Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Accesos
Accesos
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO
Compartir
Revista mexicana de fitopatología
versión On-line ISSN 2007-8080versión impresa ISSN 0185-3309
Rev. mex. fitopatol vol.41 no.spe Texcoco 2023 Epub 31-Mar-2025
https://doi.org/10.18781/r.mex.fit.2023-5
Phytopathological Notes
Comparison of molecular protocols to detect Tomato brown rugose fruit virus in solanaceae hosts
1 Departamento de Preparatoria Agrícola, Universidad Autónoma Chapingo, Km 38.5 Carretera México-Texcoco, Estado de México, CP. 56230, México;
2 Departamento de Ingeniería Genética del Centro de Investigación y de Estudios Avanzados del I.P.N. Unidad Irapuato, Irapuato 36821, México;
3 USDA, Agricultural Research Service, Beltsville Agricultural Research Center, Beltsville, MD 20705, USA;
4 Colegio de Postgraduados, Posgrado en Fitosanidad-Fitopatología, Texcoco 56230, México.
Objective/Background: The Tomato brown rugose fruit virus (ToBRFV) has emerged as a significant threat to Solanaceae crops, including tomato and pepper. Its presence in Mexico since 2018 has raised concerns about its impact on agricultural production. Early and accurate detection of this pathogen is essential to prevent its spread and mitigate its effects. In Mexico, several molecular techniques are employed for its diagnosis, including endpoint RT-PCR, RT-qPCR, and multiplex RT-qPCR.
Materials and Methods: This research aimed to assess the efficiency of different RNA extraction methods in combination with specific PCR primers for detecting ToBRFV.
Results: Among the methods tested, the CTAB-Trizol RNA extraction protocol combined with nested PCR using primers reported by Dovas et al. (2004) was identified as the most sensitive molecular method for detecting the virus.
Conclusion: This finding highlights the importance of selecting the appropriate combination of extraction and amplification protocols to achieve optimal sensitivity and accuracy in ToBRFV detection.
Keywords: Tomato brown rugose fruit virus; ToBRFV; Solanaceae crops; RNA extraction; RT-PCR; molecular detection; agricultural production
Objetivo/Antecedentes: El virus del fruto rugoso marrón del jitomate (ToBRFV) ha surgido como una amenaza significativa para los cultivos de la familia Solanaceae, incluidos el tomate y el pimiento. Su presencia en México desde 2018 ha generado preocupación sobre su impacto en la producción agrícola. La detección temprana y precisa de este patógeno es crucial para prevenir su propagación y mitigar sus efectos. En México, se emplean varias técnicas moleculares para su diagnóstico, incluyendo RT-PCR convencional, RT-qPCR y RT-qPCR multiplex.
Materiales y Métodos: El objetivo de esta investigación fue evaluar la eficiencia de diferentes métodos de extracción de ARN en combinación con oligonucléotidos PCR específicos para la detección de ToBRFV.
Resultados: Entre los métodos probados, el protocolo de extracción de ARN CTAB-Trizol combinado con PCR anidada utilizando oligonucleótidos reportados por Dovas et al. (2004) se identificó como el método molecular más sensible para detectar el virus.
Conclusión: Este hallazgo destaca la importancia de seleccionar la combinación adecuada de protocolos de extracción y amplificación para lograr la sensibilidad y precisión óptimas en la detección de ToBRFV.
Palabras clave: Virus del fruto rugoso marrón del jitomate; ToBRFV; cultivos de Solanaceae; extracción de ARN; RT-PCR; detección molecular; producción agrícola
Tomato brown rugose fruit virus (ToBRFV) poses a grave threat to tomato production. Initially identified in Israel in 2014 (Solanum lycopersicum) (Luria et al., 2017), it extended to Jordan in 2015 (Salem et al., 2016). Since then, its footprint has expanded dramatically. ToBRFV has been identified in tomato crops across diverse regions, including Mexico (Cambrón-Crisantos et al., 2018), Germany (Menzel et al., 2019), Turkey (Fidan et al., 2019), the United Kingdom (Skelton et al., 2019), China (Yan et al., 2019), Netherlands (Van de Vossenberg et al., 2020), the United States (Ling et al., 2019), Italy (Panno et al., 2019), Palestine (Alkowni, 2019), Greece (EPPO, 2019/210), Spain (EPPO, 2019/238), and France (MAA, 2020). Notably, in 2020, it infiltrated chili greenhouses in Italy and Jordan (Salem et al., 2020; Panno et al., 2020a), while in 2018, commercial pepper greenhouses in Mexico also were affected (Cambrón-Crisantos et al., 2018). This pervasive spread of ToBRFV across continents underscores its substantial threat to tomato cultivation. Urgent measures are imperative to combat its rampant expansion and protect agricultural ecosystems.
