Notas Fitopatológicas
Genetic diversity of Fusarium oxysporum associated to the banana Fusarium wilt
-
Publication dates-
October 13, 2025
Sep-Dec , 2025
- Article in PDF
- Article in XML
- Automatic translation
- Send this article by e-mail
- Share this article +
ABSTRACT
Background/Objective.
The research objective was to analyze Fusarium oxysporum
(Fo) isolates from different banana regions from México using SSR markers.
Materials and Methods.
Pseudostems of Musa plants exhibiting wilt-disease were sampled from ‘Pisang Awak’ (ABB), ‘Silk’ (AAB), and ‘Bluggoe’ (ABB) varieties collected on commercial orchards of five banana-producing areas of México: Colima, Michoacán, Jalisco, Nayarit, and Yucatán states. Six SSR loci were analyzed for population structure based on 96 Fusarium oxysporum isolates. The number and frequency of haplotypes were calculated in POPGENE. With this data, using the dissimilarity index of Morisita in the R package vegan, the genetic similarity of the populations was estimated. With the haplotype data, we also conducted a molecular analysis of variance (AMOVA) to evaluate the genetic differentiation between regions (FCT), among populations in each region (FSC), and among all populations included in the analysis (FST). Additionally, to illustrate the relations among haplotypes, and their distribution and frequency among sampled populations we generate a minimum spanning network.
Results.
After seven days of PDA culture, all isolates had cottony white mycelia, which can be flat to aerial, and can be colored on the upper surfaces from light violet to dark violet. But on the bottom surfaces, the colony can have colorations from orange to dark red. We observed short monophialides, unicellular or bicellular with oval microconidia. Macroconidia were falciform, and with three to four septa. Putatively, these characteristics fit the F. oxysporum taxonomic criteria. All tested loci were polymorphic across the 96 Foc. We detect 16 haplotypes, of which the most frequent and common was the H16; the others were rare and could be widely dispersed or restricted to one or two populations. Also, we did not find genetic differences among the three analyzed populations, as well as we did not find genetic structure or differentiation at any level. These findings are related to Foc isolates as a complex not necessarily to those related to Foc races.
Conclusion
. Our analyses did not detect significant genetic differences between the Fo populations studied, nor did we find genetic structure or differentiation at any level. Therefore, our data suggests that the analyzed populations constitute a single metapopulation, despite the presence of infrequent or unique haplotypes.
Keywords::
Foc R4T, Banana genotypes, SSR markers
Introduction
Bananas and plantains (Musa spp.) are the fourth most important crop in the world, surpassed only by rice, wheat, and corn (FAOSTAT, 2024). This fruit is available throughout the year and is grown in 135 countries in the tropical and subtropical regions. Although banana represents a significant source of income in the Latin American region, only 15% of the production is exported as a commodity since most harvest is in small farms or backyards, for domestic consumption or local markets (FAOSTAT, 2024).
Bananas and plantains are affected by several pathogens worldwide, however, Fusarium wilt, which is caused by a complex of isolates of the fungus Fusarium oxysporum (Fo). The specific complex of F. oxysporum f. sp. cubense (Foc), constituted for at least four races, is considered the most destructive diseases for this crop (van Westerhoven et al., 2022). Foc produces three types of asexual spores: macroconidia, microconidia, and chlamydospores, which have roles in dispersal, reproduction, and survival (Pegg et al., 2019). Once Foc invades a plantation, it is hard to control, because the pathogen persists in the soil for long periods (Maryani et al., 2019). Thus, the most effective alternative to control the disease is the replacement of susceptible cultivars with others resistant to the pathogen.
-
van Westerhoven et al.,
2022Uncontained spread of Fusarium wilt of banana threatens African food securityPLOS Pathogens, 2022
-
Pegg et al.,
2019The Epidemiology of Fusarium wilt of bananaFrontiers in Plant Science, 2019
-
Maryani et al., 2019Phylogeny and genetic diversity of the bananaFusarium wilt pathogen Fusarium oxysporum f. sp. cubense in the Indonesian centre of originStudies in Mycology, 2019
In the past, nearly all commercial plantations in the world changed their susceptible varieties by Cavendish subgroup genotypes (Genome AAA). The recent emergence of Foc Tropical race 4 (Foc R4T in Spanish), which is capable to infect members of the Cavendish subgroup, has become the main threat to the banana and plantains industry. This is more problematic because currently, the markets demand primarily fruits of the ‘Giant Naine’ clone from the Cavendish subgroup (Genome AAA), and there are no other resistant banana genotypes available (Ordoñez, 2018). In México, nearly forty thousand hectares of ‘Gros Michel’ cultivar was devastated in the mid-1950s due to Foc race 1, forcing its replacement by resistant clones of the ʻCavendishʼ subgroup, such as ʻGrand Naineʼ. For example, a decade ago, the state of Nayarit had 3,000 ha of ʻSilkʼ and 3,500 ha of ʻGros Michelʼ cultivars. Recently, banana and plantain production in México has been concentrated in 16 states, with Chiapas, Tabasco, Colima, and Veracruz state being the main producers, accounting for 72.5% of the planted area, 75.3% of the volume produced, and 66.3% of the value of the crop’s production. The volume obtained in 2023 was 2,642,338 tons, 1.9% higher than the previous year’s production (SIAP, 2024).
