Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Accesos
Accesos
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO
Compartir
Revista mexicana de fitopatología
versión On-line ISSN 2007-8080versión impresa ISSN 0185-3309
Rev. mex. fitopatol vol.42 no.3 Texcoco sep. 2024 Epub 27-Mayo-2025
https://doi.org/10.18781/r.mex.fit.2403-2
Phytopathological notes
Identification of phytoplasmas associated with Bunchy Top disease of papaya in Colima, Mexico
1 Postgrado en Fitosanidad-Fitopatología, Colegio de Postgraduados, km 36.5 Carretera México-Texcoco, Montecillo, Texcoco, Estado de México, CP 56264. México
2 Universidad de Colima, Facultad de Ciencias Biológicas y Agropecuarias, Autopista Colima-Manzanillo km 40, Tecomán, Colima, México, CP 28930.
Background/Objective:
Phytoplasmas, rickettsiae and viruses have been detected in papaya plants with Bunchy Top disease (BT). In 2019, papaya plants with BTlike symptoms were observed in agroecosystems of Colima, Mexico. In order to determine the BT-associated phytoplasmas species or subgroups, asymptomatic and symptomatic plants were collected from papaya agroecosystems in four papaya producer municipalities, as well as papaya-associated weeds and insects.
Materials and Methods:
Phytoplasma detection and identification was conducted by PCR, sequencing and phylogenetics of translocase subunit SecA (secA) and 16S ribosomal RNA (16Sr) genes, and PCR-RFLPs in vitro and in silico for 16Sr gene.
Results:
In papaya, phytoplasma groups 16SrI (subgroup AF), 16SrX, and 16SrXIII were identified in 2.08% (4 out of 192) symptomatic samples. The results of RFLPs in silico analysis showing the presence of 16SrX and 16SrXIII (sub)groups. In papaya-associated weeds and insects, phytoplasmas of group 16SrI (subgroups AF and B) were identified in 1.7% (3 out of 174) and 1.1% (2 out of 185) evaluated samples, respectively. Phytoplasma-carrying weeds were Amaranthus palmeri and Echinochloa colona; positive insects were Micrutalis calva and Balclutha mexicana.
Conclusion:
It is the first time that phytoplasmas 16SrI-AF, 16SrX y 16SrXIII are associated with Bunchy Top disease of papaya in agroecosystems from Colima, Mexico. Phytoplasmas 16SrX y 16SrXIII are first reported in papaya plants at the world level and in Mexico, respectively. Phytoplasma-carrying weeds and insects are new records as natural reservoirs and potential vectors.
Key words: Carica papaya; ‘Candidatus Phytoplasma’; weeds; insects
Antecedentes/Objetivo:
Fitoplasmas, rickettsias y virus se han detectado en plantas de papaya con la enfermedad Bunchy Top (BT). En agroecosistemas de papaya ubicados en Colima, México, durante 2019 se observaron plantas con síntomas similares al BT. Con la finalidad de determinar la presencia de fitoplasmas e identificar las especies y/o subgrupos asociados a esta enfermedad, se recolectaron plantas asintomáticas y con síntomas en cuatro municipios productores del estado, así como malezas e insectos asociados al cultivo.
Materiales y Métodos:
La detección e identificación de fitoplasmas se hizo a través de PCR, secuenciación y análisis filogenéticos de la subunidad SecA de la translocasa (secA) y 16S RNA ribosomal (16Sr), y PCR-RFLPs in vitro e in silico del gen 16Sr.
Resultados:
En papaya se identificaron fitoplasmas de los grupos 16SrI (subgrupo AF), 16SrX y 16SrXIII, en el 2.08% (4 de 192) de las muestras con síntomas. Los resultados del análisis RFLPs in silico del gen 16Sr mostraron la presencia de los (sub)grupos 16SrX y 16SrXIII. En malezas e insectos, se identificaron fitoplasmas del grupo 16SrI (subgrupos AF y B) en el 1.7% (3 de 174) y 1.1% (2 de 185) de las muestras analizadas, respectivamente. Las malezas portadoras de fitoplasmas fueron Amaranthus palmeri y Echinochloa colona, mientras que los insectos fueron Micrutalis calva y Balclutha mexicana.
Conclusión:
Se reporta por primera vez la presencia de fitoplasmas 16SrI-AF, 16SrX y 16SrXIII en plantas de papaya con síntomas de Bunchy Top en agroecosistemas del estado de Colima, México. Los fitoplasmas de los grupos 16SrX y 16SrXIII se registran por primera vez a nivel mundial y en México en papaya, respectivamente. Las malezas e insectos portadores de fitoplasmas constituyen nuevos registros como reservorios naturales y potenciales vectores de las bacterias.
Palabras-clave: Carica papaya; ‘Candidatus Phytoplasma’; malezas; insectos
Introduction
Mexico is the third largest producer of papaya (Carica papaya) in the world and the main supplier of fresh fruit for the United States of America and Canada, where annual sales are greater than 350,000 tons, with a value of 335 million dollars (SIAP, 2020). In this country, the production of the caricaceae is located in the South-Southeast and Center-West regions, with the states of Oaxaca, Colima and Chiapas being the main producer states (SIAP, 2020). The state of Colima, located in the West on the coast of the Pacific Ocean, has a record of papaya production of 192,417 tons and is the main national exporter (Colima State Papaya Farmers Council [COEPAPAYA A.C.], 2017; SIAP, 2020).
In 2019, in 64 papaya agroecosystems visited in four municipalities of the state of Colima, plants showed stems with shortened internodes that caused the bunching of apical leaves. These plants also displayed leaves with chlorosis, yellowing, deformation and/or marginal necrosis. These symptoms are typical of the Bunchy Top (BT) disease (Cook, 1931; Story and Halliwell, 1969), and similar to those described in Cuba as “papaya bunchy top” (PBT) and “bunchy top symptom” (BTS) syndromes (Acosta et al., 2013).
