Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Accesos
Accesos
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO
Compartir
Revista mexicana de fitopatología
versión On-line ISSN 2007-8080versión impresa ISSN 0185-3309
Rev. mex. fitopatol vol.42 no.2 Texcoco may. 2024 Epub 24-Feb-2025
https://doi.org/10.18781/r.mex.fit.2311-2
Phytopathological Note
Genetic variability of two Mexican Tomato brown rugose fruit virus isolates and expression of symptoms in tomato and pepper
11Departamento de Fitotecnia, Colegio de Postgraduados, Km 36.5 Carretera México-Texcoco, Montecillo, Texcoco, Edo. México, México CP 56230.
22Fitosanidad, Colegio de Postgraduados, Km 36.5 Carretera México-Texcoco, Montecillo, , Texcoco, Edo. México, México CP 56230.
33Preparatoria Agrícola, Universidad Autónoma Chapingo, Km 38.5 Carretera México-Texcoco, , Texcoco, Edo. México, México, C.P. 56230
44Departamento de Parasitología Agrícola, Universidad Autónoma Chapingo, Km 38.5 Carretera México-Texcoco, , Texcoco, Edo. México, México, C.P. 56230;
Objective/Background.
The objective was to analyze the variability of two Mexican isolates of ToBRFV after a process of inoculation and multiplication in different commercial and Mexican landraces of tomato (Solanum lycopersicum) (15 materials) and pepper (Capsicum annuum) (20 materials), and to evaluate the expression of symptoms under greenhouse conditions.
Materials and Methods.
In greenhouses, the post-infection variability of two isolates was analyzed: EM-JI2021 (State of Mexico) and C-JI2021 (Colima) in 15 genotypes of tomato and 20 of pepper. Each isolate was mechanically inoculated on five plants per genotype with a total of 150 plants (56 days old) of tomato and 200 of pepper. Three plants per genotype were used as controls. Sixty-one days after inoculation, one leaf per plant was collected for RT-PCR. Incidence and symptom expression were recorded. RNA extraction was by 2% CTAB. ToBRFV-F/ ToBRFV-R primers amplifying 475 bpb of the RpRd gene were used (SENASICACNRF). 24 RT-PCR products were sequenced, cleaned and aligned with NCBI Genbank records using MEGAv11.0.13. Based on epidemiological criteria, 34 sequences were selected from GenBank for variability analysis.
Results.
Ten days after inoculation, tomato genotypes exhibited severe mosaic, mild mosaic, and reduced leaf area. In pepper, symptoms differentiated by genotype were observed, including hypersensitivity reaction, leaf deformation, stem necrosis, mosaic, yellowing, necrotic lesions, and asymptomatic condition. Between position 2,124 to 2,500 bp there was 99.74 % homology with the first report of ToBRFV in Jordan (KT383474.1). Homology >99.74 % was found with isolates from USA (MT002973.1) and Canada (OQ674195.1). C-JI2021 exhibited no variability, while EM-JI2021 generated three haplotypes: One nucleotide change (c.2,355T>C) was detected in Mulato (pepper) and Don R (tomato), while two substitutions (c.2,278A>T; c.2,355T>C) were detected in Santawest, Altius, Sahariana and Nebula (tomato).
Conclusion.
The pathogenic intensity of ToBRFV varied from asymptomatic to severe depending on the combination of host, genotype, and haplotype. In short periods of infection, three haplotypes were detected, suggesting host-dependent mutagenic capacity of the virus.
Keywords: Solanum lycopersicum; Capsicum annuum; ToBRFV; isolates.
Objetivo/Antecedentes.
El objetivo fue analizar la variabilidad de dos aislados mexicanos de ToBRFV posterior a un proceso de inoculación y multiplicación en diferentes variedades comerciales y criollos mexicanos de jitomate (Solanum lycopersicum) (15 materiales) y chile (Capsicum annuum) (20 materiales), y evaluar la expresión de síntomas en condiciones de invernadero.
Materiales y métodos.
En invernadero, se analizó la variabilidad postinfección de dos aislados: EM-JI2021 (Edo. de México) y C-JI2021 (Colima) en 15 genotipos de jitomate y 20 de chile. Cada aislado se inoculó mecánicamente en cinco plantas por genotipo con un total de 150 plantas (56 días de edad) de jitomate y 200 de chile. Se emplearon tres plantas por genotipo como testigos. Sesenta y un días después de inoculación se colectó una hoja por planta para RT-PCR. Se registró incidencia y expresión de síntomas. La extracción de ARN fue por CTAB 2 %. Se utilizó oligos ToBRFV-F/ToBRFV-R que amplifican 475pb del gen RpRd (SENASICA-CNRF). Se secuenciaron 24 productos RT-PCR, se limpiaron y alinearon con registros del Genbank NCBI mediante MEGAv11.0.13. Con criterio epidemiológico, se seleccionaron 34 secuencias del GenBank para análisis de variabilidad.