ToBRFV, classified as a Tobamovirus, exhibits a distinctive rigid rod-shaped morphology in its particles. Its genome comprises a straightforward single-stranded RNA chain (+ssRNA) containing four open reading frames (ORFs). Within these ORFs, ORF1 and ORF2 govern the RNA-dependent RNA polymerase (RdRp) function. ORF3 orchestrates the movement protein (MP), while the concise ORF4 oversees the capsid protein (CP) synthesis (Luria et al., 2017). This virus is notable for its particle stability, facilitating its remarkably easy mechanical transmission. Consequently, greenhouse cultivation practices serve as the primary conduit for its dissemination (Levitzky et al., 2019; Panno et al., 2020b). The robustness of ToBRFV’s particles amplifies its potential to spread rapidly, emphasizing the pressing need for stringent measures to curtail its propagation and safeguard crop ecosystems.
In this context, the detection of ToBRFV in plants has been approached through a range of techniques, encompassing the Enzyme-Linked Immunosorbent Assay (ELISA), electronic transmission microscopy, RT-PCR, second-generation sequencing, qRT-PCR, and Loop-mediated Isothermal Amplification (LAMP) (Oladokun et al., 2019; Ling et al., 2019; Panno et al., 2019; Sarkes et al., 2020). These methods differ in terms of their sensitivity limits, cost implications, and the level of expertise required for their execution. Selecting the optimal technique hinges on the specific diagnostic objectives, the type of plant material under scrutiny, budget constraints, and the time available for analysis. The array of detection techniques underscores the need for a tailored approach, aligning the chosen method with the particular demands of the diagnosis. This pragmatic approach ensures both accuracy and resource optimization in combating ToBRFV’s proliferation.
In Mexico, the National Center of Phytosanitary Reference (CNRF) initially advocated the use of molecular techniques for ToBRFV identification, specifically endpoint RT-PCR employing the ToBRFV F/ToBRFV-R primer pair (RodríguezMendoza et al., 2019). However, a recent update in the molecular diagnostic protocols has introduced comprehensive enhancements. The revised procedures now encompass a combination of endpoint RT-PCR, RT-qPCR, and multiplex RT-qPCR. According to the latest guidelines issued by CNRF, the diagnosis of ToBRFV requires an integrated approach, whereby a positive result should be corroborated by at least two of the aforementioned techniques (SENASICA, 2022). This shift towards a multifaceted diagnostic strategy underscores the significance of robust and cross-validated results, aligning with international best practices. The heightened emphasis on diagnostic accuracy is pivotal in containing the spread of ToBRFV within Mexico’s agricultural landscape.
Indeed, the aim of this study was to undertake a comprehensive comparison of diverse RNA extraction protocols, evaluating their efficacy in detecting ToBRFV. In addition, the research aimed to assess the limit of detection for three distinct primer pairs, employing both RT-PCR and nested-RT-PCR methodologies. By conducting this comparative analysis, our study sought to determine the most efficient and sensitive combination of RNA extraction methods and primer pairs for accurate ToBRFV detection. This research contributes to enhancing the precision and reliability of diagnostic techniques crucial for managing the spread of ToBRFV in agricultural settings.
Plant material for inoculum source. During September 2020, tomato plants (Solanum lycopersicum), showing chlorosis, leaf narrowing, and mosaics (typical symptoms of ToBRFV infection) were collected from commercial greenhouses in Colima, Mexico (Figure 1). The inoculum source tested positive for ToBRFV by RT-PCR. A Nicotiana glutinosa plant was mechanically inoculated using this tissue, and local necrotic lesions were observed. One of these lesions was cut to inoculate a second N. glutinosa plant. This process was repeated twice. A local lesion of the fourth N. glutinosa plant was used to inoculate a tomato plant. Then, 30 days post-inoculation (dpi), it was analyzed by RT-PCR using the specific primers (ToBRFV-FMX: AACCAGAGTCTTCCTATACTCGGAA/ ToBRFVRMX: CTCWCCATCTCTTAATAATCTCCT: 475 bp Region ORF 1-RdRp) and conditions described by Rodríguez-Mendoza et al. (2018), Table 1. The expected fragments were then sequenced and compared in the GenBank. The inoculum source was increased by mechanical inoculation on tomato plants to perform the different host bioassays.