BT is induced by phytoplasmas, rickettsiae and viruses, occasionally in mixed infections (phytoplasmas + rickettsiae or phytoplasmas + viruses), and it has been reported in the Dominican Republic, Cuba, Haiti, Jamaica, Puerto Rico, Costa Rica, Peru (Martorell and Adsuar, 1952; Story and Halliwell, 1969; Davis et al., 1998; Arocha et al., 2003; Acosta et al., 2013; Luis-Pantoja et al., 2015; Wei et al., 2017), and more recently, in Nigeria (Kazeem et al., 2021).
Considering that in Mexico phytoplasmas have been found in papaya plants with symptoms similar to BT (Pogoshyan et al., 2004; Navarrete-Yabur et al., 2005; Rojas-Martínez et al., 2011), and that in Colima this disease has become more economically important and its study has been scarcely covered (Colima papaya farmers, personal communication), the following goals have been set: a) to determine the presence of phytoplasmas in papaya plants with BT symptoms, as well as in weeds and insects related with the crop, and b) to establish their identity at the 16Sr group and/or subgroup level.
In November and December of 2019, 256 papaya samples were gathered from 64 commercial agroecosystems located in the municipalities of Tecomán, Colima, Ixtlahuacán and Armería. Out of all the samples, 64 were asymptomatic and 192 displayed the symptoms described earlier, related to BT. A compound sample was taken from every plant, consisting of three leaves (blade and petiole), each one collected from the upper, middle and lower strata of the plant. From every papaya plant (symptomatic or asymptomatic sampled) up to five species of weeds present within a 2-meter diameter were gathered (depending on their availability and greater abundance). Insects were also gathered with a beating entomological net using three downward sweeps on the papaya plant and adjacent weeds. The insects were kept in 96% ethanol for later classification into morphospecies. The weeds and insects that tested positive for phytoplasmas were identified at the species level. From the central veins and petioles of papaya leaves and weeds, as well as from the entire specimens of insects (one to five, depending on size and availability), the total DNA was extracted using a phytoplasma DNA enrichment protocol based on CTAB (Ahrens and Seemuller, 1992) and with the help of a homogenizer (Tissue Lyser, Qiagen, Germany) (Stillson and Szendrei, 2020). The DNA was evaluated by UV spectrophotometry (NadoDrop 2000, Fisher Scientific, USA) and by PCR amplification of the ribosomal S16 (rps16) gene of plants (Oxelman et al., 1997) or cytochrome oxidase subunit I (COI) of insects (Folmer et al., 1994), as internal controls in reaction volumes of 25 μL with 50 ng of DNA.
The detection of phytoplasmas was performed on 200 ng of DNA by nested endpoint PCR of the 16Sr genes (first round: primers P1/P7 and nested: primers R16F2n/R16R2 and R16mF2/R16mR1) (Deng and Hiruki, 1991; Schneider et al., 1995; Gundersen and Lee, 1996; Lee et al., 1993) and secA (first round: primers secAFor1/secARev3 and nested: primers SecAFor5/SecARev2) (Dickinson and Hodgetts, 2013).
Identification was performed by sequencing, PCR-RFLPs in vitro and in silico of the gene 16Sr and phylogenetic analysis of secA and 16Sr. For sequencing, the nested PCR amplicons for 16Sr (primers R16mF2/R16mR1) and secA of the positive samples were purified with the Wizard® SV Gel and PCR Clean-Up System kit (Promega, USA) and sequenced in both directions with the primers R16mF2/ R16mR1 and SecAFor5/SecARev2, using the dideoxy or chain termination method at Macrogen Inc. (Seoul, South Korea). The resulting sequences were analyzed with the MEGAX program (Kumar et al., 2018) and the BLASTn function of the National Center for Biotechnology Information (NCBI) (https://blast.ncbi.nlm.nih. gov/Blast.cgi), for deposition in the GenBank (https://submit.ncbi.nlm.nih.gov/).
For the in vitro PCR-RFLPs, approximately 1 μg of each purified product of the nested PCR for the 16Sr gene (primers R16F2n/R16R2) of the positive samples, quantified by UV spectrophotometry (NanoDrop 2000, Fisher Scientific, USA) was conducted separately with 10 U of the restriction enzymes Taq I, Hae III, Sau 3AI, Kpn I, Tru I1 (Mse I), Alu I and Rsa I (Thermo Scientific, USA), following the instructions by the manufacturer to confirm that the amplified fragment corresponded to the DNA of phytoplasmas (Lee et al., 1998). The products were separated by electrophoresis in 3% agarose gels (w/v) in a 1×TAE buffer and dyed with ethidium bromide (Sambrook and Russell, 2001).
For the in silico RFLPs, the sequences of the amplicons of the nested PCR (primers R16mF2/R16mR1), edited manually with MEGAX (Kumar et al., 2018) were analyzed using the software iPhyClassifier (https://acortar.link/AikrtQ) to assign and/or confirm species, 16Sr groups and subgroups.
The phylogenetic analyses were carried out independently with the secA and 16Sr genes. To do this, the sequences of each one of the analyzed genes from different groups of phytoplasmas reported worldwide (49 for secA and 51 for 16Sr) and from Bacillus subtilis (GenBank: X62035 for secA) or from Acholeplasma laidlawii (GenBank: U14905 for 16Sr) as outgroups were downloaded from the GenBank and aligned with Clustal W. Phylogenies were built with the neighborjoining method with 1000 bootstrap repetitions. All analyses were carried out in MEGA X (Kumar et al., 2018).