Resultados.
Diez días después de la inoculación, los genotipos de jitomate exhibieron mosaico severo, leve, reducción del área foliar. En chile se observaron síntomas diferenciados por genotipo, incluyendo reacción de hipersensibilidad, deformación foliar, necrosis en tallo, mosaico, amarillamiento, lesiones necróticas y condición asintomática. Entre la posición 2,124 al 2,500 pb se tuvo 99.74 % de homología con el primer reporte de ToBRFV en Jordania (KT383474.1). Se encontró homología >99.74 % con aislados de USA (MT002973.1) y Canadá (OQ674195.1). C-JI2021 no exhibió variabilidad, mientras que EM-JI2021 generó tres haplotipos: En Mulato (chile) y Don R (jitomate) se detectó cambio de un nucleótido (c.2,355T>C), mientras que en Santawest, Altius, Sahariana y Nebula (jitomate) se detectaron dos sustituciones (c.2,278A>T; c.2,355T>C).
Conclusión.
La intensidad patogénica de ToBRFV varió de asintomática a severa según combinación de hospedero, genotipo y haplotipo. En periodos cortos de infección se detectaron tres haplotipos lo que sugiere capacidad mutagénica del virus en función del hospedero.
Palabras clave: Solanum lycopersicum; Capsicum annuum; ToBRFV; aislados.
Introduction
In Jordan, in 2015, reports were made of symptoms of brown roughness in tomato fruits (Solanum lycopersicum), with a 100% incidence at a greenhouse level. Etiological studies determined a new Tobamovirus called Tomato brown rugose fruit virus (ToBRFV) (Salem et al., 2016). It has been postulated that ToBRFV may have come from the recombination of 314 nucleotides of the region 534-848 of the replicase gene (Maayan et al., 2018). The Tomato mottle mosaic virus (ToMMV) and the Tobacco mosaic virus (TMV) have been proposed as direct ancestors, with the latter being the main one (Salem et al., 2016). The ToBRFV genome from different countries usually display low variability, which suggests an evolutionary process from a common descendant (Salem et al., 2016). In addition, Maayan and collabrators (2018), through phylogenetic studies of Tobamovirus, concluded that ToBRFV has undergone a divergent evolutionary process with adaptation to different hosts, but with a low mutation rate over a period of 3-4 years. In general, viruses have the potential to evolve and adapt quickly under the pressure of natural selection due to high population rates resulting from the efficient intraspecies replication, the occurrence of quasispecies and the lack of genome correction mechanisms (RNA virus), which enables genetic variation and short generation times (Hanssen et al., 2010).
In Mexico, the ToBRFV was first reported in 2018 (Cambrón-Crisantos et al., 2018). This Tobamovirus is spread mainly seed-borne. Currently, it has been reported in 20 states of the country, including the main tomato (Solanum lycopersicum) and pepper (Capsicum annuum) producing entities (Camacho-Beltran et al., 2019; Cambrón-Crisantos et al., 2018). Fruits on tomato are associated with a yellow coloration, green spot and deformation, streaking and irregular brown spots.
Mosaic, mottling and yellowing are observed on foliage. The first reports of viruses included total loss of production (Cambrón-Crisantos et al., 2018). Due to this, the aim of this study was to analyze the variability of two Mexican ToBRFV isolates after an inoculation and multiplication process in different commercial and native Mexican tomato (S. lycopersicum) (15 materials) and pepper (C. annuum) varieties (20 materials), and to evaluate the expression of symptoms under greenhouse conditions until the flowering stage.
Inoculation of ToBRFV. Two isolates of the virus from the State of Mexico (EMJI2021) and Colima (C-JI2021) were used. The isolated were inoculated in a total of 15 tomato genotypes, 13 commercials and two Mexican ones, along with 20 of C. annuum, four varieties, 15 genotypes and one native. The isolates were performed under greenhouse conditions on 56-day-old plants. Before inoculation, Imidacloprid (1.5 mL L-1) was applied as a preventive measure against vector insects. The inoculation was carried out on the second youngest leaf, by spraying carborondum followed by the phosphates buffer with the macerated tomato tissue infected with the virus (two isolates). Five plants were considered for every genotype, 150 tomato plants (for both isolations) and 200 pepper plants. The control plants (three plants per plant material) were isolated to avoid contamination. The variables evaluated were the incidence of the virus in plants and the type of symptoms in the tomato and pepper materials was recorded. Tissue was taken from young leaves 61 days after inoculation, it was photographed and preserved at -20 °C. The tissue was macerated with liquid nitrogen for molecular study.
