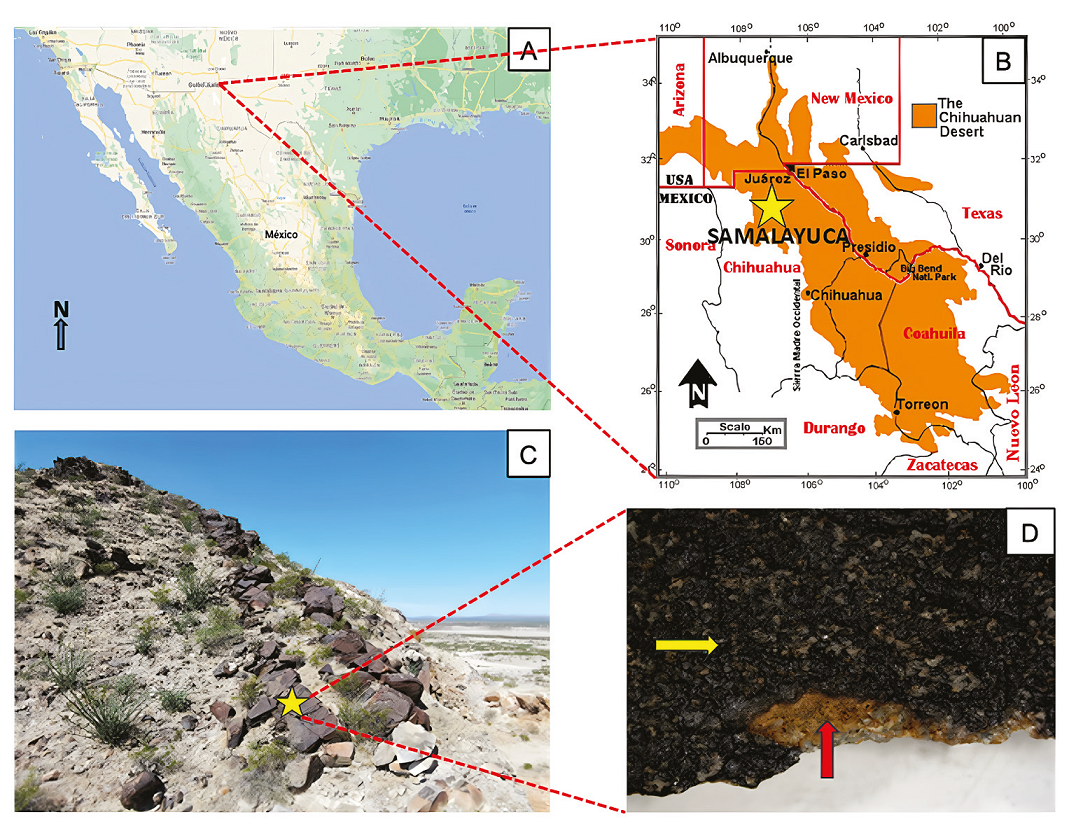
Figure 1 A and B. Location of the Chihuahuan Desert and Samalayuca collecting area (yellow star), Image taken and modified from Rivera-Rivera, 2006; C, Sampling site in Samalayuca. Prosopis glandulosa and Larrea tridentata are the dominant vegetation; D, Sample kind of collected varnish for this work. Dark varnish enriched with Mn-oxides (yellow arrow) and contact with underlying rock, sandstone (red arrow).
The annual average temperature ranges between 12-18°C. The prevailing winds, which make up 45% of the wind patterns, come from the west at speeds of approximately 20-40 km/h, although the fastest winds, reaching around 50 km/h, are from the southeast. More information on regional settings can be found in Martínez-Pabello et al., 2022.
3. Materials and methods
3.1. COLLECTION OF SAMPLES
Desert varnish was collected in the Sierra de Samalayuca, municipality of Juárez, northeast of the Chihuahuan desert, Mexico (31º 20’ 25.1124’’, 106º 30’ 21.0924’; Figure 1A to C). Sample selection was based on the appearance of rock varnish, where the samples with greater thickness, homogeneity, and dark pigmentation in the varnish were selected. The varnish was extracted with a cleaned geological hammer and stored in sterile conical Corning® tubes (50 mL) at 21ºC until use.
3.2. SAMPLE PREPARATION
Since the objective of this study was to provide an initial approach to isolating microorganisms from the varnish, one sample was selected for the analysis. The sample was prepared according to the modified protocol of Taylor-George et al. (1983). Briefly, 1 cm2 of varnish was separated from the source rock.
