Articles
Effect of Glutathione on the Stability, Dynamics and Catalysis of Two Different Classes of Glutathione Transferases from Taenia solium
-
Publication dates-
November 10, 2025
Jan-Mar , 2025
- Article in PDF
- Article in XML
- Automatic translation
- Send this article by e-mail
- Share this article +
Abstract.
In this work we compare the effect of glutathione (GSH) on the stability and dynamics of two different classes of glutathione transferases from Taenia solium, Ts26GST ((/( class) and Ts24GST (( class). The purpose was to explore why Ts24GST has low catalytic activity for the conjugation of glutathione (GSH) to hydrophobic substrates such as 1-chloro-2,4-dinitrobenzene (CDNB) compared to the very active Ts26GST but can instead use GSH to isomerize prostaglandin H2 to prostaglandin D2 by reducing its peroxide bond, a reaction described just for ( class of cytosolic GSTs. Using our recently deposited structure of Ts24GST in the Protein Data Bank, and a previous model for Ts26GST, we determined by molecular dynamics simulations that the presence of GSH decreased the number of intramolecular hydrogen bonds of Ts24GST and increased its radius of gyration, while in Ts26GST the effect was to increase its number of intramolecular hydrogen bonds without significantly changing its radius of gyration. Consistent with this, the experimental thermal stability of Ts26GST increased markedly while that of Ts24GST decreased in the presence of GSH, as determined by intrinsic fluorescence measurements. On the other hand, the binding site for the hydrophobic substrate (H site) of Ts24GST is wider than the H site of Ts26GST, with a 31 % greater solvent-accessible surface area.
Keywords::
Glutathione transferases, Taenia solium, molecular dynamics simulations, sigma class GST
Introduction
Glutathione transferases (GSTs) are a family of multifunctional enzymes whose main activity is to catalyze the conjugation of the tripeptide glutathione (GSH) to toxic compounds or xenobiotics to facilitate their excretion or further metabolism. In Taenia solium, GSTs participate in immunomodulation, assisting in the establishment of the parasite within its host, as well as in the transport of non-substrate compounds at the so-called ligandin site, among other functions. In this parasite the subfamily of cytosolic GSTs (cGSTs) possibly constitutes its main detoxification system, since it lacks other enzymes such as catalase, and has low activities of cytochrome P450 and glutathione peroxidase [1]. This is why T. solium cGSTs have been proposed as possible pharmacological targets to combat diseases caused by this parasite [2,3]. Three cGSTs classes have been identified in T. solium according to the classification established for mammalian GSTs [1]: the most abundant (/(-class GST called Ts26GST with a molecular mass of 25.9 kDa, the least abundant (-class GST with a molecular mass of 25.5 kDa denoted as Ts25GST and a moderately abundant 24.3 kDa (-class named Ts24GST; all of them are functional only in their dimeric form. The promoters of the mammalian and cestodes GST genes have xenobiotic regulatory elements and are induced by oxidative stress, infections, drugs, and xenobiotics such as H2O2 and proinflammatory agents [3-5].
-
1Acta Trop, 2008
-
2J. Mol. Graph. Model, 2020
-
3FEBS Open Bio, 2024
-
1Acta Trop, 2008
-
3FEBS Open Bio, 2024
-
5Acta Trop, 2021
While Ts26GST exhibits the highest specific GSH conjugation activity of the three enzymes, the corresponding activity of Ts24GST is the lowest and only a small fraction of the other two. This difference directly impacts how both proteins are purified from cell extracts, while Ts26GST is purified chromatographically with a glutathione Sepharose-4B affinity column, Ts24GST does not bind to this resin and must be purified by ion and filtration chromatographies. These two classes of GSTs differ quite a bit in their amino acid sequence as can be seen in Fig. 1. The percentage of identity is 25.3 %.
Thumbnail
Fig. 1
Amino acid sequence alignment of Ts24GST((( class) and Ts26GST ((/( class) classes. The alignment was performed with Clustal Omega [6]. Amino acid residues that contact the GSH molecule (3.5 Å or less away) are highlighted in blue (G site).
Amino acid sequence alignment of Ts24GST((( class) and Ts26GST ((/( class) classes. The alignment was performed with Clustal Omega [6]. Amino acid residues that contact the GSH molecule (3.5 Å or less away) are highlighted in blue (G site).
