INTRODUCTION
In the context of climate change and global environmental pollution, important efforts are being made to develop new sustainable and environmentally friendly processes (Kouzuma et al. 2018, Li et al. 2018). Microbial fuel cells (MFCs) have been shown to be an attractive alternative for environmental remediation due to their capability to generate electric energy while removing contaminants from wastewater (González-Nava et al. 2016, Li et al. 2018). Basically, an MFC is a bioelectrochemical system involving a modified anode and a cathode separated by a membrane. An electroactive biofilm grows on the surface of the anode in such a way that the electrons produced by the microorganisms during the biological degradation of the contaminants travel to the cathode through an external electrical circuit generating an electric current (ter Heijne et al. 2021).
The attractiveness of MFCs depends on their potential to complement the existing wastewater treatment processes with a technology that produces energy and could, therefore, be energetically self-sustainable (Guo et al. 2019, Katuri et al. 2019). Previous reports, for example, have estimated that the amount of energy contained in wastewater is over nine times larger than the amount of energy required for wastewater treatment (Toczyłowska-Mamińska et al. 2020). In this regard, MFCs have been successfully used to treat several municipal and industrial types of wastewater (Bose et al. 2019, Guo et al. 2019).
On the other hand, the microbial transfer of electrons to the anode substrate occurs by either direct or mediated mechanisms. In direct electron transfer, microbes require physical contact with the electrode (Khilyas et al. 2017), while in mediated electron transfer mechanisms, microorganisms produce or use soluble redox compounds to shuttle electrons between the microbial cell and the anode surface (Matsutani et al. 2014). The most common mechanism among these two is direct electron transfer (ter Heijne et al. 2021). Therefore, any disruption or reduced microbial attachment on the electrode surface limits the electron transfer kinetics between the microorganism and the anode, thus affecting the overall MFC performance and generating low power output values (Zhang et al. 2018, ter Heijne et al. 2021). On the other hand, a previous report has shown that organic contaminants used as a substrate determine the distribution of the bacterial community growing on the anode and affect MFC performance, particularly the generated power density and the Coulombic efficiency (ter Heijne et al. 2021).
To develop the electroactive microbial biofilm (EAB), the most popular strategy is using complex inoculums obtained from different sources, for instance, sediments, soils, or activated sludge (Zhang et al. 2018, Li et al. 2021). The selection and optimization of an appropriate consortium for a specific substrate is of paramount importance if the goal is to maximize energy production from wastewater (Kouzuma et al. 2018, Toczyłowska-Mamińska et al. 2020). Basically, the EAB can be developed by (1) acclimation of the inoculum and (2) transplantation of successive generations of EAB (ter Heijne et al. 2021).
In the context of the development of MFC technology, it is clear that microbial identification and assessment of the growing dynamics of microorganisms involved in the MFCs are mandatory to properly determine how the development of the microbial consortium correlates with electricity production in the MFC device (Kouzuma et al. 2018, Li et al. 2018, Toczyłowska-Mamińska et al. 2020). In this regard, metagenomics has become an important tool for studying population dynamics and the ecophysiology of microorganisms in natural ecosystems. Previous studies in MFCs have used 16 rRNA gene metabarcoding to characterize the bacteria associated with anodes and/or cathodes (Kiseleva et al. 2015, Kouzuma et al. 2018). Nowadays, microbial studies have identified the genera Shewanella, Geobacter, Clostridium, Proteus, Pseudomonas, and Rhodoferax as the microorganisms that can transfer electrons, and chromosomal libraries of these species have been developed (Li et al. 2021). Also, a previous study has shown that Firmicutes, Proteobacteria, and Actinobacteria were the main phyla that generate electricity in a MFC using wastewater as a substrate (Bose et al. 2019).
Metagenomics has also been used as a tool to analyze the functional genomes of biofilms. For example, Kiseleva et al. (2015) identified the microbial communities growing in the anodes at two different pilot-scale MFCs, Khilyas et al. (2017) identified the differences between microbial communities used as inoculum in two laboratory-scale MFCs, and Zhang et al. (2018) identified the composition and distribution of the functional genes involved in the accelerating extracellular electron transfer in MFC when rhamnolipids were added.