RNA extraction and RT-PCR. The nucleic acids were extracted using the CTAB 2% method (Yu, 2012; modified by CP-LANREF, 2021). The concentrations and characteristics of the RNA were measured in the NanoDrop 2000 (Thermo Fisher Scientific 2000, USA). For the RT-PCR, primers ToBRFV-F 5-AACCAGAGTCTTCCTATACTGGGAA-3 and ToBRFV-R 5’CTCWCCATCTCTTAATAATCTCCT-3 were used, which amplify part of the small subunit of the RpRd replicase with 475 bp (SENASICA, 2018). Retrotranscription and the polymerase chain reaction (RT-PCR) were carried out in the T-100 (BioRad) thermocycler. For the retrotranscription (RT), primer R (10uM), water and 2 µL of RNA (50 ng µL-1) were used to obtain a volume of 16.375 µL and it was incubated at 85 °C for 3 min. Later, to each previous reaction, the mixture of Buffer-RT (5X), dNTPs (10mM), RNAsin (40 U µL-1) and M-MLV-RT (200 U µL-1) was added, with a volume of 8.625 µL. The mixture was incubated at 44 °C for 60 min and 92 °C for 10 min. For the PCR, MgCl2, dNTPs, the oligos, Taq polymerase, nuclease-free water and cDNA were used in a final volume of 25 µL. The conditions were an initial denaturalization at 98 °C for 90 s, denaturalization at 98 °C for 10 s, alignment at 55 °C for 20 s, extension at 72 °C for 40 s, final extension at 72 °C for 5 min, and finally, 72 °C (SENASICA, 2018). The PCR products were analyzed by electrophoresis in agarose gel at 1% stained with ethidium bromide and they were viewed under UV light in a photodocumenter (UVP, Biolmaging Systems, Epi Chemi II Darkroom).
Phylogenetic analysis. PCR products (24 samples) were sent to Macrogen® (Seoul, Korea) to be sequenced. The sequences (both ways) were cleaned and the ends were eliminated using the program SeqAssem (https://science.do-mix.de/ software_seqassem.php). A consensus was made using the sequences to identify and compare the homology with sequences from the Genbank of the National Center for Biotechnology Information (NCBI). Sequences were chosen from complete ToBRFV genomes (three sequences) from Mexico and other countries (30 sequences) from the Genbank to perform the alignment using sequences of the fragment from the gene used in this study. The alignment was carried out using Mega 11.0.13 and Geneious 2023.0.4 (www.geneious.com) to determine the variability between sequences. The criterion for the selection of the viral sequences was according to the first reports of the virus, worldwide diversity, as well as sequences from the American continent (Table 1).
ToBRFV-related symptoms in tomato and pepper. All the commercial and native tomato material expressed symptoms 10 days after the inoculation of isolations EM-JI2021 and C-JI2021. The symptoms observed in tomato induced by ToBRFV were mainly mild to severe mosaic. A severe leaf deformation and reduction with remanence of veins was also observed (Table 2 and Figure 1). In some varieties, the clearing of veins was also observed.
Zhi-Yong et al. (2021) evaluated 50 tomato cultivars and no material expressed resistance to ToBRFV, showing different symptoms such as mild to severe mosaic, the formation of blisters on the leaves, necrosis in sepals and pedicels, deformation, and in fruits, yellow spots, as well as necrotic lesions with brown roughness. They also inoculated C. annumm, Nicotiana benthamiana, N. tabacum, Solanum melongena and S. tuberosum cv. Kexin 1, where symptoms of necrosis in inoculated leaves and stems, as well as dwarfism.
In pepper, all materials displayed a diversity of symptoms after inoculation, particularly a reaction of hypersensitivity with the falling of inoculated leaves. It was initially observed in necrotic inoculated lesions and later, the detaching of the leaf. Severe symptoms were observed in some materials such as deformation in apex leaves, stem necrosis, necrosis in veins and necrotic lesions in non-inoculated leaves (Table 2, Figure 2). These symptoms coincide with Fidan et al. (2021), who observed necrosis in inoculated leaves, necrotic lesions in the stem and the yellowing of leaves.