On the other hand, it has been reported that sigma class GSTs from different helminths also present prostaglandin synthetase activity. These enzymes catalyze the reduction of the peroxide bond of prostaglandin PGH2 with GSH to produce prostaglandin PGD2 [7]; this prostaglandin not only impact the reproduction and growth of the parasite, but also manipulates the host's immune system to its advantage. We have previously obtained the modeled atomic structure of Ts26GST show in Fig. 2(A), which was successfully used to find inhibitors by virtual screening [2]. The crystallographic structure of Ts24GST recently deposited by our group (PDB ID 9C0A), is also shown in Fig. 2(B). Even though both structures show the same folding pattern, conserving the G and H sites that bind GSH and the hydrophobic substrate to be conjugated, respectively, the superposition of both proteins (Fig. 2(C)) yields an RMSD of 7.4 Å considering only the Ca, which indicates that the arrangement of the similar secondary structures is some different in these GSTs.
-
7Trop. Med. Infect. Dis, 2024
-
2J. Mol. Graph. Model, 2020
Thumbnail
Fig. 2
Structures of Ts26GST and Ts24GST. (A) Modeled structure of Ts26GST previously used to find inhibitors by virtual screening [2]. (B) Crystallographic structure of Ts24GST. Glutathione molecules are shown in stick representation.
Structures of Ts26GST and Ts24GST. (A) Modeled structure of Ts26GST previously used to find inhibitors by virtual screening [2]. (B) Crystallographic structure of Ts24GST. Glutathione molecules are shown in stick representation.
In this work we compared these two structures, particularly their H sites, and analyzed the effect of GSH binding on the stability and mobility of both enzymes through MD simulations. We also determined the thermal denaturation profiles of both GSTs in the absence and presence of GSH by monitoring their intrinsic fluorescence and compared their parameters of enzymatic conjugation kinetics in order to try to explain the different activities of these two T. solium GSTs.
Materials and methods
Expression and purification of rTs24GST
Recombinant Ts24GST (rTs24GST) was expressed in E. coli and purified in two chromatographic steps, a cationic exchange followed by molecular exclusion, as previously reported [3]. rTs26GST was also expressed in E. coli and purified in one step using a GSH-Sepharose affinity column [2].
-
3FEBS Open Bio, 2024
-
2J. Mol. Graph. Model, 2020
Enzymatic transferase activity
The activity of rTs24GST was determined spectroscopically according to the method of Habig et al., 1974 [8], using 7.0 mM 1-chloro-2,4-dinitrobenzene (CDNB) dissolved in pure DMSO as the hydrophobic substrate with 5.0 mM reduced GSH in 100 mM Tris-HCl buffer, pH 7.4, in the standard assay. The reaction was carried out in a final volume of 1.0 ml at 25 °C, adding 5.0 µg of rTs24GST to initiate it. The increase in absorbance at 340 nm were monitored for three minutes, but just the first 20 second were used to determine the initial velocities. The reference cell was filled with the reaction mixture without enzyme, which allows to remove the contribution of the intrinsic CDNB-GSH conjugation. To measure the kinetic parameters for CDNB substrate, GSH was maintained at 5.0 mM while CDNB was varied in the range of 0.50 mM to 7.5 mM. Similarly, the parameters for GSH were obtained fixing the CDNB concentration at 7.0 mM and varying GSH concentration in the range of 0.5 to 8 mM. Changes in absorbance per minute were converted to (moles of conjugated substrate min-1 mg enzyme-1 using the molar extinction coefficient for CDNB ε340 = 9.6 mM−1 cm−1. Activity measurements were performed in triplicate.
-
8Methods Enzymol, 1981
Molecular dynamics simulations
3D structural model of the complex Ts24GST-GSH was obtained by superimposing the Ts24GST structure with the binary complex GST-GSH of the sigma class Bombyx mori GST (PDB ID 3VPQ) and removing the orthologous protein. Molecular dynamic simulations were performed using Gromacs 2023 version [9] with the OPLS-AA force field. The topology file for GSH ligand was generated using LigParGen Tool [10] and the charges were computed with MOE 2014 [11]. Previously, free Ts24GST and bound Ts24GST-GSH systems were analyzed with PropKa algorithm [12] to assign protonation states of protein ionizable residues considering a pH value of 7.4. The free and bound systems were solvated with Tip4p-( water model [13], Na+ and Cl- ions were added to neutralize the system and achieve a 0.15 M concentration. The energy of the systems was minimized for 10 steps using the steeper and descent gradient conjugate algorithm. Subsequently, the solvent and ions were equilibrated for 500 ps and 310 K at constant volume (NVT ensemble), and then 500 ps at a constant pressure of 1 bar (NPT ensemble), while the protein substrate was harmonically restrained using a force constant of 1000 kJ mol−1 nm−2. Electrostatic interactions were computed using the particle mesh Ewald (PME) [14] simulation method with a 1 nm short-range electrostatic cutoff. This same short-range cutoff was used for van der Waals interactions. For the temperature coupling, the solvated structure was divided into two groups (protein and non-protein) using velocity rescaling with a stochastic term. The isotropic pressure coupling was employed in the MD simulations by using the Parrinello-Rahman method [15] with a compressibility of 4.5 × 10−5 bar−1. During the simulation, constraints were deployed in all bonds using the LINear Constraint Solver (LINCS) algorithm [16], with parameters LINC-order of four. Each system was subjected to three independent 300 ns MD simulations. Solvent Accessible Surface Area (SASA) was determined with the GetArea server [17].