A manual grinding machine (Dremel ® 300) was adapted with a previously cleaned (hypochlorite-reagent grade) diamond tip to eliminate as much of the source rock as possible. Under sterile conditions, the varnish was placed in a Corning® conical tube with 40 mL of sodium phosphate buffer (NaCl 8.0 g/L, KCl 0.2 g/L, Na2HPO4 1.44 g/L, and KH2PO4 0.24 g/L; pH 7.4, J.T. Baker®), and vortexed for 10 min to remove traces of surface dust. Subsequently, the varnish was irradiated with UV-AB light for 30 min on each side of the varnish (face/back) to eliminate surficial microbial populations.
Finally, this fragment was pulverized in a porcelain mortar until fragments of ≈ 0.5 mm in length were obtained. The sample was separated into two fractions according to size: A) >0.5 and B) <0.5 mm. Fraction A was used for scanning electron microscopy (SEM), and fraction B was used for microbial cultures.
3.3. SEM
Fraction A consisted of 10 fragments (>0.5mm) that were analysed in a Zeiss Evo MA 10 microscope. Prior to observation, the samples were coated with carbon in a cathode sputter (Denton Vacuum® Desk V) for 10 min. Different areas of each fragment were scanned, looking for biological structures.
3.4. MICROBIAL CULTURES
Four culture solid media were used to cultivate the microorganisms: i) PDA, potato dextrose agar (39 g per litre, BD BioxonTM); ii) AA, water agar (8.5 g agar per litre, BD BioxonTM); iii) CZ (Czapek medium, 30 g sucrose, 2 g NaNO3, 1 g K2HPO4, 0.5 g MgSO4·7H2O, 0.5 g KCI, 0.01 g FeSO4, 8.5 g agar per litre, J.T. Baker®); and iv) TM (Thornton medium, MT, 1 g mannitol, 1 g K2HPO4, 0.5 g KNO3, 0.5 g asparagine, 0.2 g MgSO4·7H2O, 0.1 g CaCl2, 0.1 g NaCl, 0.002 g FeCl3, 8.5 g agar per litre, J.T. Baker®). All culture media were sterilised 15 min at 121°C, 15 psi.
20 mg of sterile fraction B were suspended in 1.5 mL of sterile water-peptone (0.01%) and vigorously shaken. 10 μL of the suspension was spread with a Drigalski spatula in triplicate in each medium. The cultures were incubated for 22 days at 21º ± 1ºC (room temperature seeking lichen growth), 25ºC (according to Wei (2020), most mesophilic microorganisms grow mostly at this temperature), and 37ºC (standard bacterial growth temperature).
3.5. SEQUENCING ANALYSIS
Two morphologically different microorganisms were chosen to sequence from the isolates: samples from the AA medium with a colony morphology like a bacterium and one from the PDA medium corresponding to a fungus (strain 5-1 and 13-3, respectively). The strains were identified by amplifying the gene 16S rRNA of 5-1 strain, using oligonucleotides 27F (5’-AGA GTT TGA TCC TGG CTC AG-3’) and 1492R (5’-GGT TAC CTT GTT ACG ACT T-3’) both at 0.3 μM (Lane, 1991) and according to the instructions of the GoTaq® G2 Flexi DNA Polymerase kit (Promega©). The reactions were run in the program: 5’ at 95°C; 35 cycles of 1’ at 95°C, 30’’ at 55°C and 1.5’ at 72°C; 10 min at 72°C and an indeterminate time of 4°C.
In the case of the 13-3 strain, we used the ITS molecular marker using the oligonucleotides, ITS1-F (5’-CTTGGTCATTTAGAGGAAGTAA-3’) and ITS4 (5’-TCCTCCGCTTATTGATATGC-3’), both at 0.3 μM (Gardes and Bruns, 1993). The reactions were run in the program: 5’ at 95°C; 35 cycles of 1’ at 95°C, 30’’ at 55°C and 45’’ at 72°C; 5 min at 72°C and an indeterminate time of 4°C.
A 1% agarose gel in 1X TAE (Tris, glacial acetic acid, and EDTA, Sigma-Aldrich®) was made to verify the amplifications and purification of fragments. The runs were at 100 volts for 40 minutes (ENDUROTM, LabNet© chamber). The gels were stained with ethidium bromide at 500 nM for 10 minutes. To identify the size of the amplicons, the 1kb Ladder (Promega©) was used and the purification of the amplicons was carried out with the gel extraction kit (QIAGEN©). The McLab company in San Francisco (USA) sequenced the fragments by the Sanger method with ABI 3730XL sequencers.
3.6. BIOINFORMATICS ANALYSIS
The sequences were cured using BioEdit© (Version 5.09). For bacteria identification, each sequence was subjected to two alignments with BLAST from NCBI©; the first excluded sequences of non-culturable bacteria; the second considered the genus with the highest percentage of identity and coverage from the first alignment was also excluded. Sequences of at least two strains of each species with similarity were downloaded. Finally, the ITS sequence was aligned in the UNITE 8.3 (Kõljalg et al., 2020) database for fungus identification.