Phytoplasmas were detected in 2.08% (4 out of 192) papaya plants with symptoms related to BT (Table 1). No phytoplasmas were found in asymptomatic plants. The positive plants were gathered in the municipalities of Colima and Ixtlahuacán (Table 1) and they displayed stems with shortened internodes and yellow chlorotic leaves (Figure 1A). Positive samples were confirmed by sequencing the secA (GenBank: ON303285-ON303288) and 16Sr (GenBank: PP348058-PP348061) genes. Sequence analysis of the secA and 16Sr genes with BLASTn indicate that, in two of the four positive samples, the phytoplasmas belong to the 16SrI group (GenBank secA: ON303286 and ON303287; 16Sr: PP348059 and PP348060), whereas the remaining two belong to the 16SrX (GenBank secA: ON303288; 16Sr: PP348061) and 16SrXIII (GenBank secA: ON303285; 16Sr: PP348058) groups, respectively.
Table 1 Detection of phytoplasmas in papaya (Carica papaya), weeds and insects associated with the crop in Colima, Mexico, in November and December of 2019.
| Municipality | Number of positive samples to phytoplasmas / Number of analyzed samples (%) | ||
|---|---|---|---|
| Papaya | Weeds | Insects | |
| Armería | 0/64 (0.0) | 1/74 (1.4) | 1/60 (1.7) |
| Colima | 2/64 (3.1) | 0/31 (0.0) | 0/32 (0.0) |
| Ixtlahuacán | 2/64 (3.1) | 1/35 (2.9) | 0/46 (0.0) |
| Tecomán | 0/64 (0.0) | 1/34 (2.9) | 1/47 (2.1) |
| Total | 4/256 (1.6) | 3/174 (1.7) | 2/185 (1.1) |
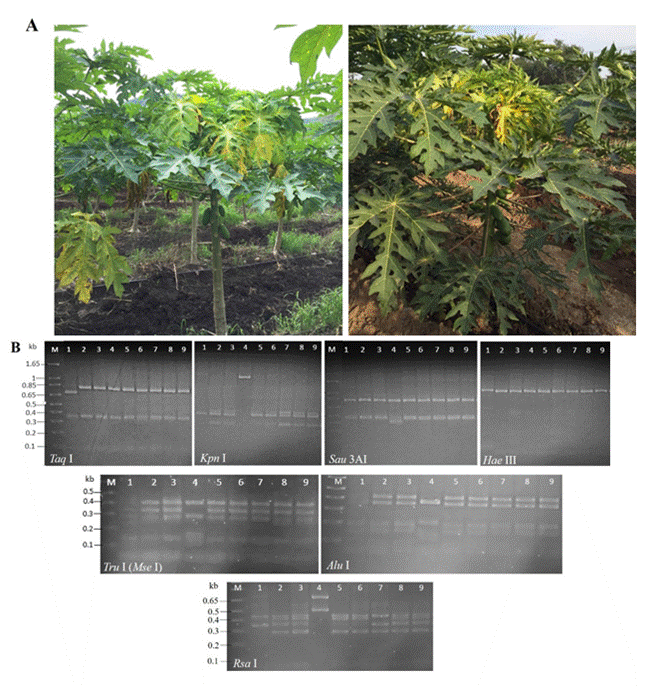
Figure 1 Detection of phytoplasmas in agroecosystems papaya with Bunchy Top disease in Colima, Mexico. A) Papaya plants showing stems with shortened internodes and yellow, chlorotic leaves, positive to phytoplasmas. B) In vitro RFLPs profiles of gene 16Sr (primers R16F2n/R16R2) of the phytoplasmas under study. Lanes 1-4: samples 1S-4Spapaya, Lanes 5-6: samples 5S-6S-Amaranthus palmeri; Lane 7: sample 7S-Echinochloa colona; Lane 8: sample 8S-Micrutalis calva; and Lane 9: sample 9S-Balclutha mexicana. The DNA was digested with 10 U of every enzyme and the products were separated by electrophoresis in 3% (w/v) agarose gels in 1×TAE buffer. Lane M: 1 kb plus molecular weight marker (Thermo Scientific, EUA).
The profiles of the in vitro RFLPs of the 16Sr gene (Figure 1B) were consistent with those reported by Lee et al. (1998) and with those obtained using iPhyClassifer (Figure 2) for the restriction enzymes used, which helped authenticate the DNA of the phytoplasmas found in this study. The in silico RFLPs analysis of the 16Sr gene confirmed the presence of the ‘Candidatus Phytoplasma asteris’ [group 16SrI (percentages of identity: 99.44 and 99.60) species and subgroup 16SrI AF (coefficient of similarity: 1.00)], ‘Ca. Phytoplasma rhamni’ [group 16SrX (percentage of identity: 93.31), undetermined subgroup] and ‘Ca. Phytoplasma hispanicum’ [group 16SrXIII (percentage of identity: 99.37), subgroup 16SrXIII-D (coefficient of similarity: 0.95)] (Table 2). Regarding the percentages of identity

Figure 2 In silico RFLPs profiles of gene 16Sr of the phytoplasmas under study, generated with iPhyClassifier with 17 restriction enzymes. A) sample 1S-papaya; B) sample 2S-papaya; C) sample 3S-papaya; D) sample 4S-papaya, E) sample 5S-Amaranthus palmeri; F) sample 6S-Amaranthus palmeri; G) sample 7S-Echinochloa colona; H) sample 8S-Micrutalis calva; I) sample 9S-Balclutha mexicana. Lanes MW: φX174 DNA digested with Hae III.