-
Ordoñez, 2018A global geneticdiversity analysis of Fusarium oxysporum f. sp. cubense, 2018
Knowledge of the genetic diversity of populations of phytopathogenic fungi and their mode of reproduction is relevant for crop management programs, and to reduce the impact of the disease (McDonald and Linde, 2002). In the case of Foc, it has a relatively diverse genetic structure for an asexual reproductive fungus (Martínez-de la Parte et al., 2024). The genetic information of the pathogen can be used to design effective management practices for the disease, as well as for breeding programs to develop a resistant or tolerant banana cultivar, and for quarantine purposes in a particular country or geographical location (Magdama et al., 2020; van Westerhoven et al., 2022; Chittarath et al., 2022; Baruah et al., 2025). In México, despite the sustained increase in the number of plantations of banana and plantain, information regarding the diversity and genetic structure of Foc is scarce. This reserach present the first analysis that studies Fo isolates from different regions using SSR markers as a preliminary approximation to justify deep Foc diversity studies.
-
McDonald and
Linde, 2002The population genetics of plant pathogens and breeding strategies for durable resistanceEuphytica, 2002
-
Martínez-de la Parte et al., 2024Genetic diversity of the banana Fusarium wilt pathogen in Cuba and across Latin America and the CaribbeanEnvironmental Microbiology, 2024
-
Magdama et
al., 2020Genetic Diversity of Fusarium oxysporum f. sp. cubense, the Fusarium Wilt Pathogen of Banana, in EcuadorPlants, 2020
-
van
Westerhoven et al., 2022Uncontained spread of Fusarium wilt of banana threatens African food securityPLOS Pathogens, 2022
-
Chittarath et al., 2022Geographical distribution and genetic diversity of the banana fusarium wilt fungus in Laos and VietnamJournal of Fungi, 2022
-
Baruah et al., 2025Patho-Ecological distribution and genetic diversity of Fusarium oxysporum f. sp. cubense in Malbhog banana belts of Assam, IndiaJournal of Fungi, 2025
Sampling was conducted directly on Musa plants from ‘Pisang Awak’ (ABB), ‘Silk’ (AAB), and ‘Bluggoe’ (ABB) varieties, affected by Fusarium wilt in five banana-producing areas of México: Colima, Michoacán, Jalisco, Nayarit, and Yucatán states. (Table 1). In each sampled plantation, we gather pseudostems of six- to eight-month-old wilting symptomatic plants. Discolored vascular strands were cut into 2-4 pieces and stored in a paper bag. In the laboratory, it was obtained single-spore cultures using the protocol proposed by Mostert et al. (2017). The morphology and cultural characteristics of Foc isolates were determined as described by Leslie and Summerell (2006). The isolates were stored in sterile glycerol 15% (v/v) at -80 °C. Further testing to verify the isolate identity to F. oxysporum f. sp. cubense was not conducted. For genomic DNA extraction of 96 monosporic Foc cultures, we followed the protocol of Möller and Bahnweg (1992). We analyzed six SSR loci for haplotypes characterization. The primers used are listed in Table 2. Because these SSR were suggested to study the structure of F. oxysporum (Fo) (Bogale et al., 2005), it is assumed that this work targeted isolates of Fo as well as those belonging to Foc races but race 4 (Foc R4T), which is officially not reported in Mexico. PCR cycles were done as specified by Bogale et al. (2005). The PCR products were analyzed by electrophoresis in 1.5% agarose gel and then stained in ethidium bromide solution and documented in a ChemiBis MF-2 system (Bio-Imaging Systems, Neve Yamin, Israel). A binary matrix for the six loci and a total of 96 isolates was constructed.
-
Mostert
et al. (2017The distribution and host range of the banana Fusariumwilt fungus, Fusarium oxysporum f. sp. cubense, in AsiaPlos ONE, 2017
-
Leslie and Summerell (2006The Fusarium Laboratory Manual, 2006
-
Möller and Bahnweg (1992A simple and efficient protocol for isolation of high molecular weight DNA from filamentous fungi, fruit bodies and infected plant tissuesNucleic Acids Research, 1992
-
Bogale et al., 2005Simple sequence repeat markers for species in the Fusarium oxysporum complexMolecular Ecology Notes, 2005
Table 1
Sampling for F. oxysporum from banana commercial orchards at five Mexican states.
Sampling for F. oxysporum from banana commercial orchards at five Mexican states.