For phylogenetic analysis, in both cases, alignments were done using MAFFT© (Version 7), applying the “Q-INS-i” strategy. A suitable evolutionary model was calculated for each group of sequences using JModelTest© (Version 2.1.10). Cladograms were created with Bayesian inference; for this, the MrBayes program (Version 3.2.7) was configured with the evolutionary models and applied at least 1,000,000 generations and sampling frequencies every 1,000 generations. Each generated file was edited with FigTree© (Version 1.4.4).
3.7. MICROSCOPIC ANALYSIS OF CULTURES
The primary cultures were observed at different incubation times in the Leica MZ® 12.5 with camera (DFC300) and the Zeiss® Axio Zoom V16 with camera Axiocam ICC5 microscopes, using transmitted light at 40X. For strain 5-1, the Gram test was performed using crystal violet (5% alcoholic solution, Hycel©), Lugol (5% in water, Hycel©), alcohol-acetone (Hycel©) in a proportion of 1:1, and safranin (1% in water, Hycel©). The preparation was observed under a microscope (Zeiss© PrimoStar) at 100X. The catalase and oxidase tests were performed on this strain, dripping 10 % H2O2 or BBL® Oxidase Reagent over isolated colonies.
3.8. BENZIDINE SPOT TEST
Reagent-grade benzidine (1,1’-biphenyl-4,4’-diamine, Sigma-Aldrich®) was used to prepare a 1% solution in acetic acid (7%). After 22 days of incubation, the primary cultures were flooded with 2 mL of the solution to determine the presence of Fe- and Mn-oxidising microorganisms. The colour changes from pale yellow to deep blue were considered positive results. Both chosen strains (5-1 and 13-3) were also tested.
4. Results and discussion
4.1. VARNISH SURFACE ANALYSIS
Ten fragments ≥ 0.5 mm were selected to be analysed by SEM to show the micromorphology of the microbiota present in varnish before pulverising. The varnish micrographs show microorganisms embedded in (Figure 2). We observed bacteria biofilm (Figure 2A) coexisting with fungus mycelium and spores (Figure 2B). The distribution of the different types of microorganisms is heterogeneous, frequently accumulated in small grooves, gaps, and microcavities on the surface of the varnish. These biological structures and distribution have also been observed on the surface of varnishes fromthe Sonoran Desert (Taylor-George et al., 1983; Martínez-Pabello et al., 2021) and the Mojave Desert (Perry et al., 2003; Kuhlman et al., 2006; Lebedeva et al., 2019), such as the brown filaments typical of Mn-oxidising bacteria described in New Mexico varnishes (Sjöberg et al., 2021).
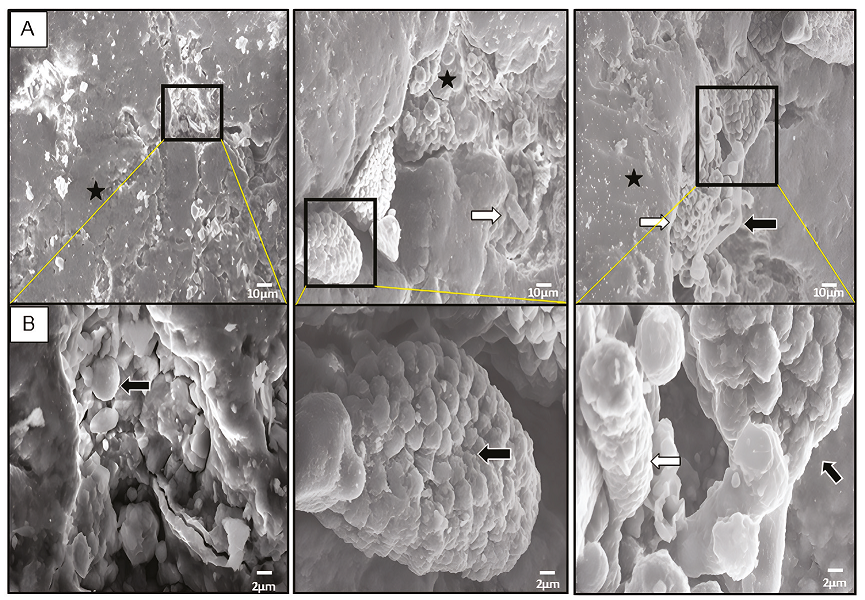
Figure 2 Analysis of varnish surface by SEM. A, Micrographs show many microorganisms morphologies embedded in the varnish such as: botryoidal biofilm (black star), ascomycetes colonies, hyphae and spores (black arrows), and tubular cells (white arrows). B, Magnification areas delimited by black rectangle (in A). Scale is indicated at bottom right by white bar (10 μm in A and 2 μm in B, respectively).