Table 2 Phytoplasmas identified in papaya, weeds and insects associated with the crop in Colima, Mexico, in NovemberDecember of 2019.
| Sample (GenBank 16Sr) | ‘Candidatus Phytoplasma sp.’ | Group 16Sr (% similarity) | Subgroup (coefficient of similarity) |
|---|---|---|---|
| 1S-papaya (PP348058) | ‘Ca. Phytoplasma hispanicum’ | 16SrXIII (99.37) | D (0.95)y |
| 2S-papaya (PP348059) | ‘Ca. Phytoplasma asteris’ | 16SrI (99.44) | AF (1.00) |
| 3S-papaya (PP348060) | ‘Ca. Phytoplasma asteris’ | 16SrI (99.60) | AF (1.00) |
| 4S-papaya (PP348061) | ‘Ca. Phytoplasma rhamni’ | 16SrX (93.31) | NAz |
| 5S-Amaranthus palmeri (PP348062) | ‘Ca. Phytoplasma asteris’ | 16SrI (99.44) | AF (1.00) |
| 6S-Amaranthus palmeri (PP348063) | ‘Ca. Phytoplasma asteris’ | 16SrI (99.68) | B (0.98) |
| 7S-Echinochloa colona (PP348064) | ‘Ca. Phytoplasma asteris’ | 16SrI (99.60) | AF (1.00) |
| 8S-Micrutalis calva (PP348065) | ‘Ca. Phytoplasma asteris’ | 16SrI (99.52) | AF (1.00) |
| 9S-Balclutha mexicana (PP348066) | ‘Ca. Phytoplasma asteris’ | 16SrI (99.68) | B (0.98) |
yPotential new subgroup.
zNA: does not apply due to the low percentage of identity of the group assigned in iPhyClassifier.
and coefficients of similarity obtained for the phytoplasmas of groups 16SrX and 16SrXIII, this will have to be analyzed with other studies, if we are dealing with new (sub)groups.
The identification of the groups of phytoplasmas obtained here was also corroborated by phylogenetic inferences of the secA and 16Sr genes, as the phylogenies obtained for both genes of the phytoplasmas detected in this study grouped with sequences corresponding to the same 16Sr groups of phytoplasmas previously identified in Mexico (GenBank 16Sr: MN807429) or in other countries (Figure 3).
To our knowledge, this study is the first to document the presence of 16SrIAF, 16SrX and 16SrXIII phytoplasmas in papaya plants with BT in Mexico and Colima. In addition, globally, it is the first time that the presence of the 16SrX has been reported in papaya. In Mexico, the first detection of phytoplasmas in papaya was made by scanning electron microscopy in Baja California Sur (Poghosyan et al., 2004). Subsequently, via molecular biology in Yucatan, Campeche, Quintana Roo (Navarrete-Yabur et al., 2005), Michoacan and Veracruz, where the positive samples were assigned to the subgroup 16SrI-C (Rojas-Martínez et al., 2011).
Along with its detection in Mexico, the group 16SrI has been reported in papaya in Cuba (Acosta et al., 2011; 2017; Acosta-Pérez et al., 2017), Peru (Hodgetts et al., 2009), Sri Lanka (Abeysinghe et al., 2014) and China (Yu et al., 2023), whereas 16SrXIII, only in Brazil (Melo et al., 2013). Other groups of economic importance in papaya are 16SrII (White et al., 1998; Panda et al., 2022), 16SrXII (Gibb et al., 1996; Barbieri et al., 2023), 16SrXV (Wei et al., 2017) and 16SrXVII (Arocha et al., 2005).
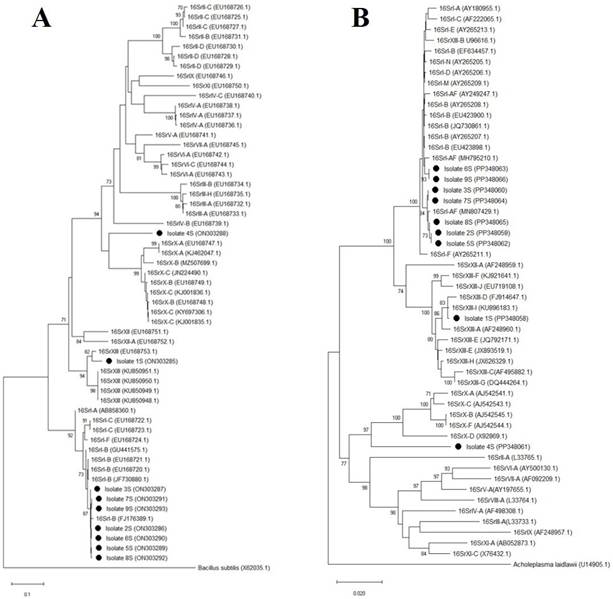

Figure 3 Phylogenetic trees of the secA (A) and 16Sr (B) trees obtained using the neighbor-joining method of 16Sr groups of phytoplasmas identified in papaya agroecosystems in Colima, Mexico (black circles) and from phytoplasma groups from other parts of the world. Accession number are shown in parentheses. Samples 1S-4S: papaya, samples 5S-6S: Amaranthus palmeri, sample 7S: Echinochloa colona, sample 8S: Micrutalis calva, sample 9S: Balclutha mexicana. The gene sequence of secA of Bacillus subtilis and the 16Sr gene sequence of Acholeplasma laidlawii were used as outgroups. In each phylogeny, bootstrap values (from 1000 replicates, greater than 70%) are shown on the branches. The scale bar indicates the number of nucleotide substitutions per site.
In Mexico, the presence of group 16SrX has never been reported (PogoshyanMelkonyan et al., 2019), whereas 16SrXIII has only been associated with potato and tomato plants in Sinaloa (Santos-Cervantes et al., 2010) and Baja California Sur (Holguín-Peña et al., 2007). In the future, whether the low percentage of the detection of phytoplasmas is due to the presence of other organisms such as rickettsiae or viruses that may be inducing the studied symptoms is something that must be determined (Davis et al., 1998; Arocha et al., 2003; Acosta et al., 2013; Luis-Pantoja et al., 2015).
In the case of weeds and insects, the detection of phytoplasmas was positive in three out of 174 (1.7%) and two out of 185 (1.1%) samples, respectively, and with the exception of Ixtlahuacán, the positive samples were obtained regardless of not having recorded phytoplasmas in papaya plants (Table 1). Colima was the only municipality in which there were no weeds nor insects that were positive to phytoplasmas (Table 1).