| Isolate | County | State | Cultivar | Geographic location | Altitude (msnm) |
|---|---|---|---|---|---|
| ACM1 | Silk (AAB) | 18°53´23”N, 104° 0´ 0” O | 11 | ||
| ACM2 | Silk (AAB) | 18°53´23”N, 104° 0´ 0” O | 11 | ||
| ACP1 | Bluggoe (ABB) | 18° 54´ 52”N, 103° 59´38” O | 11 | ||
| ACP2 | Bluggoe (ABB) | 18° 54´ 52”N, 103° 59´38” O | 11 | ||
| ACM3 | Armería | Colima | Silk (AAB) | 18°54´5”N, 103°54´31”O | 11 |
| ACM4 | Silk (AAB) | 18°54´5”N, 103°54´31”O | 11 | ||
| ACM10 | Silk (AAB) | 18°53´10”N, 103°59´37”O | 11 | ||
| ACP11 | Bluggoe (ABB) | 18°55´24”N, 103°57´49”O | 11 | ||
| ACP15 | Bluggoe (ABB) | 18°55´24”N, 103°57´49”O | 11 | ||
| ACM13 | Silk (AAB) | 18°55´34”N, 103°58´2”O | 11 | ||
| TCM1 | 18°55´3”N, 103°5´ 13´´O | 40 | |||
| TCM2 | 18°55´3”N, 103°5´ 13´´O | 40 | |||
| TCM8 | 18°48´44”N, 103°42´49” O | 69 | |||
| TCM9 | Tecomán | Colima | Silk (AAB) | 18°48´44”N, 103°42´49” O | 69 |
| TCM10 | 18°48´44”N, 103°42´49” O | 69 | |||
| TCM11 | 18°48´44”N, 103°42´49” O | 69 | |||
| TCM12 | 18°48´44”N, 103°42´49” O | 69 | |||
| TCM13 | 18°50´ 54´´N, 163°99´ 28´´ O | 27 | |||
| TCM14 | 18°50´ 54´´N, 163°99´ 28´´ O | 27 | |||
| TCM15 | 18°50´ 54´´N, 163°99´ 28´´ O | 27 | |||
| TCM16 | 18°50´ 54´´N, 163°99´ 28´´ O | 27 | |||
| TCM1 | 18°58´2”N, 103° 50´33” O | 62 | |||
| MCP1 | Bluggoe (ABB) | 18°57´0”N, 104°3´40”O | 22 | ||
| MCT3 | Pisang awak (ABB) | 109°02´48”N, 104°06´35”O | 70 | ||
| MCT4 | Pisang awak (ABB) | 109°02´48”N, 104°06´35”O | 70 | ||
| MCT5 | Pisang awak (ABB) | 109°02´48”N, 104°06´35”O | 70 | ||
| MCT6 | Pisang awak (ABB) | 109°02´48”N, 104°06´35”O | 70 | ||
| MCM7 MCM9 | Manzanillo | Colima | Silk (AAB) Silk (AAB) | 109°0´52”N, 104°9´22”O 109°0´52”N, 104°9´22”O | 21 21 |
| MCM10 | Silk (AAB) | 109°0´52”N, 104°9´22”O | 21 | ||
| MCM11 | Silk (AAB) | 109°0´52”N, 104°9´22”O | 21 | ||
| MCM13 | Silk (AAB) | 19°00´46”O, 104°09´25”O | 21 | ||
| MCM14 | Silk (AAB) | 19°00´46”O, 104°09´25”O | 21 | ||
| MCM15 | Silk (AAB) | 19°02´21”N, 104°14´08”O | 16 | ||
| SNM1 | Silk (AAB) | 21°32´25”N, 105°07´51”O | 325 | ||
| SNP1 | Bluggoe (ABB) | 21°32´32”N, 105°08´01”O | 206 | ||
| SNM2 | Silk (AAB) | 21°32´32”N, 105°08´05”O | 206 | ||
| SNM3 | Silk (AAB) | 21°32´32”N, 105°08´05”O | 206 | ||
| SNM4 | Silk (AAB) | 21°32´41”N, 105°08´05”O | 206 | ||
| SNM5 | Silk (AAB) | 21°32´44”N, 105°08´12”O | 206 | ||
| SNM5.2 | Silk (AAB) | 21°32´44”N, 105°08´12”O | 206 | ||
| SNM6 | Silk (AAB) | 21°32´44”N, 105°08´12”O | 206 | ||
| SNM7 SNM8 | San Blas | Nayarit | Silk (AAB) Silk (AAB) | 21°32´44”N, 105°08´12”O 21°32´44”N, 105°08´17”O | 206 206 |
| SNM9 | Silk (AAB) | 21°32´44”N, 105°08´17”O | 206 | ||
| SNM10 | Pisang awak (ABB) | 21°26´36”N, 105°11´20”O | 33 | ||
| SNT1 | Pisang awak (ABB) | 21°26´36”N, 105°11´20”O | 33 | ||
| SNT2 | Pisang awak (ABB) | 21°26´36”N, 105°11´20”O | 33 | ||
| SNT3 | Pisang awak (ABB) | 21°26´36”N, 105°11´20”O | 33 | ||
| SNT4 | Pisang awak (ABB) | 21°26´36”N, 105°11´20”O | 33 | ||
| SNT5 | Pisang awak (ABB) | 21°26´36”N, 105°11´20”O | 33 | ||