Table 1 Complete Tomato brown rugose fruit virus sequences obtained from the GenBank (NCBI) used for the alignment and compares with ToBRFV sequences from the study.
| No. of accession | Base pairs | Host | Plant tissue | Location |
|---|---|---|---|---|
| MZ945420.1 | 6379 | Solanum lycopersicum | Belgium | |
| OQ674194.1 | 6374 | S. lycopersicum | Canada | |
| OQ674195.1 | 6242 | S. lycopersicum cultivar Yari | Canada | |
| OM515230.1 | 6375 | S. lycopersicum | Netherlands | |
| OM515231.1 | 6373 | S. lycopersicum | United Kingdom | |
| OM515232.1 | 6373 | S. lycopersicum | United Kingdom | |
| MZ323110.1 | 6394 | S. lycopersicum | Jordan | |
| MK648157.1 | 6388 | Capsicum annuum | Jordan | |
| KT383474.1 | 6393 | S. lycopersicum | Jordan | |
| MN882030.1 | 6379 | S. lycopersicum | Egypt | |
| MN882031.1 | 6379 | S. lycopersicum | Egypt | |
| MW349655.1 | 6379 | C. annuum cultivar Tampiqueno | Mexico | |
| OM515233.1 | 6369 | S. lycopersicum | Seeds | Peru |
| OM515258.1 | 6376 | S. lycopersicum | Seeds | Peru |
| OM515256.1 | 6361 | S. lycopersicum | Seeds | China |
| MT018320.1 | 6392 | S. lycopersicum | China | |
| OM515237.1 | 6377 | S. lycopersicum | Seeds | Israel |
| OM515234.1 | 6371 | C. annuum | Seeds | Israel |
| OM515257.1 | 6367 | S. lycopersicum | Seeds | Israel |
| OM515266.1 | 6364 | S. lycopersicum | Seeds | Israel |
| OM515250.1 | 6371 | S. lycopersicum | Seeds | Israel |
| OM515261.1 | 6376 | S. lycopersicum | Fruit | Belgium |
| OM515265.1 | 6375 | S. lycopersicum | Fruit | Belgium |
| MN815773.1 | 6354 | S. lycopersicum | Greece | |
| OM305070.1 | 6386 | S. lycopersicum | Switzerland | |
| MT107885.1 | 6386 | S. lycopersicum | Turkey | |
| OM515234.1 | 6371 | C. annuum | Seeds | Israel |
| OM782671.1 | 6356 | S. lycopersicum | Mexico | |
| OM892675.1 | 6384 | S. lycopersicum | Imported fruit | Mexico |
| OM892676.1 | 6381 | S. lycopersicum | Seeds | Peru |
| OM892677.1 | 6392 | S. lycopersicum | Store fruit | USA |
| OM892678.1 | 6393 | S. lycopersicum | Imported fruit | Peru |
| OM892679.1 | 6375 | S. lycopersicum | Store fruit | USA |
| OM892681.1 | 6357 | S. lycopersicum | Leaf | USA |
Variability in ToBRFV isolates. Out of the total of tomato and pepper genotypes inoculated, five pepper samples were sent to inoculate, one with C-JI2021 and four with EM-JI2021, along with 19 tomato samples, six with EM-JI2021 and 13 with C-JI2021. The sequences in the study were aligned with 80.2% of the total of the fragment of the partial RpRd gene (475 pb). The sequences displayed an identity with accessions of the GenBank with a homology that fluctuates between 99.74 and 100%, including the sequence of the virus originally described by Salem et
Table 2 Symptoms pf Tomato brown rugose fruit virus in commercial and native tomato and pepper materials expressed under greenhouse conditions.