In spite of the use of metagenomics studies for the successful characterization of EAB and their relationship between population dynamics and different MFC performance criteria, there is not much information on the use of this tool for the characterization of microbiological metabolic functions of the microorganisms growing in the biofilm. Therefore, it is necessary to study the microbial communities and their microbial and metabolic functions, particularly those related to their electron transfer capacity.
In this context, this work aimed to determine the characteristics of an electroactive biofilm growing on the anode surface of a two-chamber type H microbial fuel cell, using the sequencing of its genomic material. Specifically, DNA analysis tools will allow to identification of population dynamics, taxonomy, and microbiological and metabolic functions that can be correlated with the bioelectrochemical performance of the biofilm in the cell. In this regard, this approach is a novel contribution to understanding the performance of an MFC operating under different experimental conditions and inoculums.
MATERIALS AND METHODS
Inoculum
The inoculum used in this experiment was obtained from the sediment of an extended aeration biological reactor used to treat wastewater at the Center for Research and Technological Development in Electrochemistry (CIDETEQ), located in Queretaro, Mexico (González-Nava et al. 2016).
Microbial fuel cell construction and operation
The experiments were carried out in an acrylic MFC H-type cell with two separate compartments bearing a proton exchange membrane (Nafion 117, DuPont) and 6 × 3 cm carbon cloths working as an anode and a cathode (Roe group). The anode compartment was constantly stirred, and the cathodic one was continuously aerated with a pump (Elite 800). All solutions in the anode chamber were bubbled with nitrogen gas for 15 min before use to displace air and oxygen (Logan 2005, Patil et al. 2011, González-Nava et al. 2016).
Biofilm formation
The anode chamber was fed with 150 mL of inoculum, 223 mL of a 10 mM sodium acetate solution (Richter et al. 2009, Patil et al. 2011), and 0.4 mL of each of the following nutrient solutions: 2.25% MgSO4·7H2O, 2.75% CaCl2 and 6.5% phosphate buffer pH 7.2 (8.5 g KH2PO4, 21.75 g K2HPO4, 33.4 g Na2HPO4·7H2O, and 1.7 g NH4Cl in approximately 500 mL of distilled water which was later diluted to 1 L, according to Bridgewater (2012) and González-Nava et al. (2016). All reagents were obtained from J. T. Baker.
Electrochemical analysis
The voltage (V) of the cells was measured every 60 min using an external resistance of 5 KΩ and a data acquisition system connected to a PC. The polarization curves were obtained by linear sweep voltammetry (LSV) at a scanning rate of 1 mV/s (Liu et al. 2012). The current and power densities were calculated using the area of the anode surface. The electrode potential was measured using a three-electrode system. Cyclic voltammetry (CV) was carried out in a potential window of -1.0 to 1.0 V at a scan rate of 20 mV/s. During these experiments, stirring and aeration were stopped to set diffusional control and to avoid noise in the electrochemical response. All experiments were performed with a BASi Epsilon-EC (Bioanalytical Systems) potentiostat and an Ag/AgCl reference electrode (0.209 V vs. NHE).
Metagenomic DNA extraction
To identify the microbial community in the biofilm electrodes, the growth over the anode was removed and homogenized in 10 mL of the analyte. The protocol established by the Zymo fungal/bacterial DNA kit (Zymo Research) was employed to obtain the total genomic DNA. To confirm the presence of DNA, 3 μL of DNA sample from each extraction protocol was were. The DNA sample was placed on a 1% prepared agarose gel, electrophoresed at 80 V for 60 min, and visualized using a UV light lamp.