The weeds in which phytoplasmas were found were asymptomatic, two of which were identified as Amaranthus palmeri (Amaranthaceae) (GenBank 16Sr: PP348062-PP348063, secA: ON303289-ON303290), and the other one, as Echinochloa colona (Poaceae) (GenBank 16Sr: PP348064, secA; ON303291. The insects were identified as Micrutalis calva (Hemiptera: Membracidae) (GenBank 16Sr: PP348065, secA: ON303292) and Balclutha mexicana (Hemiptera: Cicadellidae) (GenBank 16Sr: PP348066, secA: ON303292) (Table 2).
The in silico RFLPs analyses of the 16Sr gene helped assign the phytoplasmas detected to the species ‘Ca. Phytoplasma asteris’ (group 16SrI, percentage of identity: 99.44-99.68), subgroups 16SrI-AF (coefficient of similarity: 1.00) in A. palmeri (sample 5S), E. colona (sample 7S) and M. calva (sample 8S), and 16SrI-B (coefficient of similarity: 0.98) in A. palmeri (sample 6S) and B. mexicana (sample 9S) (Table 2).
The detection of phytoplasmas in insects and weeds related with the papaya agroecosystems suggests that the arthropods spread bacteria between plants and that weeds can play an important part as a source of inoculum (Duduk et al., 2018). For Mexico, this investigation pioneers in questioning whether weeds and insects related to papaya are carriers/sources of phytoplasma inoculum, since it is the first time they are reported as carrying species of the group 16SrI phytoplasmas in papaya agroecosystems. Weeds add to the records of those that have been associated with papaya crops and have been reported as hosts of 16SrII phytoplasmas in Cuba and Ethiopia, such as Anoda acerifolia (Malvaceae), Euphorbia heterophylla (Euphorbiaceae), Malvastrum coromandelianum (Malvaceae) and Rhynchosia minima (Fabaceae) (Arocha et al., 2007; Bekele et al., 2011). Although A. palmeri is a common weed in different Mexican agroecosystems, it has only been reported as a host of phytoplasmas of 16SrIII group phytoplasmas as a weed associated with the cultivation of luffa (Luffa acutangula) (Cucurbitaceae) (Santos-Cervantes et al., 2021).
Regarding insects B. mexicana and M. calva, it is the first time these are recorded as carriers of phytoplasmas in papaya agroecosystems. Previously, the positive detection of phytoplasmas was reported in specimens of the genus Orosius in Australia (Padovan and Gibb, 2001), E. papayae in Cuba (Acosta-Pérez et al., 2010; Acosta et al., 2017) and E. stevensi in Trinidad (Haque and Parasram, 1973). However, to date, only the vectorial capacity of E. stevensi in Trinidad (Haque and Parasram, 1973) and E. papayae in Puerto Rico (Adsuar, 1946) and Cuba (AcostaPérez et al., 2010; Acosta et al., 2017) has been proven.
Regarding Balclutha, in Mexico this genus is part of the entomofauna in Vaccinium corymbosum (Pérez-Mejía et al., 2020), Vitis vinifera (AlmendraPaxtlan et al., 2021), Capsicum annuum (Velásquez-Valle et al., 2018) and Zea mays (Pinedo-Escatel and Moya-Raygoza, 2018) agroecosystems, but neither the genus nor any species have been reported as carriers of phytoplasmas. In other countries, only in specimens of the B. hebe species have phytoplasmas of group 16SrXIII been found in Brassica oleracea (Canale and Bedendo, 2020), 16SrI in Solanum tuberosum (Girsova et al., 2016), 16SrII in Trigonella foenum-graecum (Malik et al., 2020) and 16SrIX in Prunus dulcis (Dakhil et al., 2011).
Regarding Micrutalis, its presence in Mexico has been recorded in amaranth (Amaranthus hypocondriacus) and Prosopis agroecosystems (Salas-Araiza and Boradonenko 2006), and the species M. calva in Z. mays (Pinedo-Escatel, 2014). Micrutalis calva has the vectorial capacity for the Tomato pseudo-curly top virus (TPCTV) in tomato in Florida (Simons and Coe, 1958).
Based on the importance of papaya in Mexico and other countries in which it is planted, it will soon be determined if B. mexicana and M. calva have the capacity to transmit phytoplasmas in papaya and in the weeds that make up its agroecosystem. This study reports for the first time the presence of phytoplasmas 16SrI-AF, 16SrX and 16SrXIII in papaya plants with symptoms of Bunchy Top disease in agroecosystems in the state of Colima, Mexico. The phytoplasmas of groups 16SrX and 16SrXIII are reported for the first time worldwide and in Mexico in papaya, respectively. The weeds Amaranthus palmeri and Echinochloa colona, and the insects Micrutalis calva and Balclutha mexicana, associated with papaya agroecosystems, are carriers of phytoplasma subgroups 16SrI-AF and 16SrI-B, making them new records as natural reservoirs and potential vectors of the bacteria.
Acknowledgments
To the National Agrifood Health, Safety and Quality Service (SENASICA Mexico) for funding this project. To the papaya farmers of the state of Colima for the collection of the samples. To the anonymous reviewers of this manuscript for their valuable comments. Pedro Valadez-Ramírez thanks the Colegio de Postgraduados for the support provided to carry out his doctoral studies.