| SNT6 | Pisang awak (ABB) | 21°26´36”N, 105°11´20”O | 33 | ||
| AYP1 | Bluggoe (ABB) | 20°15´53”N, 89°21´37”O | 32 | ||
| AYP2 AYM1 | Akil | Yucatán | Bluggoe (ABB) Silk (AAB) | 20°5´55”N, 89°21´37”O 20°15´52”N, 89°2´37”O | 32 32 |
| AYM2 | Silk (AAB) | 20°15´52”N, 89°21´22”O | 32 | ||
| OYM4 | Silk (AAB) | 20°13´38”N, 89°27´32”O | 49 | ||
| OYM5 OYM6 | Oxkutzcab | Yucatán | Silk (AAB) Silk (AAB) | 20°13´38”N, 89°27´32”O 20°13´14”N, 89°26´30”O | 49 49 |
| OYM7 | Silk (AAB) | 20°13´38”N, 89°27´32”O | 49 | ||
| OYM8 | Silk (AAB) | 20°13´38”N, 89°27´32”O | 49 | ||
| OYM9 | Silk (AAB) | 20°13´38”N, 89°27´32”O | 49 | ||
| OYP3 | Bluggoe (ABB) | 20°13´38”N, 89°27´32”O | 49 | ||
| OYM10 | Silk (AAB) | 20°16´42”N, 89°22´52”O | 32 | ||
| OYM11 | Silk (AAB) | 20°16´42”N, 89°22´52”O | 32 | ||
| OYM12 | Silk (AAB) | 20°16´42”N, 89°22´52”O | 32 | ||
| OYM13 | Silk (AAB) | 20°16´42”N, 89°22´52”O | 32 | ||
| OYM14 | Silk (AAB) | 20°16´42”N, 89°22´52”O | 32 | ||
| CMM1 | 18°40´41”N, 103°40´29”O | 21 | |||
| CMM2 | 18°40´43”N, 103°40´31”O | 21 | |||
| CMM3 | 18°40´22”N, 103°41´22”O | 29 | |||
| CMM4 | 18°39´57”N, 103°41´27”O | 29 | |||
| CMM5 | Coahuayana | Michoacán | Silk (AAB) | 18°44´04”N, 103°40´01”O | 27 |
| CMM6 | 18°44´04”N, 103°40´01”O | 27 | |||
| CMM7 | 18°42´59”N, 103°42´03”O | 27 | |||
| CMM8 | 18°42´59”N, 103°42´03”O | 27 | |||
| CMM9 | 18°42´59”N, 103°42´03”O | 27 | |||
| RJP1 | Silk (AAB) | 19°10´30.42´´N 104°36´01.13´´O | 12 | ||
| RJP2 | Silk (AAB) | 19°10´30.22´´N, 104°36´01.25´´O | 12 | ||
| RJP3 | Silk (AAB) | 19°10´30.22´´N, 104°36´01.57´´O | 12 | ||
| RJP4 | Silk (AAB) | 19°10´30.04´´N, 104°36´01.74´´O | 12 | ||
| RJP5 | Silk (AAB) | 19°10´29.93´´N, 104°36´02.17´´O | 12 | ||
| RJP6 | Silk (AAB) | 19°10´29.83´´N 104°36´02.41´´O | 12 | ||
| RJP7 | Silk (AAB) | 19°10´39.42´´N 104°36´07.12´´O | 12 | ||
| RJP8 | Silk (AAB) | 19°10´38.67´´N 104°36´06.60´´O | 12 | ||
| RJP9 | Silk (AAB) | 19°10´38.38´´N 104°36´05.71´´O | 12 | ||
| RJP10 RJP11 | El Rebalse | Jalisco | Silk (AAB) Bluggoe (ABB) | 19°10´38.86´´N 104°36´07.77´´O 19°10´38.66´´N 104°36´06.80´´O | 12 14 |
| RJM12 | Bluggoe (ABB) | 19°10´39.20´´N 104°36´09.30´´O | 14 | ||
| RJM13 | Bluggoe (ABB) | 19°10´38.68´´N 104°36´09.20´´O | 14 | ||
| RJM14 | Bluggoe (ABB) | 19°11´15.86´´N, 104°35´29.56´´O | 14 | ||
| RJM15 | Bluggoe (ABB) | 19°11´16.26´´N, 104°35´29.50´´O | 14 | ||
| RJM16 | Bluggoe (ABB) | 19°11´17.05´´N, 104°35´17.05´´O | 14 | ||
| RJM17 | Bluggoe (ABB) | 19°11´17.35´´N, 104°35´27.92´´O | 14 | ||
| RJM18 | Bluggoe (ABB) | 19°11´17.72´´N, 104°35´26.70´´O | 14 | ||
| RJM19 | Bluggoe (ABB) | 19°11´21.94´´N, 104°35´09.52´´O | 14 | ||
| RJM20 | Bluggoe (ABB) | 19°11´21.83´´N, 104°35´08.53´´O | 14 |
Table 2
Simple sequence repeat (SSR) primers used to analyze Fusarium oxysporum isolates genetic diversity (modified from Bogale et al., 2005).
Simple sequence repeat (SSR) primers used to analyze Fusarium oxysporum isolates genetic diversity (modified from Bogale et al., 2005).