| Name | Crop | Symptoms | Name | Crop | Symptoms |
|---|---|---|---|---|---|
| Santawest | Tomatoy | MM | Conga | Pepperz | A |
| Citali | Tomato | MM, LF, LR | Fascinato | Pepper | A |
| IR143466 | Tomato | SM, LF, LR | Felicitas | Pepper | NLNI |
| Sicilia | Tomato | SM, LF, LR | Botaron | Pepper | A, SN |
| Sahariana | Tomato | SM, LF, LR | Godzilla | Pepper | NLNI |
| Altius | Tomato | SM, LF, LR, GI SM, LF, LR, CV | Kathia | Pepper | A |
| Don R | Tomato | MS, LF, LD | Almuden | Pepper | LD, NLNI, DAA, SN, NLN |
| Nebula | Tomato | MM, LF | Bachia | Pepper | LNI |
| Volcano | Tomato | MM, LF, CV | Cannon | Pepper | A |
| Ametrino | Tomato | MM, LF, CV | Gina | Pepper | NLNI, SN |
| Angelle | Tomato | SM, LF, LD, CV | Confidaro | Pepper | A |
| Olmeca | Tomato | SM, LF, LR | |||
| UAM-X | Tomato | SM, LF, LR, CV | Cayenne | Pepper | DAA, NL |
| Rio Grande | Tomato | SM, LF, CV | |||
| Totolapan, Mor. | Tomato | MM | mulato | Pepper | A |
| Cavanna | Pepper | A | Serrano | Pepper | SM, GI |
| Shir | Pepper | A | Zongolica | Pepper | MM, CR |
| Orangela | Pepper | A | Chile pasilla | Pepper | A |
| Manzano-Ver | Pepper | A |
yTomato. MM: mild mosaic; SM: Severe mosaic; LF: Leaf deformation; LD: Leaf deformation; LR: Leaf reduction; CV: Clearing of veins; GI: Green islands.
zPepper: A: Asymptomatic; MM: mild mosaic; SM|: Severe mosaic; LD: Leaf deformation; NL: necrotic lesions on leaves; NLNI: necrotic lesions on non-inoculated leaves; SN: Stem necrosis; DAA: Deformation of apical area; NLN: necrosis in leaf nervations; GI: Green islands; CR: Concentric rings.
al. (2016) (Table 3, Figure 3). The sequences that displayed an identity of 99.74 and 99.75% were samples inoculated with EM-JI2021 (Table 3). All the sequences were aligned with accessions from the USA and Canada (Table 3). Five sequences with an identity of less than 100% were aligned with OQ6741195.1 from Canada and one sequence from the USA (MT002973.1).
According to the analysis of the alignment based on the complete genome with 34 ToBRFV sequences and the partial genome of the replicase of the 24 sequences of this investigation, variability was observed at the nucleotide level in six sequences obtained from plants inoculated with EM-JI2021. Two changes were observed: in nucleotides c.2,278A>T and in c.2355T>C (Figure 3). The sequences under study were aligned between the positions 2,124 and 2,500 corresponding to the gene of the replicase. The nucleotide substitutions were also observed in accessions MW349655.1 (pepper tissue), OQ674195.1 (tomato leaves), and
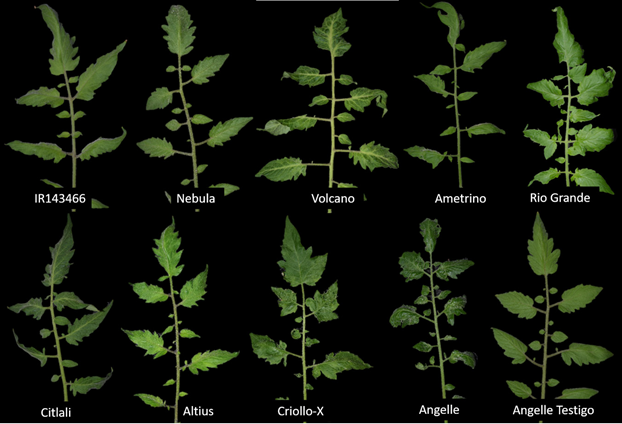

Figure 1 Symptoms in commercial and native tomato material inoculated with Tomato brown rugose fruit virus. Symptoms of mild mosaic (IR143466, Nebula, Ametrino, Rio Grande, Citlali, Altius); severe mosaic, deformation and clearing of main leaf nervations (Volcano, Criollo-X and Angelle); healthy plant (Angelle control).
OM515234.1 (pepper seeds), which correspond to Mexico, Canada and Israel. However, in another investigation, no mutations of ToBRFV were found after an inoculation process in tomato and pepper (Eichmeier et al., 2023). These results coincide with a study that reports between 2 and 39 nucleotide substitutions at the level of the complete genome (Abrahamian et al., 2022). Although the nucleotide changes are low, these changes could eventually lead to variability in the pathogenic and/or epidemiological behavior of the virus. This is supported with observations on tomato plants inoculated with the isolation from the State of Mexico (EMJI2021), which displayed a greater recorded height and stem diameter than the plants inoculated with the isolation from Colima (C-JI2021) (Data not shown). Symptoms were also observed in tomato plants inoculated with isolation C-JI2021, which were associated to European symptoms and more severe than those caused by the isolate from the State of Mexico.