REFERENCES
Abeysinghe, S., Kumari, W.G.S.M., Arachchi, I.M.M., Dickinson, M. (2014). First report of the molecular identification of a phytoplasma associated with dieback disease of papaya in Sri Lanka. New Disease Reports, 29, 13. https://doi.org/10.5197/j.2044-0588.2014.029.013. [ Links ]
Acosta, K.I., Zamora, L., Piñol, B., Quiñones, M.L., Ramos, P.L., Luis, M., Leyva, N.E., Arocha, Y. (2017). Empoasca papayae Oman, 1937 (Hemiptera: Cicadellidae) the simultaneous vector of phytoplasmas and rickettsia associated with “Bunchy Top Symptom” in Cuba. Annals of Biology, 39, 35-42. https://doi.org/10.6018/analesbio.39.03. [ Links ]
Acosta, K., Zamora, L., Fernández, A., Arocha, Y., Martínez, Y., Santos, M.E., Méndez, J., Chávez, A., Leyva, N.E. (2011). First report of ‘Candidatus Phytoplasma asteris’ (group 16SrI) affecting papaya in Cuba. New Disease Reports, 24, 29. https://doi.org/10.5197/j.2044-0588.2011.024.029. [ Links ]
Acosta, K., Zamora, L., Piñol, B., Fernández, A., Chávez, A., Flores, G., Méndez, J., Santos, M.E., Leyva, N.E., Arocha, Y. (2013). Identification and molecular characterization of phytoplasmas and rickettsia pathogens associated with ‘Bunchy top symptom’ (BTS) and ‘Papaya bunchy top’ (PBT) of papaya in Cuba. Crop Protection, 45, 49-56. https://doi.org/10.1016/j.cropro.2012.11.016. [ Links ]
Acosta-Pérez, K.I., Piñol-Pérez, B.E., Zamora-Gutiérrez, L., Quiñones-Pantoja, M.I., Miranda-Cabrera, I., Leyva-López, N.E., Arocha-Rosete, Y. (2017). A phytoplasma representative of a new subgroup 16SrI-Z associated with Bunchy Top Symptoms (BTS) on papaya in Cuba. Revista de Protección Vegetal, 32, 52-59. [ Links ]
Acosta-Pérez, K., Piñol, B., Arocha-Rosete, Y., Wilson, M., Boa, E., Lucas, J. (2010). Transmission of the phytoplasma associated with bunchy top symptom of papaya by Empoasca papaya Oman. Journal of Phytopathology, 158, 194-196. https://doi.org/10.1111/j.1439-0434.2009.01590.x. [ Links ]
Adsuar, J. (1946). Transmission of papaya bunchy top by a leafhopper of the genus Empoasca. Science, 103, 316. https://doi.org/10.1126/science.103.2671.316. [ Links ]
Ahrens, U., Seemüller, E. (1992). Detection of DNA of plant pathogenic mycoplasma-like organisms by a polymerase chain reaction that amplifies a sequence of the 16S rRNA gene. Phytopathology, 82, 828-832. https://doi.org/10.1094/Phyto-82-828. [ Links ]
Almendra-Paxtlan, L., García-Martínez, O., Robles-Hernández, V.E., Sánchez-Peña, S.R. (2021). Cicadomorpha in a vineyard at Parras Coahuila, Mexico, and vectors of diseases. Southwestern Entomologist, 46, 147-152. https://doi.org/10.3958/059.046.0114. [ Links ]
Arocha, I., Horta, D., Peralta, E., Jones, P. (2003). First report on molecular detection of phytoplasmas in papaya in Cuba. Plant Disease, 87, 1148. https://doi.org/10.1094/PDIS.2003.87.9.1148C. [ Links ]
Arocha, Y., López, M., Piñol, B., Fernández, M., Picornell, B., Almeida, R., Palenzuela, I., Wilson, M.R., Jones, P. (2005). ‘Candidatus Phytoplasma graminis’ and ‘Candidatus Phytoplasma caricae’, two novel phytoplasmas associated with diseases of sugarcane, weeds and papaya in Cuba. International Journal of Systematic and Evolutionary Microbiology, 55, 2451-2463. https://doi.org/10.1099/ijs.0.63797-0. [ Links ]
Arocha, Y., Piñol, B., López, M., Miranda, I., Almeida, R., Wilson, M., Jones, P. (2007). Bunchy top symptom of papaya in Cuba: new insights. Bulletin of Insectology, 60, 393-394. [ Links ]
Barbieri, C., Contaldo, N., Sciovolone, A., Bertaccini, A. (2023). Phytoplasmas in papaya: detection and identification. Phytopathogenic Mollicutes, 13, 145-146. https://doi.org/10.5958/2249-4677.2023.00073.7. [ Links ]
Bekele, B., Hodgetts, J., Tomlinson, J., Boonham, N., Nikolić Swarbrick, P., Dickinson, M. (2011). Use of a real-time LAMP isothermal assay for detecting 16SrII and XII phytoplasmas in fruit and weeds of the Ethiopian Rift Valley. Plant Pathology, 60, 345-355. https://doi.org/10.1111/j.1365-3059.2010.02384.x. [ Links ]
Canale, M.C., Bedendo, I.P. (2020). Report of 'Candidatus Phytoplasma hispanicum' (16SrXIII-E) associated with cauliflower stunt in São Paulo State, Brazil, and Balclutha hebe as its potential vector. Plant Disease, 104, 967. https://doi.org/10.1094/PDIS-10-19-2111-PDN. [ Links ]
COEPAPAYA, Consejo Estatal de Productores de Papaya de Colima A.C. (2017). Calidad y mejora continua ante un mercado internacional exigente. Red Innovagro. https://www.redinnovagro.in/casosexito/2017/Papaya_COEPAPAYA.pdf (consulta, agosto 2020). [ Links ]
Cook, M.T. (1931). New virus diseases of plants in Porto Rico. The Journal of the Department of Agriculture of Porto Rico, 15, 193-195. https://doi.org/10.46429/jaupr.v15i2.14232. [ Links ]
Dakhil, H.A., Hammad, E.A.F., El-Mohtar, C., Abou-Jawdah, Y. (2011). Survey of leafhopper species in almond orchards infected with almond witches'-broom phytoplasma in Lebanon. Journal of Insect Science, 11, 60. https://doi.org/10.1673/031.011.6001. [ Links ]