Evaluation of total RNA extraction methods. Twenty seeds were placed in each Petri dish (three dishes per plant species), using bell pepper seeds and saladette tomato seeds that were germinated in Petri dishes with paper towels moistened with
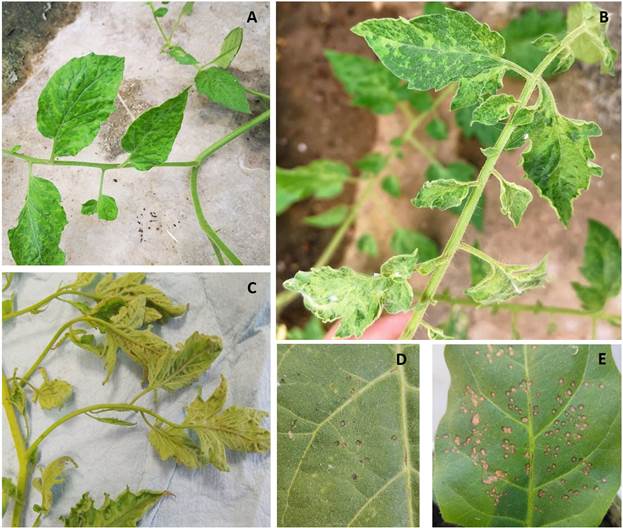

Figure 1 A-C) Symptoms of mosaic, leaf deformation, and leaf narrowing were evident in tomato saladette plants gathered from greenhouse settings. These plants tested positive for Tomato brown rugose fruit virus (ToBRFV); D-E) visual observations on inoculated Nicotiana leaves revealed the presence of localized chlorotic and necrotic lesions caused by ToBRFV infection. The virus-positive plants utilized for the study were sourced from Tecoman, Colima, Mexico.
sterile distilled water. After 10 to 20 days, when the cotyledonary leaves emerged and expanded, 100 mg of complete seedlings (stem, leaves, and root) of each plant species were separately used to perform total RNA extraction. Likewise, three seeds were planted in 1 L Styrofoam cups with peat of most of each plant species. One plant was used as a replicate. The plants were kept in a greenhouse with 12 hours of light and 12 hours of darkness, with minimum temperatures of 15 °C and maximum temperatures of 32 °C. After 45 days, 100 mg of foliar tissue (pedicels, leaves, and shoots) were collected to perform total RNA extraction. A total of four
Table 1 Primers tested in this study for ToBRFV detection in tomato, tomatillo, and eggplant.
| Nucleotide sequences of the primers (5’ to 3’) | Expected fragment size and region for the PCR amplification | RT-PCR conditions | Reference |
|---|---|---|---|
| ToBRFV-FMX: AACCAGAGTCTTCCTATACTCGGAA | 475 pb (ORF 1-RdRp)θ | - 95° 3 minβ. | Rodríguez- Mendoza et al., 2018 |
| -32 cycles: 95° 50 sΩ, | |||
| ToBRFV-RMX: CTCWCCATCTCTTAATAATCTCCT | 53° 50 s y 72° 55 s. | ||
| -72°C 10 min. | |||
| TobN up3: GGCGYTGCARACIATHGTITAYCA | 400 pbα (ORF 1-RdRp) θ | -95° 3 minα. | Dovas et al., 2004 |
| -4 cycles: 95° 30 s, | |||
| TobN do4: GTRTTICCIATRAAIGTIGTIACRTC | 51° 30 s y 72° 30 s. | ||
| -26 cycles: 95° 30 s, | |||
| TobN do4G: GCCGATRAAGGTGGTGACRTC | 51° 30 s y 72° 30 s. | ||
| -72°C 10min. | |||
| ToBRFV-F: GAAGTCCCGATGTCTGTAAGG | 842 pb (ORF 3- MP) θ | -95° 3 min. | Ling et al., 2019 |
| -32 cycles: 95° 50 s, | |||
| ToBRFV-R: TGCCTACGGATGTGTATGA | 53° 50 s y 72° 55 s. | ||
| -72°C 10min. |
θ Section of the ToBRFV gene amplified, αNested RT-PCR, βminutes, Ωseconds.
RNA extraction methods were evaluated: PlantRNAeasy miniKit®, following the manufacturer’s instructions; Trizol®, CTAB 2 % and CTAB 2%-Trizol® following the protocol described by Jordon-Thaden et al. (2015) (minor modifications, appendix 1). Plant tissue (pedicels, leaves, and shoots) maceration was performed with liquid nitrogen in a sterile mortar. The amount and quality of the extracted RNA were quantified using a Nanodrop 2000®, and the absorbance measurements 260/280 and 260/230 were recorded.
Primer evaluation and sensitivity limit. Three replicates of tomato (S. lycopersicum), eggplant (Solanum melongena), and tomatillo (Physalis ixocarpa) seedlings were mechanically inoculated. Inoculations were carried out when the first true leaf (leaflet) emerged and developed. Plants that were not inoculated were also kept, as controls. All plants were maintained in the greenhouse under the previously described conditions. Then, at 30 dpi, when plants showed chlorosis and mosaic symptoms, the total RNA extraction using the previously described CTABTrizol protocol was performed. The quality and quantity of the extracted RNA were verified using a Nanodrop 2000®. The final concentration was adjusted to 300 ng µL-1. Subsequently, serial dilutions of each tRNA extraction were made and adjusted to 10, 1, 0.1, 0.01, 0.001, and 0.0001 ng µL-1. Pairs of specific primers, described by Ling et al. (2019) and Rodríguez-Mendoza et al. (2018), and reported by Dovas et al. (2004), were tested, Table 1. We employed this protocol as it yielded a higher concentration of nanograms compared to using Trizol and the kit, while also being more cost-effective. The specifications described in the protocols reported for each primer were followed.