4.2. CULTIVABLE MICROBIAL VARNISH STRAINS
Viable colonies were recovered from the second fraction (B) of the same varnish that was cultured in four different media at three different temperatures (21º ± 1ºC, 25ºC, and 37ºC). After 22 days of incubation, bacterial colonies and mycelial morphologies were identified and photographed at different times (Figure 3): as time passed, the different morphologies appeared. Also, the cultures were incubated at room temperature (21° ± 1ºC) near a natural light source to promote the growth of photosynthetic microorganisms. However, it was impossible to ensure the isolation of microorganisms with this capacity. It has been reported that the growth of cyanobacteria associated with lichens is around 25 °C (Lange et al., 1998), which was the reason to incubate them at this temperature. Finally, greater growth of mesophilic bacteria was favored by culturing at 37°C.
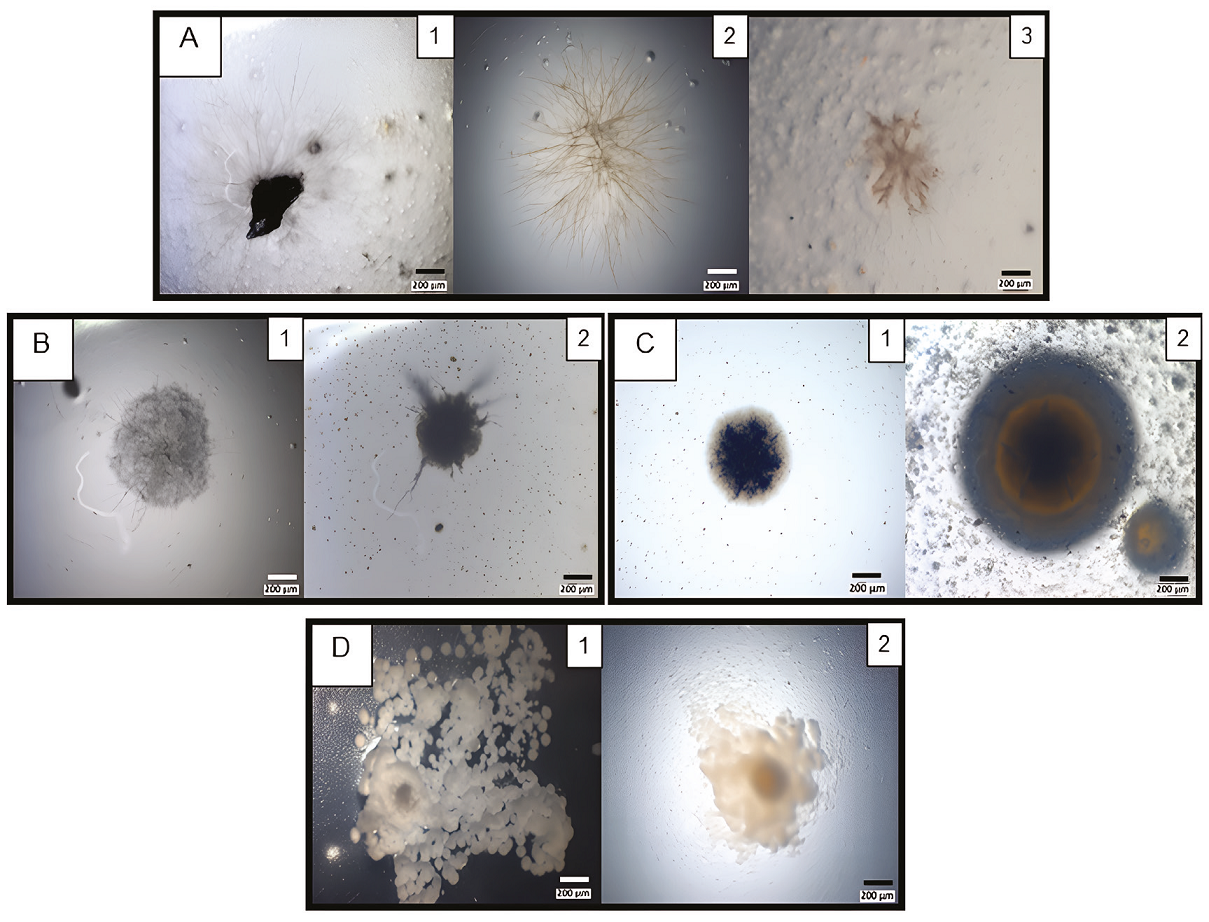
Figure 3 Strain colonial diversity recovered from pulverised varnish. A, Brown mucilage colonies with thin (A1-2) and astral mycelia (A3) morphologies. Micrographs (A1-3) correspond to AA Petri dishes incubated for 9, 10 or 8 days and, at 21º, 25º or 37ºC, respectively. B, Mycelium with sporangia morphologies. Micrographs (B1-2) correspond to AA Petri dishes incubated for 9 days at 21ºC. C, Rounded colonies. Micrographs correspond to CZ (C1), or TM (C2) Petri dishes incubated for 10 or 9 days and, at 25º or 21ºC, respectively. D, Umbonate colonies. Micrographs correspond to CZ Petri dishes incubated for 7 (D1) or 9 (D2) days and, 37º or 21ºC, respectively.