| Primer | xPrimer secuence (5´-3´) | Annealing temperature (°C) |
|---|---|---|
| G1/ MV15 | R: CTCGTCCTTTGCGAATGACC | 58 |
| G: ACCACCTCGGTGATGGTGAGACGG | ||
| R: CAAGCCGCTCTCCACGGCGAAGGCG | ||
| MB18 | F: GGTAGGAAATGACGAAGCTGAC | 57 |
| R: TGAGCACTCTAGCACTCCAAAC | ||
| R: CGTCCTCAAGAGCAGCGAC |
-
Bogale et
al., 2005Simple sequence repeat markers for species in the Fusarium oxysporum complexMolecular Ecology Notes, 2005
The total isolates were organized into three groups, to improve the robustness of the analysis and to facilitate data interpretation. Thus, we considered isolates form Colima, Jalisco and Michoacán as a single population because of their regional continuity and the lack of apparent ecological barriers. Isolates from Nayarit integrate the second population, and those from Yucatán are the third population. Later, based on geographical proximity and the banana fruit and propagule trading routes, these were further allocated into two regions: (i) East México (samples from Yucatán), and (ii) West México (isolates from Nayarit, Colima, Jalisco, and Michoacán). The number and frequency of haplotypes were calculated in POPGENE. With this data, using the dissimilarity index of Morisita in the R package vegan (Oksanen et al. 2019), the genetic similarity of the populations was estimated. With the haplotype data, we also conducted a molecular analysis of variance (AMOVA) to evaluate the genetic differentiation between regions (FCT), among populations in each region (FSC), and among all populations included in the analysis (FST). We performed the AMOVA in ARLEQUIN v. 3.5 using 1000 permutations. Additionally, to illustrate the relations among haplotypes, and their distribution and frequency among populations we generate a minimum spanning network. Genetic structure was inferred using the principal component analysis and the sparse non-negative matrix factorization (sNMF) in LEA (Frichot and François, 2015). We selected the preferred number of K using a cross-entropy criterion based on the prediction of masked genotypes to evaluate the error of ancestry estimation.
-
Oksanen et
al. 2019Vegan: Community Ecology Package. R packageversion 2.5-6, 2019
-
Frichot and François, 2015LEA: An R package for landscape and ecological association studiesMethods in Ecology and Evolution, 2015
Results of cultural characterization showed that colony size development ranged from
8.7 to 9.8 mm per day on PDA medium at 28 °C. After seven days of culture, all isolates had cottony white mycelia, which can be flat to aerial, circular or irregular in shape, colored on the upper surfaces from light violet to dark violet. On the petri dish reverse, color colonies ranged from orange to dark red. Short monophialides had an average of 3-
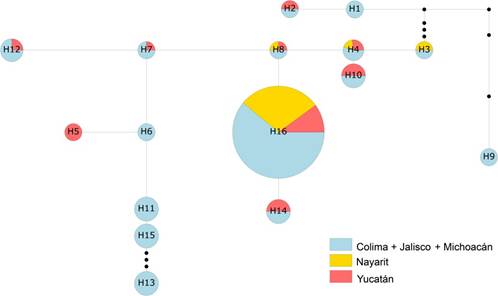
5 septa, unicellular or bicellular, with oval microconidia of 5-15 × 2.5-3.5 μm. Macroconidia were 23.5-47.8 × 3.3-4.8 μm, falciform, with three to four septa (Leslie and Summerell, 2006). These characteristics were typical to Fusarium oxysporum. All tested loci were polymorphic across the 96 Mexican isolates of Fusarium oxysporum. We detect 16 haplotypes, of which the most frequent and common was the H16 (Figure 1). The rest were widely dispersed or restricted to one or two populations.
-
Leslie and Summerell, 2006The Fusarium Laboratory Manual, 2006
Thumbnail
Figure 1
A minimum spanning network of 16 haplotypes found in the studied populations of Fusarium oxysporum. Small black dots along branches represent missing haplotypes. H withing circles is the ID haplotype, color denote population type, and circle size represents the haplotype frequency.
A minimum spanning network of 16 haplotypes found in the studied populations of Fusarium oxysporum. Small black dots along branches represent missing haplotypes. H withing circles is the ID haplotype, color denote population type, and circle size represents the haplotype frequency.
Table 3
Dissimilarity among sampled populations of Fusarium oxysporum .
Dissimilarity among sampled populations of Fusarium oxysporum .
| Population | Colima, Jalisco and Michoacán | Nayarit | Yucatán |
|---|---|---|---|
| Colima, Jalisco and Michoacán | – | – | – |
| Nayarit | 0.097 | – | – |
| Yucatán | 0.000 | 0.255 | – |
Thumbnail
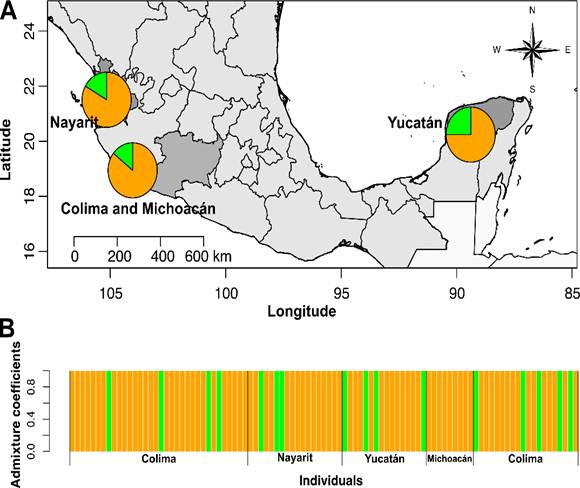
Figure 2
Genetic structure analysis of Fusarium oxysporum isolates from different locations based on sparse nonnegative matrix factorization (sNMF). (A) Population structure for K = 2 plotted on the map using the average ancestry coefficient values for each estimated population. (B) Group assignment probability of each sampled individual.