Evaluation of total RNA extraction methods. In our comparison of methodologies, it became evident that tomato and pepper seedlings yielded a notably higher concentration of total RNA (Figure 2). Significantly, the CTAB-Trizol protocol exhibited the most pronounced outcome in terms of nanograms per microliter, obtaining the highest concentration (966 ng μL-1). In contrast, the RNA isolation Kit manifested the lowest concentration (29 to 101 ng μL-1) among the evaluated methods (Figure 2).
Primer evaluation and sensitivity limit. Our findings delineate the varying sensitivity limits of different primer sets in detecting ToBRFV using total RNA extracted from tomato, tomatillo, and eggplant plants (Table 2). Specifically, the primers devised by Ling et al. (2019) exhibited the capacity to detect the virus down to a concentration of 0.01 ng µL-1 in tomato and tomatillo samples. In contrast, their sensitivity in eggplant RNA samples manifested at a higher concentration of 1 ng µL-1. In parallel, the primers developed by Rodríguez-Mendoza et al. (2018) showcased a detection limit of 0.1 ng µL-1 in tomato and 0.01 ng µL-1 in tomatillo. Irrespective of the primer selection, the sensitivity threshold for virus detection in eggplant samples consistently resided at 1 ng µL-1.
Notably, an intriguing escalation in sensitivity thresholds was observed for both tomato and tomatillo samples through the employment of the nested RTPCR technique coupled with the primer set elucidated by Dovas et al. (2004). This optimization enabled the identification of an elevated level of sensitivity in detecting ToBRFV 0.0001 ng µL-1 in tomato, 0.001 ng µL-1 in tomatillo, and 1 ng µL-1 in eggplant samples (Table 2, Figure 3). These nuanced distinctions in sensitivity thresholds emphasize the significance of primer selection and nested RT-PCR amplification in optimizing the detection capability of ToBRFV in diverse plant species.
Several RNA extraction methods have been developed with a central aim: to yield optimal concentrations of high-quality RNA. This is particularly crucial considering that plant tissues often harbor significant amounts of potentially interfering substances such as phenolic compounds, polysaccharides, pigments, and RNases (Wang et al., 2009). Phenolic content within plants is remarkably diverse and subject to fluctuations determined by various factors, including the developmental stage of the plant and its exposure to biotic and abiotic environmental influences. In the context of tomato leaves, the phenol content may exhibit a decline with
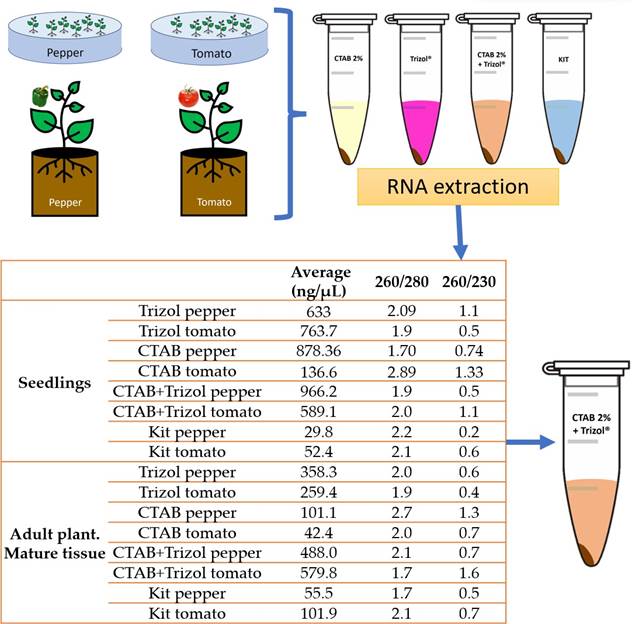
Figure 2 The evaluation of total RNA extraction methods involved comparing their effectiveness and yield, as measured by absorbance readings on a Nanodrop 2000® spectrophotometer. Among the methods tested, including Trizol®, CTAB 2%, and CTAB 2%-Trizol®, the CTAB 2%-Trizol® protocol yielded the highest concentration of total RNA, while the RNA isolation Kit produced the lowest concentration. This data was recorded through absorbance measurements at wavelengths of 260/280 and 260/230. The Nanodrop 2000® was used to quantify the extracted RNA’s concentration and assess its quality by these absorbance ratios. The results indicated that the CTAB 2%-Trizol® protocol was particularly effective in extracting high-quality RNA from the plant samples, making it a suitable choice for subsequent molecular analyses.