Pyrosequencing and data preprocessing
The sequencing of the genomic DNA was performed using the Illumina HiSeq 2000 sequencing at the Macrogen Institute in Korea (Seoul, Republic of Korea). Data processing was performed mainly on the metagenomic analysis server (Meyer et al. 2008). The data were submitted for online annotation using the quality control (QC) pipeline. The reads that passed the QC pipeline were assigned to ribosomal RNA, annotated protein, and unknown protein. The percentage of reads with predicted proteins and RNA genes annotated to the indicated taxonomic distribution when the microbial biofilm was growing on the anode surface.
Taxonomic profiles were calculated by Best Hit classification at the E-value cutoffs of 10-5 with a minimum alignment length of 50 pb on all the annotation source databases used by MG-RAST (metagenomics rapid annotation using subsystems technology). The distribution of taxonomic domains, phyla, orders, families, and genera for the annotation was analyzed in detail. Functional profiles were identified using the SEED subsystems annotation source of the MG-RAST with a maximum e-value of 1e-5, a minimum identity of 60%, and a minimum alignment length of 15 measured in amino acids for protein. The gene profile characteristic was determined using the databases of clusters of orthologous groups of proteins (COG) and the Kyoto Encyclopedia of Genes and Genomes (KEGG).
RESULTS AND DISCUSSION
Electrochemical biofilm performance
The electrochemical performance of the cell was followed for 28 days, showing an adaptation phase during the first five days, where the MFC does not generate significant voltage (Fig. 1a). Then, four sequential cycles of voltage generation were observed reaching values near 0.4 V in the first cycle and later an average near 0.35 V in subsequent cycles. In previous reports, Chen et al. (2021) reported maximum values of 0.596 and 0.627 V when an MFC with two chambers was fed glucose and sodium citrate. Also, Sciarria et al. (2019) obtained voltage values of 0.6 V on average using a single chamber MFC and sodium acetate as a carbon source.

Fig. 1 Electroactive potential of the biofilm growth in a type-H microbial fuel cell. (a) Generated voltage after inoculation; (b) cyclic voltammetry, bare electrode (gray) and electrode with biofilm (black); (c) polarization and power curves.
Cyclic voltammetry experiments were carried out to study the performance of the cells that formed biofilms. As shown in figure 1b, the electrochemical performance of the bioanodes shows wide and not well-defined peaks such as those obtained in previous works (Silveira and Schneedorf 2018, Bhowmick et al. 2020). The electrochemical signals that can be observed at potentials close to 1 V are probably related to the oxidation of compounds positioned on the surface of the bacteria.
On the other hand, the polarization and power curves showed an open circuit potential of 0.56 V, a maximum current of 41.4 mA/m2, and a maximum power density of 8.1 mW/m2 (Fig. 1c). The anodic potential corresponded to -0.483 V vs. Ag/AgCl, a characteristic value for the exoelectrogenic activity of the microorganisms (Silveira and Schneedorf 2018). The cathodic potential was 0.075 V vs. Ag/AgCl, which favored the potential difference between the anode and the cathode for the operation of the cells (Sciarria et al. 2019, Chen et al. 2021, Tholia et al. 2021).
Taxonomic composition
At the end of the last cycle of operation of the MFC, the composition of the microbial community in the biofilm was analyzed. The rRNA sequences obtained by DNA pyrosequencing analysis showed an average length of 707 pbs. After quality checking obtained by MG-RAST (ID mgp96078), 94 117 sequences (90.92%) were accepted and grouped in ribosomal RNA genes (0.25%), sequences predicted proteins with known (77.66%) and unknown functions (22.09%).
Taxonomic analysis of the biofilm in the anode also showed that 98.26% of all reads were assigned to the kingdom bacteria, identifying 84 bacterial orders, of which Rhodobacterales, Burkholderiales, Pseudomonadales, Rhizobiales, Flavobacteriales, Nitrosomonadales, Rhodospirillales, Rhodocyclales, Alteromonadales, and Verrucomicrobiales were the most abundant (Fig. 2a). These results are consistent with those obtained by Wang et al. (2013), who identified electroactive capacity in Rhizobiales and Burkholderiales. In addition, Burkholderiales has been identified as bacteria with the capability to metabolize oils, fats, and hydrocarbons.