Slow-growing filamentous colonies were observed in all media and all incubation temperatures. Some of the colonies grew directly from millimetric varnish particles dispersed over the Petri dish (Figure 3A). The greatest number of microorganisms growths was found in the AA media (Figure 3A and 3B), followed by CZ (Figure 3C1 and 3D) and TM media (Figure 3C2). In the PDA media, we observed some colonies on the 9th day of incubation compared with the above other media, where colonies appeared at least after 14 days of incubation.
Fungal structures (spores and hyphae) were also present in all samples. The hyphae were composed of spherical or slightly elongated cells, with approximately 5-10 μm in diameter and thick melanized walls typical of black meristematic fungi. Some colonies resemble Mn-oxidiser fungi or bacteria, such as Cladosporium sp. or Pedobacter sp., respectively (Sjöberg et al., 2021). Microbial communities appear to be composed mainly of bacteria with few individual coccoid cells (Antonelli et al., 2020).
Although we observed differences in growth between the isolated bacteria and the fungus, their role in varnish colonization and/or mineralization processes is still indeterminate. Further studies are needed to understand and clarify their contributions.
4.3. MANGANESE AND IRON OXIDE PRODUCTION (REDOX)
The cultures were subjected to the benzidine spot test to determine the presence of Fe- and Mn-oxidising microorganisms. This test is based on the oxidation of benzidine in the presence of Fe3+ or Mn4+ to form a benzidine radical cation (deep blue; do Nascimento et al., 2006). It was confirmed that some of the obtained microorganisms were Fe- and Mn- oxidisers, such as the brown (Figure 4A) and black filamentary colonies (Figure 4B) that were coloured deep blue. Oxidising microorganisms were found mainly in AA and TM media, at 37ºC and 25ºC, respectively. On the one hand, Fe2+ and Mn2+ were dissolved in the TM medium, and oxidizer strains could directly mineralize them; conversely, in the AA medium, the dark hue comes from dispersed varnish particles.
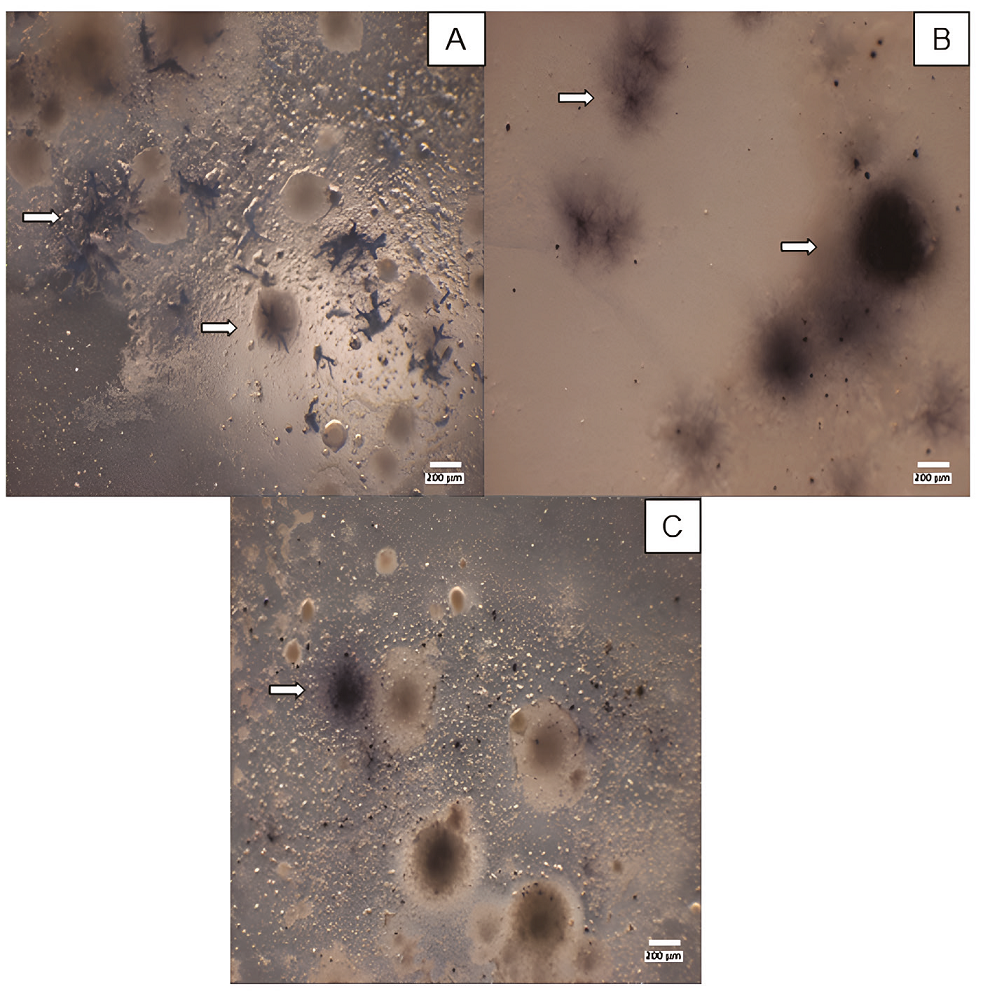
Figure 4 Revealing of oxidising microorganisms. A and B, AA medium culture incubation at 37ºC. C, TM medium culture incubation at 25ºC. All cultures were revealed in 22nd incubation day. Arrows highlight the positive presence of colonies with different morphologies.
