Introduction
Mexico is an important producer of maize (Zea mays L.) worldwide. In 2022, 6,941,031 ha were planted to maize, and 27,549,000 tons were produced with a value of $ 10.1 USD billion (SIAP, 2024). The state of Zacatecas, located in North-Central Mexico, is gaining importance as a maize producer with 75,805 ha and a production of 325,230 tons (SIAP, 2024). The most significant economic losses caused by microorganisms in maize are due to fungus diseases whose causal agents are members of the genera Aspergillus, Giberella, Fusarium and Penicillium (Arispe et al., 2019). Nevertheless, another group of phytopathogens known as phytoplasmas has been reported, which are dispersed in Mexican territory, affecting crops of economic interest, among which maize stands out (Perez-Lopez et al., 2016). Since the 2021 crop season, maize plants have shown symptoms of dwarfism, color changes in leaves from green to white, general decline, and plant death in several commercial fields in Valparaiso, Zacatecas, México (Figure 1).

Figure 1 Maize plants (Zea mays ) showing symptoms of dwarfism, stunting and white leaves with green stripes and brown edges associated with phytoplasma infection.
Phytoplasmas are small, insect-transmitted, cell-wallless bacteria that cause numerous diseases in plant species worldwide that are important from an economic and environmental point of view (Bertaccini and Lee, 2018). Phytoplasmas have been found associated with multiple diseases in plants; the symptoms related to their infection may vary depending on the host, phytoplasma strain and environmental factors, and often include yellow/purple discoloration of leaves and shoots, virescence (greening of petals), big bud (conversion of floral organs into leaflike structures), proliferation of shoots, witches’ broom, stunting, general decline, and in some cases plant death (Ermacora and Osler, 2019). Phytoplasma diseases in maize plants have been reported in Bulgaria and Turkey, where Ca. Phytoplasma solani (16SrXII) was associated with empty grain bearing and leaf reddening (Çağlar et al., 2021; Genov et al., 2014); in Mexico and India, the presence of Ca. Phytoplasma asteris (16SrI) was related to leaf redness, yellowing, chlorotic stripes, and stunting (Perez-Lopez et al., 2016; Rao et al., 2017), while Ca. Phytoplasma trifolii caused yellowing and dwarfing in Iran (Zibadoost et al., 2015); in addition, the presence of maize plants was reported with bushy growth, short internodes, leaf discoloration, and yellowing disease associated with Bermudagrass phytoplasma (16SrXIV) (Çağlar et al., 2019). This study aimed to identify phytoplasmas related to this new disease in maize plants and its possible insect vector.
Materials and methods
Plant samples
A 20-hectare maize commercial field located in the municipality of Valparaiso, Zacatecas, Mexico (22° 46’ 0” N, -103° 34’ 0” W) was visited during Spring 2021 to find symptoms related to phytoplasma infection (dwarfism, color changes in the leaves from green to white, general decline and plant death). Fifteen symptomatic and five asymptomatic maize plants in vegetative development stage Vn were randomly selected and collected. Symptoms were recorded for each symptomatic plant, and its incidence was obtained in five points within the field by counting the number of diseased plants in a hundred consecutive plants at each location. The capture and counting of 50 adult members of the species Dalbulus maidis were carried out at the same time as the maize collection and taxonomic identification was carried out with a stereo microscope using dichotomous keys (Dietrich, 2005). Total DNA was isolated separately from 100 mg of leaf and 50 insects using the method described by Dellaporta et al. (1983) and stored at -20°C. DNA samples extracted from maize and insect tissues were used as templates for PCR.
Direct and nested polymerase chain reaction (PCR) for phytoplasma
PCR was performed in a programmable thermocycler (Applied Biosystems, Foster City, California, EUA) with the following parameters: i) for direct PCR primer pair P1/Tint (Smart et al., 1996): 95 °C for 5 min, 35 cycles of 94 °C for 1 min, 56 °C for 2 min and 72 °C for 2 min, and a final extension cycle of 72 °C for 5 min; ii) for nested PCR primer pair R16F2n/R16R2 (Gundersen and Lee, 1996): 95 °C for 5 min, 35 cycles of 94 °C for 1 min, 55 °C for 2 min, and 72 °C for 2 min, and a final extension cycle of 72 °C for 5 min. PCR products were analyzed by electrophoresis on a 1 % agarose gel and visualized by staining with ethidium bromide and UV illumination.