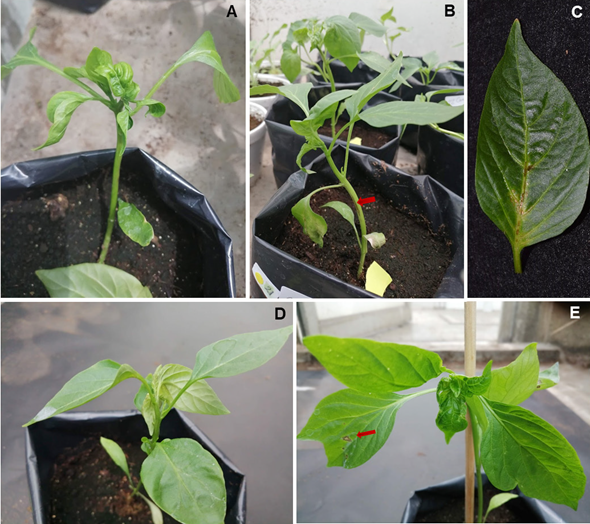

Figure 2 Symptoms in chili (Capsicum annumm) inoculated with Tomato brown rugose fruit virus. A-C) Symptoms of apical deformation, necrosis in stem and nervations of Almuden leaves; D) Symptoms of apical deformation, slight necrosis in Cayenne nervations; E) Symptoms of apical deformation, mosaic, necrotic lesions in noninoculated Felicitas leaves.
The importance of ToBRFV, from an early diagnosis of the virus to the impact on the production of tomato and pepper, is relevant. Studies have been performed on RNA extraction methods for an early and accurate diagnose in the molecular analysis (Zamora-Macorra et al., 2023), up to alternatives for the management of ToBRFV with a low environmental impact, such as the use of Beauveria peruviencis, Trichoderma longibrachiatum and Pseudomonas sp. (RamosVillanueva et al., 2023). Nevertheless, studies on the expression of symptoms in
Table 3 Percentage of coverage and identity of two ToBRFV isolations inoculated in a total of 35 tomato and pepper genotypes under greenhouse conditions.
| Isolate | Variety creolez | Crop | Isolate origin | lenght pb | Coverage | Identity | Homolaged isolate | |
|---|---|---|---|---|---|---|---|---|
| Accesion | Country | |||||||
| EM-JI2021 | Almuden | Chile | Edo. México | 398 | 100 | 100 | OQ674195.1 | Canada |
| Mulato | Chile | Edo. México | 390 | 100 | 99.74 | OQ674195.1 | Canada | |
| Congo | Chile | Edo. México | 391 | 100 | 99.74 | MT002973.1 | USA | |
| Gina | Chile | Edo. México | 381 | 98 | 100 | MT002973.1 | USA | |
| Santawest | Jitomate | Edo. México | 402 | 100 | 99.75 | OQ674195.1 | Canada | |
| Gotzilla | Jitomate | Edo. México | 383 | 100 | 100 | MT002973.1 | USA | |
| Altius | Jitomate | Edo. México | 385 | 100 | 99.74 | OQ674195.1 | Canada | |
| Don R | Jitomate | Edo. México | 407 | 100 | 100 | OQ674195.1 | Canada | |
| Sahariana | Jitomate | Edo. México | 392 | 100 | 99.74 | OQ674195.1 | Canada | |
| Nebula | Jitomate | Edo. México | 408 | 99 | 99.75 | OQ674195.1 | Canada | |
| C-JI2021 | Manzano | Chile | Colima | 402 | 100 | 100 | MT002973.1 | USA |
| Santawest | Jitomate | Colima | 410 | 100 | 100 | MT002973.1 | USA | |
| Altius | Jitomate | Colima | 407 | 100 | 100 | MT002973.1 | USA | |
| Sicilia | Jitomate | Colima | 411 | 100 | 100 | MT002973.1 | USA | |
| IR143466 | Jitomate | Colima | 418 | 100 | 100 | MT002973.1 | USA | |
| Don R | Jitomate | Colima | 408 | 100 | 100 | MT002973.1 | USA | |
| Ametrino | Jitomate | Colima | 384 | 100 | 100 | MT002973.1 | USA | |
| Olmeca | Jitomate | Colima | 399 | 100 | 100 | MT002973.1 | USA | |
| Nebula | Jitomate | Colima | 404 | 100 | 100 | MT002973.1 | USA | |
| Xochimilco | Jitomate | Colima | 384 | 100 | 100 | MT002973.1 | USA | |
| Angelle | Jitomate | Colima | 399 | 100 | 100 | MT002973.1 | USA | |
| Río Grande | Jitomate | Colima | 408 | 100 | 100 | MT002973.1 | USA | |
| Totolapan | Jitomate | Colima | 388 | 100 | 100 | MT002973.1 | USA | |
| Citlali | Jitomate | Colima | 385 | 100 | 100 | MT002973.1 | USA | |
Z Names in italics are native materials. In bold, sequences with a homology of <100%.