Manganese and iron oxidation/reduction by microorganisms contribute to desert varnish´s complex mineralogy and geochemistry. The relative abundance and distribution of manganese and iron oxides within desert varnishes can provide valuable insights into the microbial communities and environmental conditions that have shaped their deposit over time.
Manganese and iron are two abundant elements in desert varnish, often occurring in the form of oxides. Microorganisms, particularly bacteria and cyanobacteria, play a significant role in the oxidation and reduction of these metals. Manganese-oxidising bacteria, such as Bacillus, Pseudomonas (Wright et al., 2018), and Arthrobacter (Liang et al., 2017) genders, can oxidise Mn2+ to Mn4+; these facilitate the oxidation of soluble Mn2+ ions to insoluble Mn4+ oxides resulting in their precipitation. Conversely, manganese-reducing bacteria, like Geobacter sp. (Mehta et al., 2005) and Shewanella sp. (Wright et al., 2016), and some fungi species can reduce Mn4+ to Mn2+, leading to the dissolution of manganese oxides (Wei et al., 2012). Similarly, iron-oxidising bacteria, such as Leptothrix sp. and Gallionella sp., oxidise Fe(II) to Fe(III), promoting the formation of iron oxides (Eggerichs et al., 2020), while iron-reducing bacteria, such as Geobacter sp. (Straub and Schink, 2004) and Clostridium sp. (Mishra and Pradhan, 2024), reduce Fe(III) to Fe(II), resulting in the dissolution or alteration of iron oxides.
4.4. STRAINS CHARACTERIZATION
After the 22nd incubation day of the prime culture and benzidine test, we picked two strains with a bacterial (Figure 5) and fungal colony morphology (Figure 6). Both were grown in their original isolated media, AA for 5-1 strain and PDA for 13-3 strain, at 37°C and 25°C for seven or 37 days, respectively. Both strains were positive for the benzidine spot test (data not shown). Also, they were molecularly identified by sequencing its 16S rRNA gene or the ITS region, respectively.
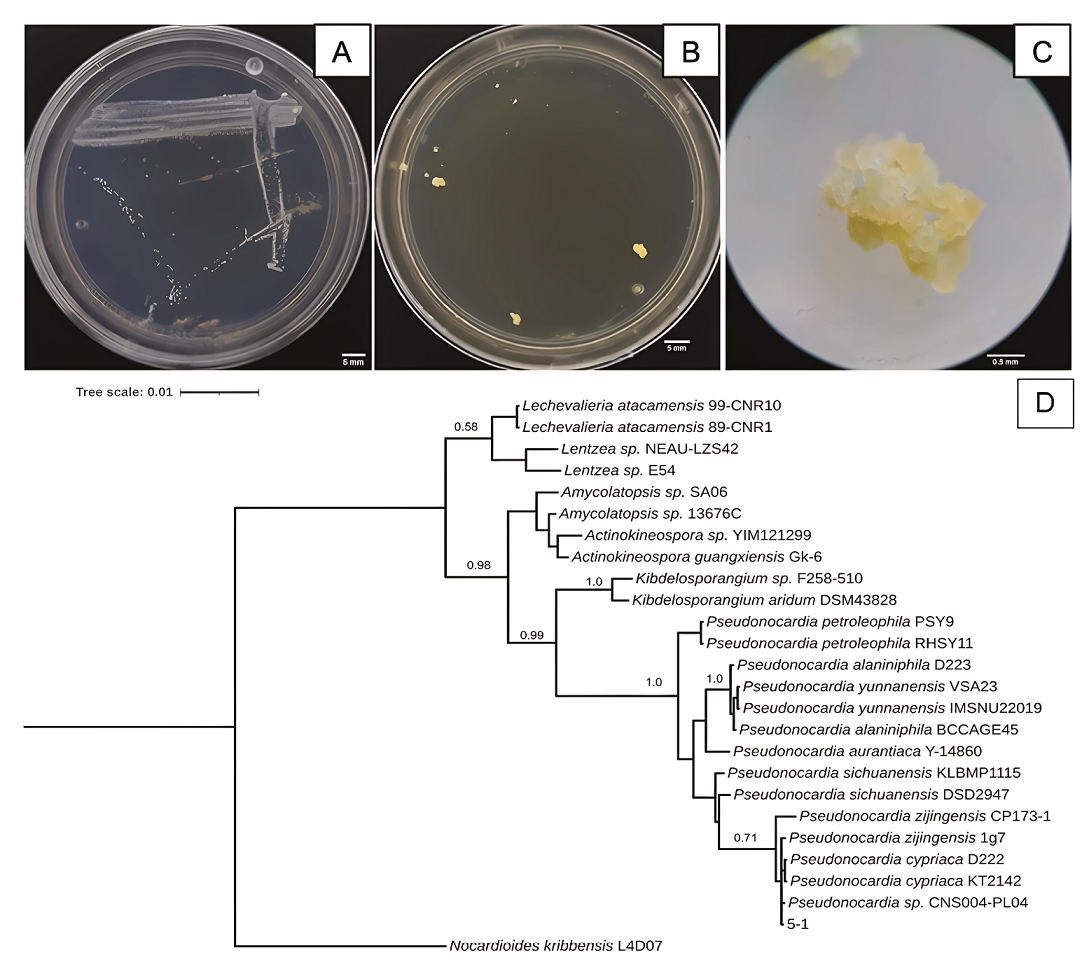