Cloning of PCR products, RFLP analysis, and sequencing of DNA
The 1.25 kb amplicons obtained from positive samples from plants and insects were ligated in the pGEM-T Easy vector (Promega, Madison, Wisconsin, EUA) as indicated by the supplier, and Escherichia coli Top-10 calcium competent cells were transformed by the heat shock method. Positive clones were assessed by restriction fragment length polymorphism (RFLP) analysis using EcoRI and HinfI enzymes. The digestion products were separated by electrophoresis on 2 % agarose gels, and PCR clones exhibiting different restriction fragment patterns were identified and sequenced at the National Laboratory of Agricultural, Medical and Environmental Biotechnology (LANBAMA) in San Luis Potosi, Mexico.
Phylogenetic and virtual RFLP analysis
A phylogenetic tree showing the 16S rDNA sequence of the phytoplasma isolated from insect and symptomatic maize plants, along with the 16S rDNA gene of 18 known phytoplasmas and Acholeplasma brassicae (used as outgroup) was constructed. The analysis of the nucleotide sequences obtained from this study was performed by comparison with available sequences at the NCBI database using the BLASTN software, and phylogeny analysis was performed using the ClustalV method of MegAling (DNASTAR software, Lasergene, London, UK). In addition, bootstrapping was performed 1000 times to estimate the stability and support of the branches.
Computer-simulated RFLP analysis of the 16S rDNA gene was performed on phytoplasma sequence isolated in this study. The aligned sequences were exported to the virtual RFLP analysis programs of the iPhyClassifier website (http://plantpathology.ba.ars.usda.gov/cgi-bin/resource/iphyclassifier.cgi) for computer-simulated restriction digestion and virtual gel plotting. Each aligned DNA fragment was digested in silico with 17 distinct restriction enzymes (AluI, BamHI, BfaI, BstUI (ThaI), DraI, EcoRI, HaeIII, HhaI, HinfI, HpaI, HpaII, KpnI, Sau3AI (MboI), MseI, RsaI, SspI and TaqI) that have been routinely used for RFLP analysis of the 16S phytoplasma rDNA gene (Reveles-Torres et al., 2018). After in silico restriction digestion, a virtual 3.0 % agarose gel electrophoresis image was plotted and captured as a device-independent pdf file. The RFLP analyses of phytoplasma sequence obtained from insects and maize plants were automatically compared with each 16S group and subgroup of reference phytoplasma strains.
Results and discusión
By evaluating the incidence of symptoms related to phytoplasmas infection, information on the risk for maize production could be obtained. In this study, the incidence of maize phytoplasmic disease (15 %) was lower than that reported for maize by Ca. Phytoplasma solani (16SrXII) affecting maize in Bulgaria (Genov et al., 2014) and Serbia (Jović et al., 2007) with a range from 30 to 85 % of incidence, while Ca. Phytoplasma asteris (16SrI) was reported in maize with an incidence range from 8 to 60 % in Mexico (Pérez-López et al., 2016) and 40 % in Brazil (Galvão et al., 2021). Reports of high incidence of plants with phytoplasmas refer to mature maize plants, whereas the 15 % incidence observed in this research could be due to maize plants being at an early stage of vegetative growth; furthermore, the range of incidence of symptomatic plants may increase later because at that time the interaction of Dalbulus maidis with maize crop was in the initial stage of development of the crop.