the different materials available in the market, as well as knowing the variability of ToBRFV that exists on the field, provides a guideline to better understand the virus epidemiology and its behavior according to the crop and the established weather conditions. It is therefore important to study the virus diversity in both the field and greenhouse over time and to observe the association between the diversity of expression of symptoms in tomato and pepper.
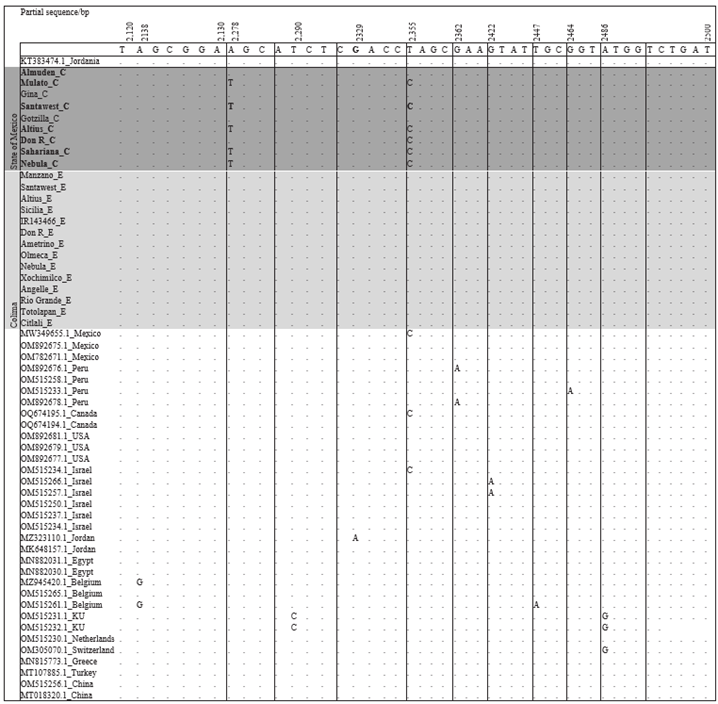
Figure 3 Alignment of the partial sequence of the tomato brown rugose fruit virus replicase gene from 24 sequences of isolates obtained from 35 genotypes inoculates with isolates EM-JI2021 and C-JI2021, and from 34 sequences from different tomato and chili-producing countries. The alignment was performed using Geneious.
Conclusions
All tomato materials expressed symptoms of mosaics, but without showing symptoms as severe as those reported for the first time, except in tomato plants inoculated with isolation C-JI2021. In pepper, lesions were recorded on inoculated leaves (hypersensitivity reaction), as well as necrotic lesions on the stem, apical deformation, necrosis in leaf nervations and asymptomatic.
According to the alignment analysis of two ToBRFV isolates (State of Mexico and Colima), in comparison with 34 isolates available in the NCBI Genbank, in five sequences of isolates from the State of Mexico (EM-JI2021) the substitution of two nucleotides were observed, nucleotides c.2,278A>T and c.2355T>C. Accessions MW349655.1 (Mexico), OQ674195.1 (Canada) and OM515234.1 (Israel) coincided with isolate EM-JI2021 (inoculated with the variety Don R) in one nucleotide change c.2355T>C. In this study, a total of three haplotypes were found. This is the first study that reports variability of ToBRFV in a short period under controlled conditions. This information may be relevant for cross-protection studies.
Acknowledgements
The first author thanks the Consejo Nacional de Humanidades, Ciencias y Tecnologías (CONAHCyT) for the postdoctoral fellowship provided to perform this investigation work. To the LANREF-CP team for the methodological and logistical support.