Fig. 2 Bacterial community composition involved in the biofilm on the anode surface analyzed by pyrosequencing and identified by metagenomics rapid annotation using subsystems technology. Top 10 of the most abundant: (a) orders, (b) families, and (c) species.
Related to family categories, the dominant family was Pseudomonadaceae, with a relative abundance of 19.9%, followed by Rhodobacteraceae (13.1%) and Comamonadaceae (12.3%), as seen in Figure 2b. This information is consistent with previous works that identified Pseudomonas as an electroactive microorganism commonly found in MFCs (Almatouq et al. 2020, Chen et al. 2021). Finally, the top ten species found in the taxonomic analysis of the electroactive biofilm in figure 2c show that the most abundant species is Hyphomonas neptunium (16.0%), followed by Opitutus terrae (14.5%), and Azotobacter vinelandii (12.8%). In a previous work, Khilyas et al. (2017) also reported the presence of Azotobacter genus on anodes in the MFCs, particularly A. vinelandii, a species identified as one of the predominant microorganisms in this study.
Functional categories description
To assess the functional profile of the electroactive biofilm on the MFC, the total reads were annotated according to categories of the Clusters of Orthologous Genes (COG) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) databases. The COG database (Fig. 3a) showed that 47% of the total reads were related to metabolism, and 22% were assigned to cellular processes and signaling. In contrast, about 18% corresponded to housekeeping genes involved in information storage and processing of the EAB. Figure 3b shows the different subcategories of the EAB based on the COG database. In the category of metabolism, the most abundant metabolic type was amino acid transport and metabolism (10.06%), followed by energy production and conversion (9.36%), as well as inorganic ion transport and metabolism (7.18%). Also, the presence of carbohydrate transport and metabolism (6.07%), lipid transport and metabolism (4.40%), and coenzyme transport and metabolism (4.24%) were detected in the metabolism category.
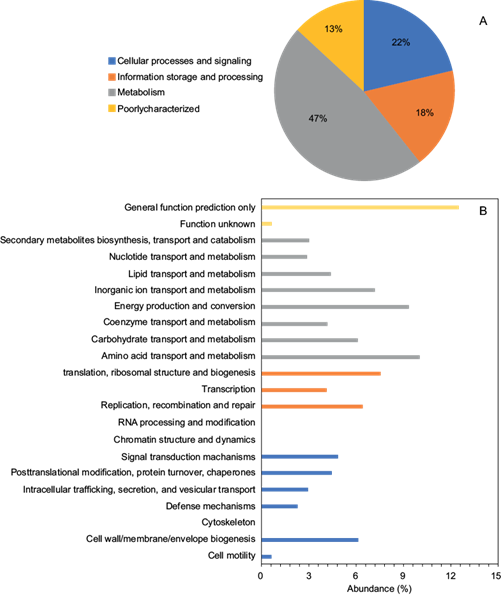
Fig. 3 Functional categories of an anodic electroactive biofilm based on the Clusters of Orthologous Genes (COG) proteins database. (a) Categories and (b) subcategories.
According to these observations, it is evident that besides the capacity to metabolize amino acids the biofilm exhibits the genetic machinery to break down carbohydrates and lipids. Particularly, there are abundant reads matching the genes for nucleoside-diphosphate-sugar epimerases, glycosidases, and dihydroxy acid dehydratase/phosphogluconate dehydratase (results not shown). The epimerases detected are enzymes that catalyze the inversion of stereochemistry in biological molecules (Tanner 2002), and glycosidases are the enzymes responsible for hydrolyzing glycosidic linkages. This is particularly important because they are involved in the carbohydrate digestion of oligo- and polysaccharides, carbohydrates, chains of glycoproteins, and glycolipids (Nishimura 1992). On the other hand, phosphogluconate dehydratase is an enzyme that participates in the Entner-Doudoroff pathway so that glucose is catabolized into pyruvate (Meloche and Wood 1964).