The identification of Dalbulus maidis individuals was based on their morphological characteristics. The adult male was 3.5-4 mm, and the female was 4.1 mm in length; the female has a dark ovipositor under the abdomen, adults are straw-yellow with two round black spots on the vertex of the head, and their rear wings are translucently extending beyond the tip of the abdomen. The identification of Dalbulus maidis was confirmed according to the taxonomic keys provided by Dietrich (2005). The presence of Dalbulus maidis is related to the presence of Maize bushy stunt, a disease caused by Ca. Phytoplasma asteris from Southern United States to Argentina (De Oliveira et al., 2011; Moya, 2001).
Typical phytoplasma DNA amplicons of 1227 bp were obtained by nested PCR using total DNA extract from the Dalbullus maidis mixture and, at the same time, in 14 out of 15 symptomatic DNA samples from maize plants. No amplification was observed in asymptomatic maize plants. BLAST analysis of the 16S rDNA sequences revealed that Phytoplasma from Dalbulus maidis and symptomatic maize plants shared 100 % sequence identity with each other and 99.7 % sequence identity with those in group 16SrI, strains of Candidatus Phytoplasma asteris. The Maize/ Dalbulus phytoplasma sequence (GenBank ON314873) was compared with those in the Iphyclassifier database (http://plantpathology.ba.ars.usda.gov/cgi-bin/resource/iphyclassifier.cgi), and the maximum identity was found with isolates of Candidatus Phytoplasma asteris.
The virtual RFLP pattern derived from the Maize/Dalbulus sequence was more similar to the reference pattern of group 16SrI and subgroup B (GenBank NC005303), with a pattern similarity coefficient of 1.0 (Figure 2). A phylogenetic tree was constructed with the Maize/ Dalbulus 16S rDNA sequence, with 18 other phytoplasmas representing distinct groups of of this pathogen (Figure 3). The Maize/Dalbulus phytoplasma strain was grouped with the Ca. Phytoplasma asteris and belongs to the 16SrI group according to the neighbor-joining method. Results show that Ca. Phytoplasma asteris is the phytoplasma present in the Dalbulus maidis population and is also related to the symptoms described in maize plants in this study.