Figure 5 Characteristics of 5-1 isolated bacterium strain from Samalayuca desert varnish. A, Colony morphology in AA medium for seven days; B, or in NA medium for 14 days; both at 37°C. C, 5-1 colony micrograph in NA medium. D, Bayesian inference phylogenetic tree formed with rRNA 16S sequences. L4D07 Nocardioides kribbensis was the external group. The probability branch supports values are shown.
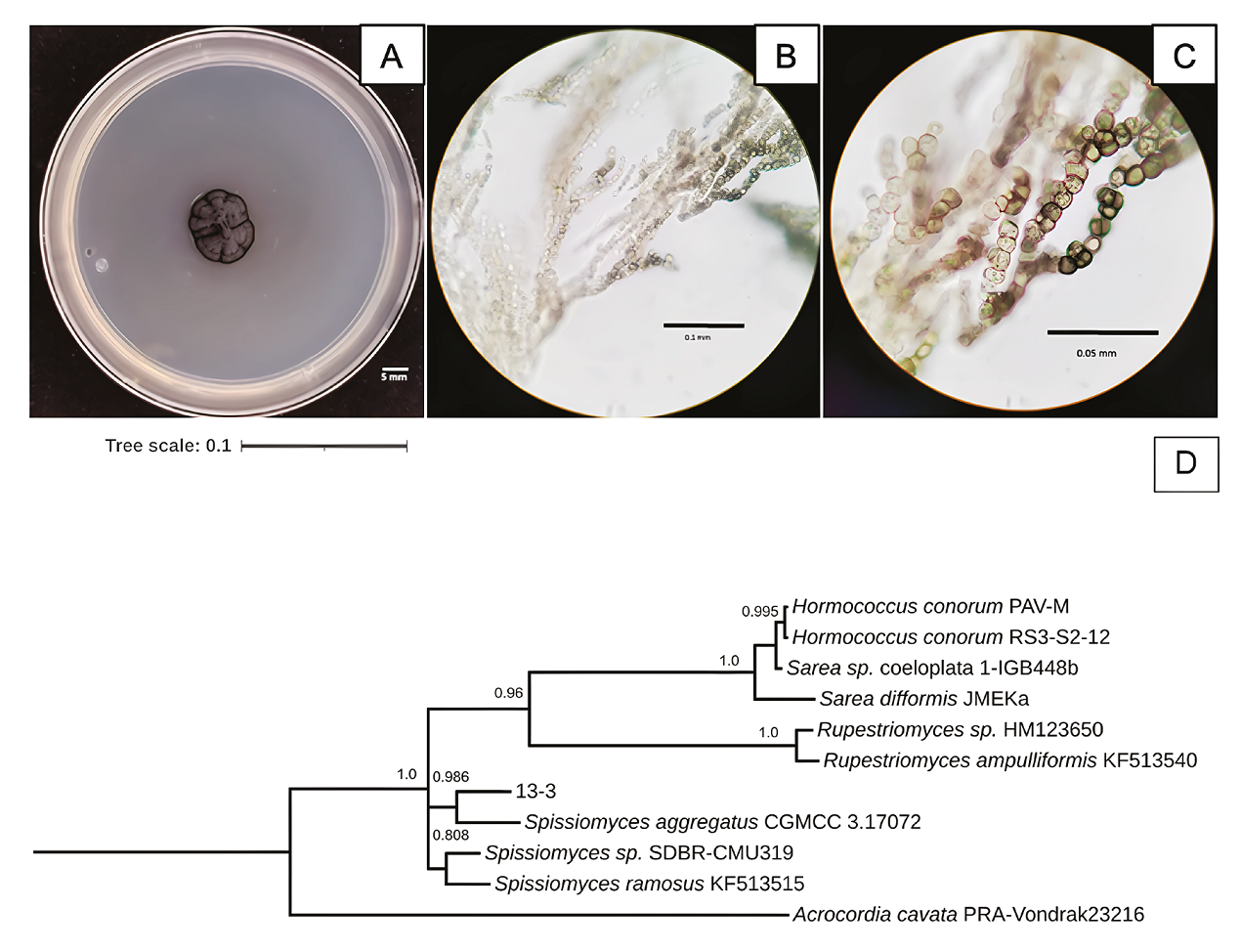
Figure 6 Characteristics of 13-3 isolated fungus strain from Samalayuca desert varnish. A, Colony morphology in PDA medium after 37 incubation days at 25°C; B and C, mycelium micrographs. D, Bayesian inference phylogenetic tree formed with ITS1 sequences. PRAVondrak23216 Acrocordia cavata was the external group. The probability branch supports values are shown.
The 5-1 strain shows white, opaque punctiform colonies in AA media (Figure 5A) on the seventh incubation day. In contrast, in Nutrient Agar (NA), the strain takes 14 days to form yellow botryoidal colonies (Figures 5B and 5C). The 5-1 strain phenotype is consistent with what is described for the gender as bacilli, catalase, and oxidase positive (Riahi et al., 2022), and botryoidal mycelium also appears yellow or yellow/brown on NA media (Thawai, 2018). However, the 5-1 strain was a Gram-negative bacillus, like P. carboxydivorans, which presents a substrate mycelium that strains Gram-positive, while the aerial mycelium and spores are Gram-negative (Tanvir et al., 2016). The 5-1 strain is related to Pseudonocardia sp. since it forms a single clade (Figure 5D). It is closer to P. cypriaca and the strain CNS004-PL04. However, the probabilistic value of the node is very low, which suggest that it could be a new strain or species. The Atacama Desert is one of theplaces where desert varnishes have been reported; P. nigra has previously been isolated in rocks from this same site (Trujillo et al., 2017); it has also been found that there are genes of the genus in biofilms that rest on sandstone rocks (Duan et al., 2021), the same type of rock on which varnishes grow in the Samalayuca mountain range.