Davis, M., Ying, Z., Brunner, B., Pantoja, A., Fewerda, F. (1998). Rickettsial relative associated with papaya bunchy top disease. Current Microbiology, 36, 80-84. https://doi.org/10.1007/s002849900283. [ Links ]
Deng, S, Hiruki, C (1991). Amplification of 16S rRNA genes from culturable and nonculturable mollicutes. Journal of Microbiological Methods, 14, 53-61. 10.1016/0167-7012(91)90007-D. [ Links ]
Dickinson, M, Hodgetts, J (2013). PCR analysis of phytoplasmas based on the secA gene. In: Dickinson M and Hodgetts J (eds.), Phytoplasma: Methods and Protocols, Methods in Molecular Biology, Vol. 938, Humana Press, 205-216. 10.1007/978-1-62703-089-2_17. [ Links ]
Duduk, B, Stepanoviç, J, Yadav, A, Rao, GP (2018). Phytoplasmas in weeds and wild plants. In: Rao G, Bertaccini A, Fiore N, Liefting L (eds.), Phytoplasmas: Plant Pathogenic Bacteria - I, Springer, 313-345. 10.1007/978-981-13-0119-3_11. [ Links ]
Folmer, O, Black, M, Hoeh, W, Lutz, R, Vrijenhoek, R (1994). DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology, 3, 294-299. [ Links ]
Girsova, NV, Bottner-Parker, KD, Bogotdinov, DZ, Meshkov, YI, Mozhaeva, KA, Kastalyeva, TB, Lee, IM (2016). Diverse phytoplasma associated with potato stolbur and other related potato diseases in Russia. European Journal of Plant Pathology, 145, 139-153. 10.1007/s10658-015-0824-3. [ Links ]
Gundersen, DE, Lee, I-M (1996). Ultrasensitive detection of Phytoplasmas by nested-PCR assays using two universal primer pairs. Phytopathologia Mediterranea, 35, 144-151. [ Links ]
Haque, SQ, Parasram, S (1973). Empoasca stevensi, a new vector of bunchy top disease of papaya. Plant Disease Reporter, 57, 412-413. [ Links ]
Hodgetts, J, Chuqillangui, C, Muller, G, Arocha, Y, Gamarra, D, Pinillos, O, Velit, E, Lozada, P, Boa, E, Boonham, N, Mumford, R, Barker, I, Dickinson, M (2009). Surveys reveal the occurrence of phytoplasmas in plants at different geographical locations in Peru. Annals of Applied Biology, 155, 15-27. 10.1111/j.1744-7348.2009.00316.x. [ Links ]
Holguín-Peña, RJ, Vázquez-Juárez, RC, Martínez-Soriano, JP (2007). First report of a 16SrI-B group phytoplasma associated with a yellows-type disease affecting tomato plants in the Baja California Peninsula of Mexico. Plant Disease, 91, 328. 10.1094/PDIS91-3-0328B. [ Links ]
Kazeem, SA, Inaba, J, Zhao, Y, Zwolinska, A, Ogunfunmilayo, AO, Arogundade, O, Wei, W (2021). Molecular identification and characterization of ‘Candidatus Phytoplasma convolvuli’-related strains (representing a new 16SrXII-O subgroup) associated with papaya bunchy top disease in Nigeria. Crop Protection, 148, 105731. 10.1016/j.cropro.2021.105731. [ Links ]
Kumar, S, Stecher, G, Li, M, Knyaz, C, Tamura, K (2018). MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35, 1547-1549. 10.1093/molbev/msy096. [ Links ]
Lee, I-M, Gundersen-Rindal, DE, Davis, RE, Bartoszyk, IM (1998). Revised classification scheme of phytoplasmas based on RFLP analyses of 16S rRNA and ribosomal protein gene sequences. International Journal of Systematic Bacteriology, 48, 1153-1169. 10.1099/00207713-48-4-1153. [ Links ]
Lee, I, Hammond, RW, Davis, RE, Gundersen, DE (1993). Universal amplification and analysis of pathogen 16S rDNA for classification and identification of mycoplasmalike organisms. Phytopathology, 83, 834-842. 10.1094/Phyto-83-834. [ Links ]
Luis-Pantoja, M, Ramos-González, PI, Naranjo, M, Hernández-Rodríguez, L, Rodríguez, J, Pérez-López, E (2015). Rickettsia-related bacteria associated with papaya plants showing bunchy top disease in Cuba. Journal of General Plant Pathology, 81, 166-168. 10.1007/s10327-014-0571-x. [ Links ]
Malik, ST, Ahmad, JN, Sharif, MZ, Trebicki, P, Tahir, M, Bertaccini, A (2020). Molecular detection and characterisation of phytoplasma in Trigonella foenum-graecum and identification of potential insect vectors in Punjab, Pakistan. Pakistan Journal of Botany, 52, 1605-1613. 10.30848/PJB2020-5(16). [ Links ]
Martorell, LF, Adsuar, J (1952). Insects associated with papaya virus diseases in the Antilles and Florida. The Journal of Agriculture of the University of Puerto Rico, 34, 319-329. 10.46429/jaupr.v36i4.12786. [ Links ]
Melo, L, Silva, E, Flôres, D, Ventura, J, Costa, H, Bedendo, I (2013). A phytoplasma representative of a new subgroup, 16SrXIII-E, associated with papaya apical curl necrosis. European Journal of Plant Pathology, 137, 445-450. 10.1007/s10658-013-0267-7. [ Links ]
Navarrete-Yabur, A, González-Losa, M, Moreno-Valenzuela, O (2005). Detection and identification of phytoplasma in cultivated and wild papaya plants in the Yucatan peninsula. Phytopathology, 95, S74. [ Links ]
Oxelman, B, Lidén, M, Berglund, D (1997). Chloroplast rps16 intron phylogeny of the tribe Sileneae (Caryophyllaceae). Plant Systematics and Evolution, 206, 393-410. 10.1007/BF00987959. [ Links ]
Padovan, AC, Gibb, KS (2001). Epidemiology of phytoplasma diseases in papaya in Northern Australia. Journal of Phytopathology, 149, 649−658. 10.1046/j.1439-0434.2001.00688.x. [ Links ]