Figure 2 Virtual restriction fragment length polymorphism (RFLP) patterns derived from in silico digestions of 16S rDNA gene fragment from Maize/Dalbulus phytoplasma sequence. Image of virtual gel of RFLP patterns of Maize/Dalbulus phytoplasma sequence with recognition sites for the following 17 restriction enzymes for simulated digestions: AluI, BamHI, BfaI, BstUI, DraI, EcoRI, HaeIII, HhaI, HinfI, HpaI, HpaII, KpnI, Sau3AI, MseI, RsaI, SspI and TaqI. MW: molecular weight standards from ɸX174 DNA digested with HaeIII.
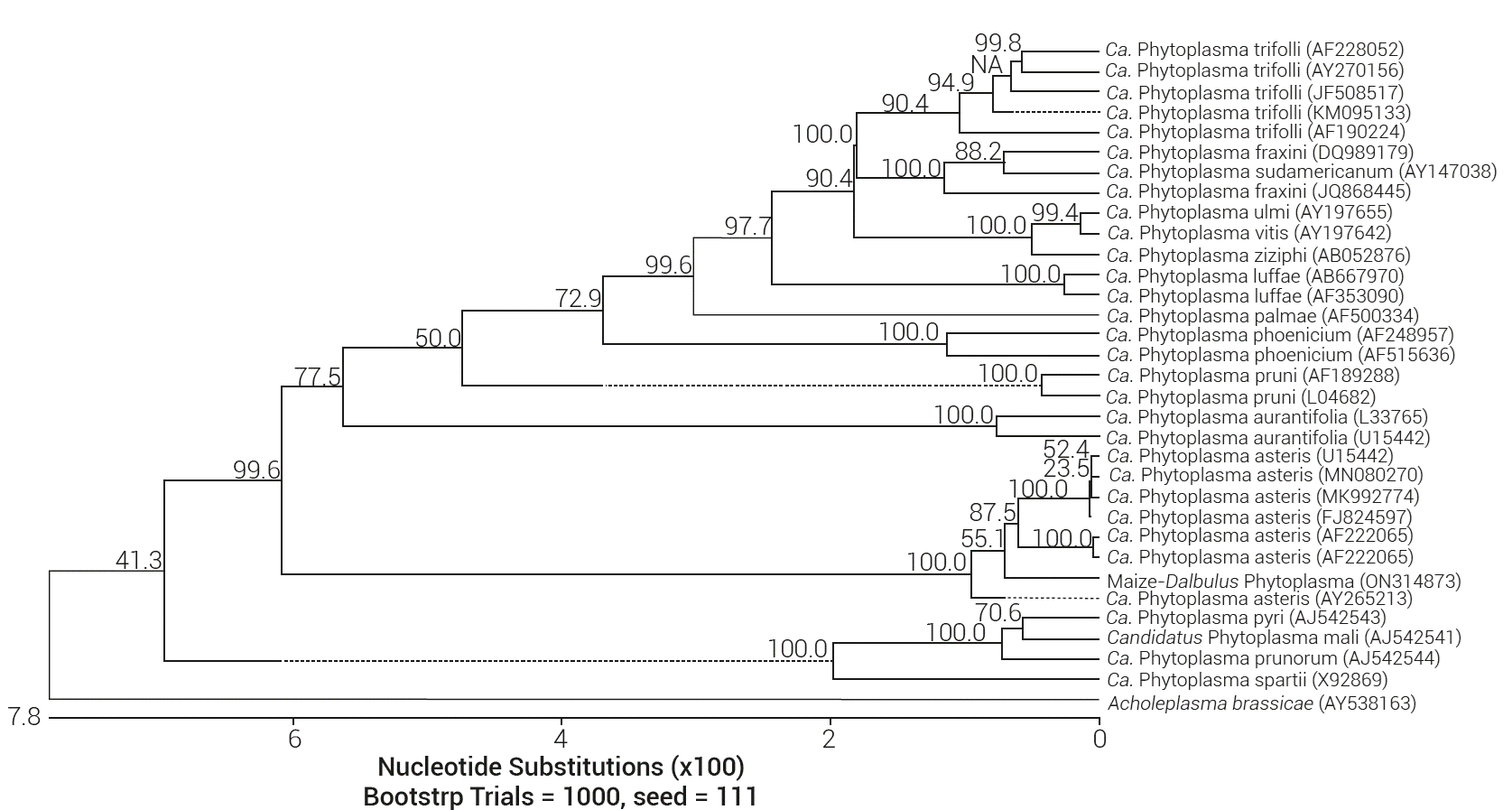
Figure 3 Phylogenetic distance tree constructed using the Clustal V alignment method (DNASTAR), which compares the 16S rDNA gene sequence from Maize/Dalbulus phytoplasma with sequences of 18 Candidatus Phytoplasma species. Acholeplasma brassicae was used as outgroup for the phylogenetic tree reconstruction. GenBank accession numbers for sequence are given in parentheses.
Phytoplasmas belonging to the 16SrI group are recognized as causal agents of maize bushy stunt disease in Mexico (Pérez-López et al., 2016), Iran (Zibadoost et al., 2016), India (Rao et al., 2017) and Peru (Gamarra et al., 2022), all of them induce symptoms such as reddish leaves, leaves with yellow or chlorotic stripes, stunting and chlorotic spots. Nevertheless, in this case the maize plants showed a new symptomatology characterized by dwarfism, stunting, and white leaves with green stripes and brown edges; except for the leaves with brown edges, similar symptoms are related to Ca. Phytoplasma asteris in wild grass belonging to the Poaceae family, as reported by Pérez-López and Dumonceaux (2017) in Mexico.
The presence of phytoplasmas belonging to the 16SrI group associated with diseases in maize could seriously constrain important economic crops in this region of Mexico; detection and identification of Ca. Phytoplasma asteris, as a potential pathogenic agent associated with maize crops in Mexico, facilitates the study of the epidemiology of the disease and contributes to the knowledge of the genetic diversity of phytoplasmas present in Mexico.