Genetic structure analysis of Fusarium oxysporum isolates from different locations based on sparse nonnegative matrix factorization (sNMF). (A) Population structure for K = 2 plotted on the map using the average ancestry coefficient values for each estimated population. (B) Group assignment probability of each sampled individual.
Table 4
Analysis of molecular variance (AMOVA) based on haplotypes associated to 96 isolates of Fusarium oxysporum from Colima, Jalisco, Michoacán, Nayarit and Yucatán state grouped into two regions (East and West México).
Analysis of molecular variance (AMOVA) based on haplotypes associated to 96 isolates of Fusarium oxysporum from Colima, Jalisco, Michoacán, Nayarit and Yucatán state grouped into two regions (East and West México).
| Source of variationX | d.f. | Sum of squares | Variance components | Variation (%) | >FI | P |
|---|---|---|---|---|---|---|
| Between regions | 1 | 0.921 | -0.00145 | -0.19 | -0.00188 | 0.66960 ± 0.019 |
| Between population within regions | 1 | 1.012 | 0.00888 | 1.15 | 0.01150 | 0.21114 ± 0.014 |
| Within populations | 94 | 71.778 | 0.76359 | 99.04 | 0.00964 | 0.23460 ± 0.011 |
| Total | 96 | 73.711 | 0.77103 | – | – | – |
According to these results, Mexican Fo populations are genetically similar (Tables 3 y 4). Magdama et al., (2020), also concluded that the populations of Foc in Ecuador comprise a single clonal lineage. In this work several Fo haplotypes were detected but only one was dominant (Figure 1). This may be related to the low variability rate of Fo as asexual organism compared to sexually reproducing organisms such as Pseudocercospora fijiensis (Manzo-Sánchez et al., 2019; McDonald and Linde, 2002). These results also agree with Bentley (1998), who detects only one Foc lineage in Mexican populations (lineage VIII). The same author describes that this lineage is unique in Central America, but he did not assign a vegetative compatibility groups (VCG) code. Ordoñez (2018) specifies that VCG 0124 found in México corresponds to the 0124/5/8/20 compatibility group complex, whose worldwide members had been identified withing the 1, 2, and 4 race. The Foc VCG found in México allows us to hypothesize that the pathogen in this country is a mixture between a native and introduced strains. If this is true, human activities are responsible for the dissemination of the clonal lineage.
-
Magdama et al., (2020Genetic Diversity of Fusarium oxysporum f. sp. cubense, the Fusarium Wilt Pathogen of Banana, in EcuadorPlants, 2020
-
Manzo-Sánchez et al.,
2019Genetic variability of Pseudocercospora fijiensis, the black Sigatoka pathogen of banana (Musa spp.) in MéxicoPlant Pathology, 2019
-
McDonald and Linde, 2002The population genetics of plant pathogens and breeding strategies for durable resistanceEuphytica, 2002
-
Bentley
(1998Geneticvariation among vegetative compatibility groups of Fusarium oxysporum f. sp. cubense analyzed by DNA fingerprintingPhytopathology, 1998
-
Ordoñez (2018A global geneticdiversity analysis of Fusarium oxysporum f. sp. cubense, 2018
The fungus Fusarium oxysporum f. sp. cubense is a genetically diverse pathogen with a cosmopolitan distribution. It is composed of four races, of which three infect plantains and bananas (Ploetz, 2015; Pegg et al., 2019; Martinez-de la Parte et al., 2024; Baruah et al., 2025). Also, more than 24 vegetative compatibility groups have been recognized (Koening et al., 1997; Ordoñez, 2018; Maryani et al., 2019), and at least nine lineages phylogenetically clustered in two clades have been detected (Bentley et al., 1998; Czislowski et al., 2018; O’Donnell et al., 1998; Pegg et al., 2019). Because Foc reproduces preferentially asexually, its genetic diversity is grouped in clonal lineages (Bentley et al., 1998; Pegg et al., 2019); this diversity can be enriched occasionally by parasexual events in which horizontal gene transfer can occur (Czislowski et al., 2018; Fourie et al., 2009; Ma et al., 2010; Vlaardingerbroek et al., 2016). Thus, the genetic structure of Foc (or the absence of it) corresponds to what is expected for an organism that preferentially reproduces asexually (McDonald and Linde, 2002). In fact, the genetic diversity of this pathogen (most VCG) is concentrated in its center of origin, and only a few VCG are scattered around the world (Bentley et al., 1998; Koening et al., 1997; Mostert et al., 2017), which can explain the absence of detectable genetic structure in the sampled populations.