REFERENCES
Abrahamian, P., Cai, W., Nunziata, S.O., Liing, K.S., Jaiswal, N., Mavrodieva, V.A., Rivera, Y., y Nakhla, M.K. (2022). Comparative analysis of Tomato brown rugose fruit virus isolates shows limited genetic diversity. Viruses, 14:2816. 10.3390/v14122816 [ Links ]
Camacho-Beltrán, E., Pérez-Villarreal, A., Leyva-López, N.E., Rodríguez-Negrete, E.A., Ceniceros-Ojeda, E.A., y Méndez-Lozano, J. (2019). Occurrence of Tomato brown rugose fruit virus Infecting Tomato Crops in Mexico. Disease Notes, 103(6), 1440. 10.1094/PDIS-11-18-1974-PDN [ Links ]
Cambrón-Crisantos, J.M., Rodríguez-Mendoza, J., Valencia-Luna, J.B., Alcasio-Rangel, S., García-Ávila, C.J., López-Buenfil, J.A., y Ochoa-Martínez, D.L. (2018). First report of Tomato brown rugose fruit virus (ToBRFV) in Michoacan, Mexico. Mexican Journal of Phytopathology, 37(1), 185-192. 10.18781/R.MEX.FIT.1810-5 [ Links ]
Eichmeier, A., Hejlova, M., Orsagova, H., Frejlichova, L., Hakalova, E., Tomankova, K., Linhartova, S., Kulich, P., Cermak, V., y Cechova, J. (2023). Characterization of Tomato brown rugose fruit virus (ToBRFV) detected in Czech Republic. Diversity, 15(2), 301. 10.3390/d15020301 [ Links ]
Fidan, H., Sarikaya, P., Yildiz, K., Topkaya, B., Erkis, G., y Calis, O. (2021). Robust molecular detection of the new Tomato brown rugose fruit virus in infected tomato and pepper plants from Turkey. Journal of Integrative Agriculture, 20(8), 2170-2179. 10.1016/S2095-3119(20)63335-4 [ Links ]
Hanssen, I.M., Lapidot, M., y Thomma, B.P.H.J. (2010). Emerging viral diseases of tomato crops. Molecular Plant-Microbe Interactions, 23(5), 539-548. 10.1094/MPMI-23-5-0539 [ Links ]
Maayan, Y., Pandaranayaka, E.P.J., Srivastava, D.A., Lapidot, M., Levin, I., Dombrovsky, A., y Harel, A. (2018). Using genomic analysis to identify tomato Tm-2 resistance-breaking mutations and their underlying evolutionary path in a new and emerging tobamovirus. Archives of Virology, 163, 1863-1875. 10.1007/s00705-018-3819-5 [ Links ]
Ramos-Villanueva, C.D., Carrillo-Benitez, G., Zamora-Macorra, E.J., Santiago-Elena, E., Ramírez-Alarcón, S., Jimenez-Vidals, J., y Ávila López, R. (2023). Use of endophytic microorganisms for the management of Tomato brown rugose fruit virus in tomato crop (Solanum lycopersicum). Mexican Journal of Phytopathology, 41(4), 1. 10.18781/R.MEX.FIT.2023-1 [ Links ]
Salem, N., Mansour, A., Ciuffo, M., Falk, B.W., y Turina, M. (2016). A new tobamovirus infecting tomato crops in Jordan. Archives of Virology, 161, 503-506. 10.1007/s00705-015-2677-7 [ Links ]
SENASICA (Servicio Nacional de Sanidad, Inocuidad y Calidad Agroalimentaria) (2018). Protocolo de Diagnóstico: Tomato brown rugose fruit virus (ToBRFV) (Virus del fruto marrón rugoso del tomate) [Versión 2.0]. Tecámac, México. 18. [ Links ]
Yu, D., Tang, H., Zhang, Y., Du, Z., Yu, H., y Chen, Q. (2012). Comparison and Improvement of Different Methods of RNA Isolation from Strawberry (Fragaria × ananassa). Journal of Agricultural Science, 4(7), 51-56. 10.5539/jas.v4n7p51 [ Links ]
Zamora-Macorra, E.J., Aviña-Padilla, K., Rosemarie, H.W., y Ochoa-Martínez, D.L. (2023). Comparison of molecular protocols to detect Tomato brown rugose fruit virus in solanaceae hosts. Mexican Journal of Phytopathology, 41(4), 2. 10.18781/R.MEX.FIT.2023-5 [ Links ]
Zhi-Yong, Y., Mei-sheng, Z., Hua-yu, M., Ling-zhi, L., Guang-ling, Y., Chao, G., Yanping, T., y Xiang-dong, L. (2021). Biological and molecular characterization of tomato brown rugose fruit virus and development of quadruplex RT-PCR detection. Journal of Integrative Agriculture, 20(7), 1871-1879. 10.1016/S2095-3119(20)63275-0 [ Links ]
Received: November 30, 2023; Accepted: March 06, 2024











 texto en
texto en 