Panda, P, Singh, SK, Mall, S, Rao, GP (2022). Multilocus gene-based characterization of peanut witches’ broom related phytoplasma strain associated with flower malformation of papaya in India. Indian Phytopathology, 75, 1211−1216. 10.1007/s42360-022-00547-z. [ Links ]
Pérez-Mejía, FA, Ortega-Arenas, LD, Bautista-Martínez, N, Blanco-Rodríguez, E, López-Buenfil, JA (2020). Cicadélidos asociados a arándano en Jalisco, México. Southwestern Entomologist, 45, 275-288. 10.3958/059.045.0129. [ Links ]
Pinedo-Escatel, JA (2014). Abundancia y riqueza de membracoideos (Hemiptera: Auchenorrhyncha) en maíz durante la estación lluviosa en Las Agujas, Zapopan, Jalisco. Dugesiana, 21, 49-53. [ Links ]
Pinedo-Escatel, JA, Moya-Raygoza, G (2018). Diversity of leafhoppers (Hemiptera: Cicadellidae) associated with border grasses and maize during the wet and dry seasons in Mexico. Environmental Entomology, 47, 282-291. 10.1093/ee/nvx204. [ Links ]
Poghosyan, AV, Lebsky, VK, Arce-Montoya, M, Landa, L (2004). Possible phytoplasma disease in papaya (Carica papaya L.) from Baja California Sur: diagnosis by scanning electron microscopy. Journal of Phytopathology, 152, 376-380. 10.1111/j.1439-0434.2004.00847.x. [ Links ]
Pogoshyan-Melkonyan, A, Lebsky, V, Rojas-Martínez, RI, Oropeza-Salín, C, Narváez-Cab, M, Córdova-Lara, I (2019). Fitoplasmas y su estudio en México. Pp. 30-70. In: Álvarez-Sánchez, F. J, Rodríguez-Guzmán, P, Alarcón, A (coord.). Biodiversidad de microorganismos de México. Importancia, aplicación y conservación. UNAM: México. 682p. [ Links ]
Rojas-Martínez, RI, Zavaleta-Mejía, E, Rivas-Valencia, P (2011). Presencia de fitoplasmas en papayo (Carica papaya) en México. Revista Chapingo. Serie Horticultura, 17, 47-50. [ Links ]
Salas-Araiza, MD, Boradonenko, A (2006). Insectos asociados al amaranto Amaranthus hypocondriacus L. (Amaranthaceae) en Irapuato, Guanajuato, México. Acta Universitaria, 16, 50-55. [ Links ]
Sambrook, J, Russell, D (2001). Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, USA. 1659p. [ Links ]
Santos-Cervantes, ME, Camacho-Bojórquez, JE, Escobedo-Rivera, UR, Méndez-Lozano, J, Leyva-López, NE (2021). Molecular characterization of group 16SrI and 16SrIII phytoplasmas associated with loofah witches’ broom disease in Sinaloa, Mexico. Canadian Journal of Plant Pathology, 43, 366-373. 10.1080/07060661.2020.1817794. [ Links ]
Santos-Cervantes, ME, Chávez-Medina, JA, Acosta-Pardini, J, Flores-Zamora, GL, Méndez-Lozano, J, Leyva-López, NE (2010). Genetic diversity and geographical distribution of phytoplasmas associated with potato purple top disease in Mexico. Plant Disease, 94, 388-395. 10.1094/PDIS-94-4-0388. [ Links ]
Schneider, B, Seemüller, E, Smart, CD, Kirkpatrick, BC (1995). Phylogenetic classification of plant pathogenic mycoplasma-like organisms or phytoplasmas. Pp. 369-380. In: Razin, S, Tully, JG (eds.). Molecular and Diagnostic Procedures in Mycoplasmology. Academic Press, San Diego. 466p. 10.1016/B978-012583805-4/50040-6. [ Links ]
SIAP (2020). Atlas agroalimentario 2020. Servicio de Información Agroalimentaria y Pesquera. https://www.gob.mx/siap/ (consulta, septiembre 2020). [ Links ]
Simons, JN, Coe, DM (1958). Transmission of pseudo-curly top virus in Florida by a treehopper. Virology, 6, 43-48. 10.1016/0042-6822(58)90058-8. [ Links ]
Stillson, PT, Szendrei, Z (2020). Identifying leafhopper targets for controlling aster yellows in carrots and celery. Insects, 11, 411. 10.3390/insects11070411. [ Links ]
Story, GE, Halliwell, RS (1969). Association of a mycoplasma-like organism with the bunchy top disease of papaya. Phytopathology, 59, 1336-1337. [ Links ]
Velásquez-Valle, R, Reveles-Torres, LR, Fernández-Martínez, M (2018). Diversity of leafhoppers (Homoptera: Cicadellidae) collected in chili parcels in northern-central Mexico. Revista Mexicana de Ciencias Agrícolas, 9, 1289-1293. [ Links ]
Wei, W, Pérez-López, E, Davis, RE, Bermúdez-Díaz, L, Granda-Wong, C, Wang, J, Zhao, Y (2017). ‘Candidatus Phytoplasma brasiliense’-related strains associated with papaya bunchy top disease in northern Peru represent a distinct geographic lineage. Crop Protection, 92, 99−106. 10.1016/j.cropro.2016.10.024. [ Links ]
White, DT, Blackall, LL, Scott, PT, Walsh, KB (1998). Phylogenetic positions of phytoplasmas associated with dieback, yellow crinkle and mosaic diseases of papaya, and their proposed inclusion in ‘Candidatus Phytoplasma australiense’ and a new taxon, ‘Candidatus Phytoplasma australasia’. International Journal of Systematic Bacteriology, 48, 941−951. 10.1099/00207713-48-3-941. [ Links ]
Yu, SS, Zhu, AN, Song, WW (2023). Carica papaya represents a new host of 16SrI-B subgroup phytoplasma associated with yellow symptoms in China. Plant Disease, 107, 211. 10.1094/PDIS-03-22-0692-PDN. [ Links ]
Received: March 05, 2024; Accepted: May 19, 2024











 texto en
texto en 