The 13-3 strain was a slow-growing black fungus in PDA media (Figure 6A). It has branched mycelium with thick-walled, cylindric, and swollen cell hyphens (Figures 6B and 6C). Spissiomyces aggregates was its close relative (Figure 6D), with whom it shares the same morphological characteristics (Su et al., 2015). The species of this gender are melanin-producing fungi with a rock-inhabiting lifestyle (Ametrano et al., 2019), also reported as Mn-oxidisers (Chaput et al., 2015).
5. Conclusions
This is the first study that explores the microorganism diversity inhabiting the rock varnishes in the Chihuahuan Desert (Mexico). Varnishes of the Samalayuca desert have a rough surface that allows the colonization of various reproductive structures of different microorganisms (spores and conidia). As expected, the Samalayuca desert varnish harbors microbial communities (mainly bacteria and fungi) that oxidise metals such as Mn or Fe. Bacterial colonies mineralise Fe and Mn directly from the varnish in media with any supplementation (e.g., PDA or AA media). These bacteria have rapid growth (~7 days) compared to isolated black fungi (at least ~22 days) in the selection media; this may be due to the metabolism of each one and the conditions in which it was experimented. However, this can be evaluated in future works. The specific role of those isolated microorganisms in the formation of rock varnishes remains unknown. Still, the positive response of some of them to benzidine is a first step to understanding the complex ecological relationship of varnish microbiota.
Also, future studies on the molecular (metagenomics or metabarcoding analysis) and physiological characterization (i.e. mycelial morphology, secondary metabolites production in different culture media) of these isolates could clarify which species are involved with desert varnish formation and which are wind-borne environmental organisms.











 nueva página del texto (beta)
nueva página del texto (beta)