-
Ploetz, 2015Fusarium wilt of bananaPhytopathology, 2015
-
Pegg et al., 2019The Epidemiology of Fusarium wilt of bananaFrontiers in Plant Science, 2019
-
Martinez-de la Parte et al., 2024Genetic diversity of the banana Fusarium wilt pathogen in Cuba and across Latin America and the CaribbeanEnvironmental Microbiology, 2024
-
Baruah et al., 2025Patho-Ecological distribution and genetic diversity of Fusarium oxysporum f. sp. cubense in Malbhog banana belts of Assam, IndiaJournal of Fungi, 2025
-
Koening et al., 1997Fusarium oxysporum f. sp. cubense consists of a small number of divergent and globally distributed clonal lineagesPhytopathology, 1997
-
Ordoñez, 2018A global geneticdiversity analysis of Fusarium oxysporum f. sp. cubense, 2018
-
Maryani et al., 2019Phylogeny and genetic diversity of the bananaFusarium wilt pathogen Fusarium oxysporum f. sp. cubense in the Indonesian centre of originStudies in Mycology, 2019
-
Bentley et al., 1998Geneticvariation among vegetative compatibility groups of Fusarium oxysporum f. sp. cubense analyzed by DNA fingerprintingPhytopathology, 1998
-
Czislowski et al., 2018Investigation of the diversity of effector genes in the banana pathogen, Fusarium oxysporum f. sp. cubense, reveals evidence of horizontal gene transferMolecular Plant Pathology, 2018
-
O’Donnell et al.,
1998Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogiesProceedings of the National Academy of Sciences, 1998
-
Fourie et al., 2009Evolutionary relationships among the Fusarium oxysporum f. sp. cubense vegetative compatibility groupsApplied and Environmental Microbiology, 2009
-
Vlaardingerbroek et al., 2016Exchange of core chromosomes and horizontal transfer of lineage-specific chromosomes in Fusarium oxysporumEnvironmental Microbiology, 2016
-
McDonald and Linde, 2002The population genetics of plant pathogens and breeding strategies for durable resistanceEuphytica, 2002
-
Mostert
et al., 2017The distribution and host range of the banana Fusariumwilt fungus, Fusarium oxysporum f. sp. cubense, in AsiaPlos ONE, 2017
Why Foc is diverse, and cosmopolitan worldwide, but locally may lack genetic structure? We found three possible explanations: one is that the pathogen had effectively spread because of the exchange of contaminated materials (rhizomes or leaf trash) (Ploetz, 2015). Another explanation is that it is a polyphyletic complex; thus, it is possible that native local strains parasitically converged to the introduction of Musa spp. (Gurr, 2011). The third possibility is that indeed Musa spp. may have dispersed together with some of the Foc clonal lineages which exchanged genetic information using parasexual or perhaps sexual mechanisms with local native linages (Ordoñez, 2018). In México it is not known the dispersal dynamics of Foc (Ordoñez, 2018), although Foc -mainly race 1- it is widely distributed in the territory.
-
Ploetz, 2015Fusarium wilt of bananaPhytopathology, 2015
-
Ordoñez, 2018A global geneticdiversity analysis of Fusarium oxysporum f. sp. cubense, 2018
In conclusion, cultural morphology of monosporic Foc isolates were consistent with what has been reported in the literature for Foc. Despite the haplotypes diversity found (16), H16 was the most frequent, while the remaining haplotypes may be either widely distributed or local, but with low frequency. No significant genetic differences were found among the studied populations or grouped by regions. Therefore, these results using six SSR markers suggest that the analyzed populations constitute a single metapopulation.
Conflicts interest
All the authors declare no conflict of interest.
Acknowledgments
This work was supported by Universidad de Colima, México.
Author contributions
First, second and four authors provided the genomic methodology and sampling. Second author extensive sampling and phytosanitary problem conception. All contributed to writing and discussion.
Referencias
- Baruah, A, Bora, P, Damodaran, T, Saikia, B, Manoharan, M, et al. (2025). Patho-Ecological distribution and genetic diversity of Fusarium oxysporum f. sp. cubense in Malbhog banana belts of Assam, India. Journal of Fungi, 11, 195. https://doi.org/ 10.3390/jof11030195 Links
- Bentley, S, Pegg, KG, Moore, NY, Davis, RD and Buddenhagen, IW. (1998). Geneticvariation among vegetative compatibility groups of Fusarium oxysporum f. sp. cubense analyzed by DNA fingerprinting. Phytopathology 88:1283-1293. https://doi.org/ 10.1094/phyto.1998.88.12.1283 Links
- Bogale, M, Wingfield, BD, Wingfield, MJ and Steenkamp, ET. (2005). Simple sequence repeat markers for species in the Fusarium oxysporum complex. Molecular Ecology Notes 5:622-624. https://doi.org/ 10.1111/j.1471-8286.2005.01015.x Links
- Chittarath, K, Nguyen, CH, Bailey, WC, Zheng, SJ, Mostert, D, et al. (2022). Geographical distribution and genetic diversity of the banana fusarium wilt fungus in Laos and Vietnam. Journal of Fungi 8: 46. https://doi.org/ 10.3390/jof8010046 Links