Table 2 Comparison of Primer References, Plant Sources, and RNA Detection Limits in source of infected plant material.
| Reference for Primers | Source of Infected Plant Material | Limit of Detected Total RNA Concentration (ng μL-1) |
|---|---|---|
| Rodriguez-Mendoza et al., 2018. | Tomato | 0.1000 |
| Tomato | 0.0100 | |
| Tomato | 1.0000 | |
| Ling et al., 2019. | Tomatillo | 0.0100 |
| Tomatillo | 0.0100 | |
| Tomatillo | 1.0000 | |
| Dovas et al., 2004. | Eggplatn | 0.0001 |
| Eggplatn | 0.0010 | |
| Eggplatn | 1.0000 |
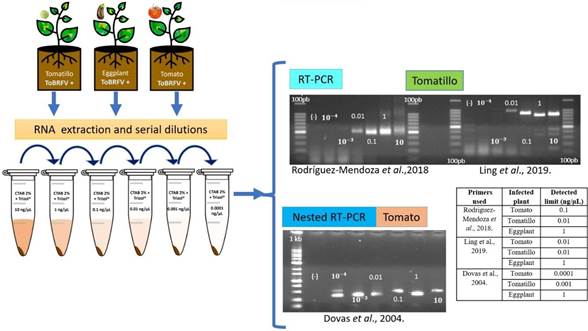
Figure 3 Evaluation and sensitivity of PCR primers. 1.5% agarose gels electrophoretic analysis of RT-PCR and Nested RT-PCR products (Ling´s primers expected size 842 pb; Rodríguez-Mendoza´s primers expected size 475 bp and Dovas´s primers expected size 400 pb). (-): sterilized water instead RNA. 100 bp = 100bp DNA Ladder (Invitrogen®). 1Kb= 1000 bp DNA ladder (Promega®). 10-3 and 10-4 = 0.001 and 0.0001 ng µL-1.
the progression of age. However, it’s noteworthy that the composition of various phenolic compounds can exhibit dynamic variations throughout different stages of leaf development (Dadáková et al., 2020). This intricate interplay between RNA extraction techniques and the complex composition of plant tissues underscores the importance of selecting appropriate methods to ensure reliable and representative RNA samples for downstream analyses.
Polyphenolic compounds notably engage in intricate interactions, forming intricate molecular aggregates with proteins and nucleic acids. This propensity to create large molecular weight complexes further complicates the extraction process. Concurrently, the presence of polysaccharides compounds the challenge, as they have a tendency to co-precipitate with RNA in the presence of alcohol-based extraction protocols. This phenomenon leads to the persistence of polysaccharide contaminants within the final extracted RNA product, thereby affecting its integrity (Salzman et al., 1999).
The effectiveness of the CTAB and Trizol methods transcends their costefficiency. These methods have gained widespread adoption across diverse plant species and tissues, consistently yielding favorable outcomes (Gambino et al., 2008). The commendable quality and purity of RNA, coupled with the expedited and budget-friendly nature of these extraction techniques, render them particularly suitable for virus detection (Mathioudakis et al., 2020). The CTAB method particularly facilitates rapid RNA extraction from challenging tissues, eliminating the need for hazardous chemicals (Chang et al., 1993). It is noteworthy that while the Trizol method yields quality RNA extractions, it also concurrently extracts DNA. Employing Trizol in conjunction with CTAB extraction further enhances the quality of total RNA extraction.
The quality and concentration of extracted RNA wields a profound impact on the precision and reliability of various plant molecular studies, including RT-PCR (Gambino et al., 2008) and gene expression analysis (Toni et al., 2018). However, in the context of large-scale detection assays, the selection of an RNA extraction method is further influenced by cost considerations. In this regard, methodologies like CTAB and Trizol emerge as cost-effective alternatives compared to commercially available kits. The versatility and efficacy of the CTAB and Trizol methods extend across diverse plant species and tissues, consistently yielding favorable outcomes (Gambino et al., 2008). These approaches offer dual benefits of attaining high-quality and pure RNA swiftly and economically, rendering them particularly advantageous for expedited virus detection (Mathioudakis et al., 2020).
Our findings unequivocally highlight the variability in sensitivity limits exhibited by each pair of primers across different plant species. This phenomenon likely arises due to variations in the expression levels and concentration of the pathogen within different host plants. Interestingly, eggplant displayed consistent sensitivity limits irrespective of the primer set employed. In the case of tomatillo, both Ling’s and Rodríguez-Mendoza’s primers demonstrated equivalent sensitivity limits, while for tomato, Ling’s primers exhibited tenfold greater sensitivity. Notably, Ling’s primers are meticulously designed to target the coat protein region, whereas Rodríguez-Mendoza’s primers detect the RdRp region. A noteworthy observation by Magaña-Álvarez et al. (2021) underscores the superior performance of coat protein-targeting primers in ToBRFV detection within tomato plants compared to RdRp-targeting primers.
It has been established that nested PCR, while more time and resource-intensive, offers heightened sensitivity and reliability in comparison to conventional PCR (Shin et al., 2004). As anticipated, the Dovas primers displayed an impressive sensitivity that was a hundredfold higher in tomato and tenfold higher in tomatillo. Altogether our results show that the optimal detection protocol for ToBRFV, within the scope of these experimental conditions, emerges as the CTAB-Trizol RNA extraction method paired with nested RT-PCR employing the primers delineated by Dovas et al. (2004).