The abundant reads obtained from metagenomics sequences match the genes for inorganic ion transport and metabolism, as shown in figure 3b, and according to the COG database, they could be involved in “Outer membrane receptor proteins, mostly Fe transport”, “Putative silver efflux pump”, and “Cation transport ATPase” (results not shown). Fe transport is a mechanism involved in the growth of dissimilatory metal-reducing microorganisms when metal reduction is stimulated by adding a suitable electron donor to enhance electron transfer (Kondo et al. 2015, Li et al. 2021). The P-type ATPases comprise a large superfamily of cation transporters that utilize the energy of ATP to transport ions against their electrochemical gradients across cell membranes (Fambrough and Inesi 1996). The abundance of the sequences related to energy production and conversion in figure 3b, could be considered indicative of the biofilm capacity to generate and transfer electrons, which is an important characteristic of microorganisms involved in MFCs.
Electron reactions and cell wall and capsule formation
Figure 4 shows the analysis of the abundance of reads related to the electron-accepting and donating reactions based on the annotation of functional genes using SEED subsystems in MG-RAST, the level 2 subsystem of respiration. Figure 4a shows that the most abundant electron-accepting reaction is related to terminal cytochrome C oxidases (44.5%), followed by anaerobic respiration reductases (18.9%) and ubiquinone menaquinone cytochrome c reductase complexes (16.6%). These proportions make sense since the anode chamber of MFC resembles anaerobic conditions when the substrate is converted into metabolic intermediates by bacteria through redox reactions and enzyme activities. Some redox-active proteins involved in the electroactive biofilm are the c-type cytochromes (Kotloski and Gralnick 2013). The metagenomic analysis revealed the presence of many cytochromes’ oxidases, such as a terminal cytochrome oxidase and terminal cytochrome d and O ubiquinol oxidases (Fig. 4a), which could be related to the fact that the c-type cytochromes are present on the outer cell membranes of electroactive microorganisms (Kotloski and Gralnick 2013).
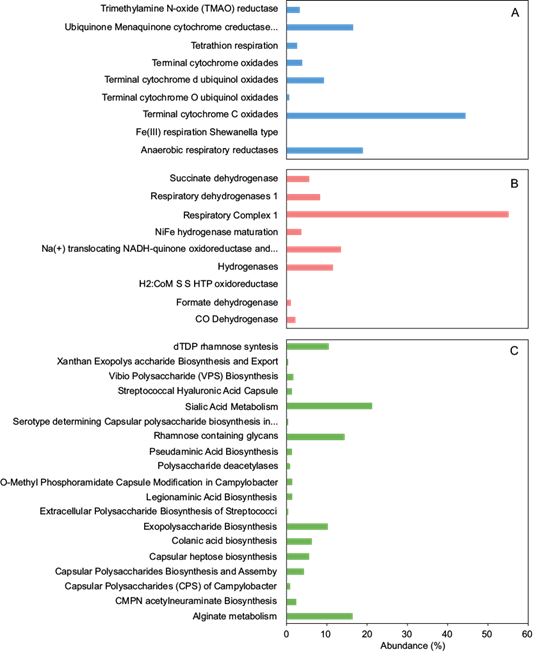
Fig. 4 Abundance of subsystems in an electroactive biofilm from level 1 subsystem of respiration based on the SEED database. (a) Electron-accepting reactions, (b) donation reactions, and (c) cell wall and capsule.
On the other hand, the larger relative abundance of the sequences corresponds to the respiratory complex I (55.0%), which, according to the SEED database, could be involved in the NADH-ubiquinone oxidoreductase chain function (Fig. 4b). This enzyme complex is characteristic of electroactive microorganisms. In fact, respiratory complex I is the first large protein complex of the respiratory chains of many organisms, such as bacteria (Weiss et al. 1991, Strickland et al. 2014). Also, it is important to note that electrons are transferred from NADH to O2 by complex I or NADH dehydrogenase. Na+-pumping NADH:ubiquinone oxidoreductase (Na+- NQR) is a respiratory enzyme reported in bacteria such as Haemophilus influenzae and Vibrio cholerae. Na+-NQR obtains energy by oxidizing NADH and reducing ubiquinone, which results in a stable sodium gradient across the inner bacterial membrane (Strickland et al. 2014).