- Czislowski, E, Fraser-Smith, S, Zander, M, O’Neill, WT, Meldrum, RA, et al. (2018). Investigation of the diversity of effector genes in the banana pathogen, Fusarium oxysporum f. sp. cubense, reveals evidence of horizontal gene transfer. Molecular Plant Pathology 19:1155-1171. https://doi.org/ 10.1111/mpp.12594 Links
- Food and Agriculture Organization of the United Nations, Statistics (FAOSTAT). (2024). FAO Rome, Italy. http://www.fao.org/faostat/es/#data/WCAD . Consulta, abril 2025. Links
- Fourie, G, Steenkamp, ET, Gordon, TR and Vijoen, A. (2009). Evolutionary relationships among the Fusarium oxysporum f. sp. cubense vegetative compatibility groups. Applied and Environmental Microbiology 75:4770-4781. https://dx.doi.org/ 10.1128/AEM.00370-09 Links
- Frichot, E and François, O. (2015). LEA: An R package for landscape and ecological association studies. Methods in Ecology and Evolution 6:925-929. https://doi.org/ 10.1111/2041-210X.12382 Links
- Koening, RL, Ploetz, RC and Kistler, HC. (1997). Fusarium oxysporum f. sp. cubense consists of a small number of divergent and globally distributed clonal lineages. Phytopathology 87:915-923. https://doi.org/ 10.1094/PHYTO.1997.87.9.915 Links
- Leslie, JF and Summerell, BA. (2006). The Fusarium Laboratory Manual. Blackwell Publishing-Wiley, Hoboken, NJ, USA. 388 p. Links
- Ma, L, Van der Does, H, Borkovich, K, Coleman, JJ, Daboussi, MJ, et al. (2010). Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 464:367-373. https://doi.org/ 10.1038/nature08850 Links
- Magdama, F, Monserrate-Maggi, L, Serrano, L, García Onofre, J, and Jiménez-Gasco, MM. (2020). Genetic Diversity of Fusarium oxysporum f. sp. cubense, the Fusarium Wilt Pathogen of Banana, in Ecuador. Plants 9:1133. https://doi.org/ 10.3390/plants9091133 Links
- Manzo-Sánchez, G, Orozco-Santos, M, Islas-Flores, I, Martínez-Bolaños, L, Guzmán-González, S, et al. (2019). Genetic variability of Pseudocercospora fijiensis, the black Sigatoka pathogen of banana (Musa spp.) in México. Plant Pathology 68:513-522. https://doi.org/ 10.1111/ppa.12965 Links
- Maryani, N, Lombard, L, Poerba, YS, Subandiyah, S, Crous, PW, et al. (2019). Phylogeny and genetic diversity of the bananaFusarium wilt pathogen Fusarium oxysporum f. sp. cubense in the Indonesian centre of origin. Studies in Mycology 92: 155-194. Links
- Martínez-de la Parte, E, Pérez-Vicente, L, Torres, DE, van Westerhoven, A, Meijer, HJ, et al. (2024). Genetic diversity of the banana Fusarium wilt pathogen in Cuba and across Latin America and the Caribbean. Environmental Microbiology, 26 : e16636. https://doi.org/ 10.1111/1462-2920.16636 Links
- McDonald, BA and Linde, C. (2002). The population genetics of plant pathogens and breeding strategies for durable resistance. Euphytica 124:163-180. Links
- Möller, EM and Bahnweg, G. (1992). A simple and efficient protocol for isolation of high molecular weight DNA from filamentous fungi, fruit bodies and infected plant tissues. Nucleic Acids Research 20: 6115-6116. https://doi.org/ 10.1093/nar/20.22.6115 Links
- Mostert, D, Molina, AB, Daniells, J, Fourie, G, Hermanto, C, et al. (2017). The distribution and host range of the banana Fusariumwilt fungus, Fusarium oxysporum f. sp. cubense, in Asia. Plos ONE 12: e0181630. https://doi.org/ 10.1371/journal.pone.0181630 Links
- O’Donnell, K, Kistler, HC, Cigelnik, E and Ploetz, R. (1998). Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogies. Proceedings of the National Academy of Sciences 95: 2044-2049. https://doi.org/ 10.1073/pnas.95.5.2044 Links
- Oksanen, J, Blanchet, FG, Friendly, M, Kindt, R, Legendre, P, et al. (2019). Vegan: Community Ecology Package. R packageversion 2.5-6. https://CRAN.R-project.org/package=vegan . Consulta, septiembre 2019. Links
- Ordoñez, N. (2018). A global geneticdiversity analysis of Fusarium oxysporum f. sp. cubense. PhD. Thesis. Wageningen University, Wageningen, The Netherlands. 156 pp. https://doi.org/ 10.18174/453455 Links
- Pegg, KG, Coates, LM, O’Neill, WT and Turner, DW. (2019). The Epidemiology of Fusarium wilt of banana. Frontiers in Plant Science 10:1395. doi: 10.3389/fpls.2019.01395 Links
- Ploetz, RC. (2015). Fusarium wilt of banana. Phytopathology 105:1512-1521. http://dx.doi.org/ 10.1094/PHYTO-04-15-0101-RVW Links
- Servicio de Información Agroalimentaria y Pesquera (SIAP). (2024). Cierre de la producción agrícolapor estado. Recuperadode http://www.siap.gob.mx/cierre-de-la-produccion-agricola-por-estado/ . Consulta, enero 2024. Links
- van Westerhoven, AC, Meijer, HJG, Seidl, MF and Kema, GHJ. (2022). Uncontained spread of Fusarium wilt of banana threatens African food security. PLOS Pathogens 18: e1010769. Links
- Vlaardingerbroek, I, Beerens, B, Rose, L, Fokkens, L, Cornelissen, BJC, et al. (2016). Exchange of core chromosomes and horizontal transfer of lineage-specific chromosomes in Fusarium oxysporum. Environmental Microbiology 18: 3702-3713. https://doi.org/ 10.1111/1462-2920.13281. Links