In conclusion, we performed a comprehensive comparison of various molecular protocols for the detection of Tomato brown rugose fruit virus (ToBRFV) in Solanaceae hosts. ToBRFV poses a significant threat to tomato production globally, and its early and accurate detection is crucial to mitigate its impact. The study evaluated RNA extraction methods and specific PCR primer pairs for their efficiency in detecting the virus. The CTAB-Trizol RNA extraction protocol combined with nested PCR using primers reported by Dovas et al. (2004) was identified as the most sensitive method for ToBRFV detection. Our study underscores the importance of selecting appropriate combinations of extraction and amplification protocols to achieve optimal sensitivity and accuracy in ToBRFV detection. These findings could contribute to enhancing the precision of diagnostic techniques crucial for managing the spread of ToBRFV in agricultural settings.
Acknowledgments
K.A.-P. (CVU:227919), received financial support from CONACyT
REFERENCES
Alkowni, R., Alabdallah, O. y Fadda, Z. (2019). Molecular identification of Tomato brown rugose fruit virus in tomato in Palestine. Journal of Plant Pathology, 101, 719-723. https://doi.org/10.1007/s42161-019-00240-7 [ Links ]
Cambrón-Crisantos, J.M., Rodríguez-Mendoza, J., Valencia-Luna, J.B., Rangel, S.A., García-Ávila, D.J.C. y López-Buenfil, J.A. (2018). First report of Tomato brown rugose fruit virus (ToBRFV) in Michoacan, Mexico. Revista Mexicana de Fitopatología, 37, 185-192. https://doi.org/10.18781/R.MEX.FIT.1810-5 [ Links ]
Chang, S., Puryear, J. y Cairney, J. (1993). A simple and efficient method for isolating RNA from pine trees. Plant Molecular Biology Reporter, 11, 113-116. https://doi.org/10.1007/BF02670468 [ Links ]
Dadáková, K., Heinrichová, T., Lochman, J. y Kašparovský, T. (2020). Production of Defense Phenolics in Tomato Leaves of Different Age. Molecules, 25(21), 4952. https://doi.org/10.3390/molecules25214952 [ Links ]
Dovas, C.I., Efthimiou, K. y Katis, N.I. (2004). Generic detection and differentiation of tobamoviruses by a spot nested RT-PCR-RFLP using dI-containing primers along with homologous dG-containing primers. Journal of Virological Methods, 117, 137-144. https://doi.org/10.1016/j.jviromet.2004.01.004 [ Links ]
European and Mediterranean Plant Protection Organization (EPPO) (2019). Reporting Service 2019/210. https://gd.eppo.int/reporting/article-6640 [ Links ]
European and Mediterranean Plant Protection Organization (EPPO) (2019). Reporting Service 2019/238. https://gd.eppo.int/reporting/article-6668 [ Links ]
Fidan, H., Sarikaya, P. y Calis, O. (2019). First report of Tomato brown rugose fruit virus on tomato in Turkey. New Disease Reports, 39, 18. https://doi.org/10.5197/j.2044-0588.2019.039.018 [ Links ]
Gambino, G., Perrone, I. y Gribaudo, I. (2008). A Rapid and effective method for RNA extraction from different tissues of grapevine and other woody plants. Phytochemical Analysis, 19(6), 520-525. https://doi.org/10.1002/pca.1078 [ Links ]
Jordon-Thaden, I.E., Chanderbali, A.S., Gitzendanner, M.A. y Soltis, D.E. (2015). Modified CTAB and Trizol protocols improve RNA extraction from chemically complex embryophyta. Applications in Plant Sciences, 3(5), 1400105. https://doi.org/10.3732/apps.1400105 [ Links ]
Ling, K.S., Tian, T., Gurung, S., Salati, R. y Gilliard, A. (2019). First report of Tomato brown rugose fruit virus infecting greenhouse tomato in the United States. Plant Disease, 103, 1439. https://doi.org/10.1094/PDIS-11-18-1959-PDN [ Links ]
Levitzky, N., Smith, E., Lachman, O., Luria, N., Mizrahi, Y., Bakelman, H., Sela, N., Laskar, O., Milrot, E. y Dombrovsky, A. (2019). The bumblebee Bombus terrestris carries a primary inoculum of Tomato brown rugose fruit virus contributing to disease spread in tomatoes. PloS One, 14(1), e0210871. https://doi.org/10.1371/journal.pone.0210871 [ Links ]