In figure 4b it is also possible to identify the presence of dehydrogenase complexes (succinate dehydrogenase, formate dehydrogenase, and CO dehydrogenase), which could be related to the electroactive activity of the biofilm on the anode. Previous reports indicate that two enzymes (phosphatase and dehydrogenase) are responsible for substrate degradation and redox reactions. Phosphatase deactivates the substrate by removing the PO3- group and prevents its conversion into products. Dehydrogenase catalyzes the redox reactions involving mediators and enhances the release of H+ and e-, which are essential for electricity generation (Reddy et al. 2010). The presence of succinate dehydrogenase is also important because this enzyme has a central role in the respiratory chain and is a component of both the TCA cycle and the electron transport chain. Additionally, this enzyme catalyzes the oxidation of succinate to fumarate as a membrane-bound component of the respiratory chain and transfers electrons to ubiquinone without pumping protons across the bacterial membrane, indicating that the biofilm could possess electro-active properties (Yu et al. 2019). Formate dehydrogenase, on the other hand, was detected with a lower relative abundance (0.9%); however, this enzyme was previously detected in Escherichia coli, a microorganism that has shown electroactive properties (Choi et al. 2018).
Identifying the cell wall and capsule components in biofilms is important because biofilms are cellular aggregations surrounded by self-produced polymeric substances and exist as mono and poly-species cellular communities (Moradali and Rehm 2019). To evaluate the genetic capacity of the microbial consortium to develop a biofilm over the anode surface, a level two subsystem of the cell wall and capsule based on SEED database analysis was performed. As shown in figure 4c, experimental results indicate that the most abundant of reads codified to sialic acid metabolism (21.19%), followed by alginate metabolism (16.38%), rhamnose containing glycans (14.26%), dTDP rhamnose synthesis (10.40%), and exopolysaccharides biosynthesis (10%). In this way, the presence of genes involved in rhamnose synthesis (dTDP rhamnose synthesis and rhamnose containing glycans) suggests that the biofilm formed over the anode is composed of rhamnose, alginate, and sialic acid molecules.
It is important to mention that while sialic acids play an important role in the stabilization of molecules and membranes, as well as in modulating interactions with the environment (Lindberg et al. 2001), alginates lead to the formation of hydrogels and cross-linked polymeric scaffolds. Consistent with the data in figure 2, alginate biosynthesis has been previously identified in P. aeruginosa and, to a lesser extent, in A. vinelandii (Moradali and Rehm 2019).
CONCLUSIONS
The microbial fuel cell implemented in the laboratory showed the typical electrochemical performance of these devices as far as voltage generation and power density. Taxonomic analysis of the sequences obtained after DNA sequencing of the biofilm revealed that the main orders found in the electroactive biofilm were Pseudomonas, Nitrosomonas, Hyphonas, and Opitutus. The metagenomic analysis of the sequences also indicates that electroactive microorganisms involved in electricity production have the codified genes to metabolize amino acids, lipids, and carbohydrates. This analysis also suggests that the biofilm has the machinery to promote ion transport and energy production/conversion. The presence of many cytochromes’ oxidases, such as terminal cytochrome oxidase, terminal cytochrome d, and O ubiquinol, could be considered a confirmation of the capacity of this biofilm to generate and transfer electrons. On the other hand, the synthesis of rhamnose, sialic acid, and alginate molecules is related to the formation of the microbial biofilm capable of growing in the anode. All this information provides a general characterization of an electroactive biofilm that can be used to support further studies aimed at reaching a deeper understanding of the individual aspects of the mechanisms that explain and eventually lead to the optimization of biofilm dynamics in MFC energy conversion performance.











 nueva página del texto (beta)
nueva página del texto (beta)