Luria, N., Smith, E., Reingold, V., Bekelman, I., Lapidot, M., Levin, I., Elad, N., Tam, Y., Sela, N., Abu-Ras, A., Ezra, N., Haberman, A., Yitzhak, L., Lachman, O. y Dombrovsky, A. (2017). A New Israeli Tobamovirus Isolate Infects Tomato Plants Harboring Tm-22 Resistance Genes. PloS One, 12(1), e0170429. https://doi.org/10.1371/journal.pone.0170429 [ Links ]
Mathioudakis, M.M., Saponari, M., Hasiów-Jaroszewska, B., Elbeaino, T. y Koubouris, G. (2020). Detection of viruses in olive cultivars in Greece, using a rapid and effective RNA extraction method, for certification of virus-tested propagation material. Phytopathologia Mediterranea, 59(1), 203-211. https://doi.org/10.14601/Phyto-11033 [ Links ]
Magaña-Álvarez, A.A., Pérez-Brito, D., Vargas-Hernández, B.Y., Ramírez-Pool, J.A., Núñez-Muñoz, L.A., Salgado-Ortiz, H., de la Torre-Almaraz, R., Ruiz-Medrano, R. y Xoconostle-Cázares, B. (2021). Detection of Tomato brown rugose fruit virus (ToBRFV) in solanaceous plants in Mexico. Journal of Plant Disease Protection, 128, 1627-1635. https://doi.org/10.1007/s41348-021-00496-1 [ Links ]
Menzel, W., Knierim, D., Winter, S., Hamacher, J. y Heupel, M. (2019). First report of Tomato brown rugose fruit virus infecting tomato in Germany. New Disease Reports, 39(1). https://doi.org/10.5197/j.2044-0588.2019.039.001 [ Links ]
Ministère de L’Agriculture et de L’Alimentation (MAA) (2020). Virus ToBRFV: le ministère confirme la contamination de tomates en serre dans le Finistère. https://agriculture.gouv.fr/virus-tobrfv-le-ministere-confirme-la-contamination-de-tomates-en-serre-dans-le-finistere [ Links ]
Oladokun, J., Halabi, M., Barua, P. y Nath, P.J. (2019). Tomato brown rugose fruit disease: current distribution, knowledge, and future prospects. Plant Pathology, 68, 1579-1586. https://doi.org/10.1111/ppa.13096 [ Links ]
Panno, S., Caruso, A.G. y Davino, S. (2019). First Report of Tomato brown rugose fruit virus on Tomato Crops in Italy. Plant Disease, 103, 1443. https://doi.org/10.1094/PDIS-12-18-2254-PDN [ Links ]
Panno, S., Caruso, G.A., Blanco, G. y Davino, S. (2020). First report of Tomato brown rugose fruit virus infecting sweet pepper in Italy. New Disease Report, 41(20). https://doi.org/10.5197/j.2044-0588.2020.041.020 [ Links ]
Panno, S., Caruso, G.A., Blanco, G., Lo Bosco, G., Ezequiel, R.A. y Davino, S. (2020). Spread of Tomato brown rugose fruit virus in Sicily and evaluation of the spatiotemporal dispersion in experimental conditions. Agronomy, 10, 834. https://doi.org/10.3390/agronomy10060834 [ Links ]
Rodríguez-Mendoza, J., García-Ávila, C.J., López-Buenfil, J.A., Araujo-Ruiz, K., Quezada-Salinas, A. y Cambrón-Crisantos, J.M. (2018). Identification of Tomato brown rugose fruit virus by RT-PCR from a coding region of replicase (RdRP). Mexican Journal of Phytopathology, 37(2), 1-12. https://doi.org/10.18781/R.MEX.FIT.1902-6 [ Links ]
Salzman, R.A., Fujita, T., Zhu-Salzman, K., Hasegawa, P.M. y Bressan, R.A. (1999). An improved RNA isolation method for plant tissues containing high levels of phenolic compounds or carbohydrates. Plant Molecular Biology Report, 17, 11-17. https://doi.org/10.1023/A:1007520314478 [ Links ]
Salem, N., Mansour, A., Ciuo, M., Falk, B.W. y Turina, M. (2016). A new tobamovirus infecting tomato crops in Jordan. Archives of Virology, 161, 503-506. https://doi.org/10.1007/s00705-015-2677-7 [ Links ]
Salem, N.M., Cao, M.J., Odeh, S., Turina, M. y Tahzima, R. (2020). First Report of Tobacco mild green mosaic virus and Tomato brown rugose fruit virus Infecting Capsicum annuum in Jordan. Plant Disease, 104, 601. https://doi.org/10.1094/PDIS-06-19-1189-PDN [ Links ]
Sarkes, A., Fu, H., Feindel, D., Harding, M.W. y Feng, J. (2020). Development and evaluation of a loop-mediated isothermal amplification (LAMP) assay for the detection of Tomato brown rugose fruit virus (ToBRFV). PLOS ONE, 15(6), e0230403. https://doi.org/10.1371/journal.pone.0230403 [ Links ]
SENASICA (2022). Procedimientos de diagnóstico: Tomato brown rugose fruit virus (ToBRFV) (Virus del fruto rugoso café del tomate). https://www.gob.mx/cms/uploads/attachment/file/759117/Procedimientos_de_Diagn_stico_ToBRFV_2022.pdf [ Links ]
Shin, Y., Cho, K., Cho, H., Kang, S., Kim, H., Kim, Y. y Park, N. (2004). Comparison of one-step RT-PCR and a nested PCR for the detection of canine distemper virus in clinical samples. Australian Veterinary Journal, 82(1-2), 83-86. https://doi.org/10.1111/j.1751-0813.2004.tb14651.x [ Links ]
Skelton, A., Buxton-Kirk, A., Ward, R., Harju, V., Frew, L., Fowkes, A., Long, M., Negus, A., Forde, S., Adams, I.P., Pufal, H., McGreig, S., Weekes, R. y Fox, A. (2019). First report of Tomato brown rugose fruit virus in tomato in the United Kingdom. New Disease Report, 40, 12. http://dx.doi.org/10.5197/j.2044-0588.2019.040.012 [ Links ]
Toni, L.S., Garcia, A.M., Jeffrey, D.A., Jiang, X., Stauffer, B.L., Miyamoto, S.D. y Sucharov, C.C. (2018). Optimization of phenol-chloroform RNA extraction. MethodsX, 5, 599-608. https://doi.org/10.1016/j.mex.2018.05.011 [ Links ]
Van de Vossenberg, T.L.H., Visser, M., Bruinsma, M., Koenraadt, H.M.S., Westenberg, M. y Botermans, M. (2020). Real-time tracking of Tomato brown rugose fruit virus (ToBRFV) outbreaks in the Netherlands using Nextstrain. PLOS ONE, 15, e0234671. https://doi.org/10.1371/journal.pone.0234671 [ Links ]
Wang, H.M., Yin, W.C., Wang, C.K. y To, K.Y. (2009). Isolation of functional RNA from different tissues of tomato suitable for developmental profiling by microarray analysis. Botanical Studies, 50(2), 115-125. https://ejournal.sinica.edu.tw/bbas/content/2009/2/Bot502-01.pdf [ Links ]
Yan, Z.Y., Ma, H.Y., Han, S.L., Geng, C., Tian, Y.P. y Li, X. (2019). First report of Tomato brown rugose fruit virus infecting tomato in China. Plant Disease, 103, 2973. https://doi.org/10.1094/PDIS-11-18-1959-PDN [ Links ]
Appendix 1
A) Trizol protocol:
Macerate 100 mg of plant tissue with liquid nitrogen and add it immediately on an Eppendorf tube with 1 mL of cold Trizol. Give a pulse of vortex and leave them for 5 minutes on ice. Add 400 ml of chloroform and mix by inversion 7 times. Leave the tube on ice for 10 minutes. Centrifuge by 10 minutes at 13,000 rpm at 4 °C. Recover carefully the supernatant (approx. 400 μL) and transfer it in a clean Eppendorf tube. Add 1.5 V of cold isoamyl alcohol and leave the tube at -20 °C for 20 minutes. Centrifuge by 15 minutes at 13,000 rpm at 4 °C. Discard the alcohol taking care of the pellet at the bottom of the tube. Wash the pellet with 1 mL of cold ethanol at 90 % and centrifuge again by 5 minutes at 13,000 rpm at 4 °C. Discard all the ethanol taking care of the pellet and let it dry at room temperature. Resuspend and dissolve the pellet in 30 to 50 μL of RNAsa free distilled water.
B) CTAB 2% protocol:
Macerate 100 mg of plant tissue with liquid nitrogen and add it immediately on an Eppendorf tube with 700 mL of CTAB 2 % + PVP (1 %) + BME (0.2%). Give a pulse with a vortex. Incube at 55 °C in a water bath for 10 minutes. Add 470 mL of cold chloroform: isoamyl alcohol (24:1) and mix by inversion 7 times. Centrifuge by 10 minutes at 13,000 rpm at 4 °C. Recover carefully the supernatant (approx. 400 μL) and transfer it in a new Eppendorf tube. Add 1 V of cold isoamyl alcohol and 1/10 V of sodium acetate (3M) and mix by inversion. Left the tube at -20 °C for 20 minutes. Centrifuge by 15 minutes at 13,000 rpm at 4 °C. Discard the liquid taking care of the pellet at the bottom of the tube. Wash the pellet with 1 mL of cold ethanol at 90 % and centrifuge again by 5 minutes at 13,000 rpm at 4 °C. Discard all the ethanol taking care of the pellet and let it dry at room temperature. Resuspend and dissolve the pellet in 50 to 100 μL of RNAsa free distilled water.
C) CTAB2% + Trizol
Follow the steps 1 to 5 of section B) of appendix 1. Add 500 μL of Trizol and mix by inversion 7 times. Follow the steps 2 to 10 of section A) of appendix 1.
Received: August 23, 2023; Accepted: November 24, 2023











 texto en
texto en 