-
9SoftwareX, 2015
-
10Nucleic Acids Res, 2017
-
11Program for Molecular Modelling. Chemical Computing Group ULC; 910-1010, 2024
-
12,
-
13J. Phys. Chem. B, 2014
-
14Comput. Phys. Commun, 1996
-
15J. Appl. Phys, 1981
-
16J. Comp. Chem, 1997
-
17J. Comp. Chem, 1998
Thermal unfolding assays
Intrinsic fluorescence of both rTs26GST and rTs24GST at 2.0 (g mL-1 in Tris-HCl 50 mM, pH 7.4 were obtained at various temperatures in the 20 to 80 (C range in the absence and presence of 10 mM GSH. The emission spectra were recorded from 300 to 360 nm in a K2 ISS spectrofluorometer (Illinois, USA) using a bandwidth of 2.0 nm and a 280 nm excitation wavelength. The spectral center of mass (SCM) for each spectrum was calculated with the expression:
where ( is the wavelength at which the fluorescence intensity (FI) was determined. For better data comparison, SCM changes were expressed as the fraction of change with respect to the maximum shift observed in each system.
Results and discussion
Enzymatic characterization
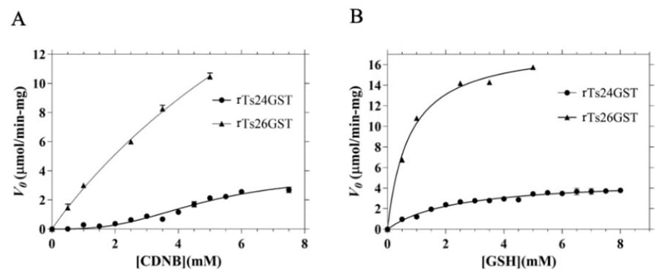
Both transferases, rTs24GST and rTs26GST were expressed and purified to homogeneity as verified by single bands in SDS-PAGE. The enzymatic parameters K m and V max for both substrates 1-chloro-2,4-dinitrobenzene (CDNB) and GSH in the conjugation reaction catalyzed by rTs24GST and rTs26GST [2] were determined from the data shown in Fig. 3. Hill´s equation was fitted to the CDNB data when the enzyme was rTs24GST in panel A, to obtain the kinetics parameters. For the rest of the kinetics, the Michaelis-Menten equation was used. Values obtained from these non-linear regressions are shown in Table 1.
-
2J. Mol. Graph. Model, 2020
Thumbnail
Fig. 3
Determination of the kinetic parameters of rTs24GST and rTs26GST by spectrophotometric assays at pH 7.4 and 25 (C. (A) Using 5.0 mM GSH and 0-7.5 mM CDNB. (B) Using 7.0 mM CDNB and 0-8.0 mM GSH. In all assays the concentration of rTs24GST or rTs26GST was 5.0 (g mL-1. Nonlinear regression analysis was fit to the mean values of three independent measurements, using the Hill equation for data corresponding to rTs24GST in panel (A) and the Michaelis-Menten equation for the rest of kinetics. The bars represent the corresponding standard deviations.
Determination of the kinetic parameters of rTs24GST and rTs26GST by spectrophotometric assays at pH 7.4 and 25 (C. (A) Using 5.0 mM GSH and 0-7.5 mM CDNB. (B) Using 7.0 mM CDNB and 0-8.0 mM GSH. In all assays the concentration of rTs24GST or rTs26GST was 5.0 (g mL-1. Nonlinear regression analysis was fit to the mean values of three independent measurements, using the Hill equation for data corresponding to rTs24GST in panel (A) and the Michaelis-Menten equation for the rest of kinetics. The bars represent the corresponding standard deviations.
Table 1
Enzymatic parameters of glutathione transferases rTs24GST and rTs26GST at pH 7.4 and 25 (C.
Enzymatic parameters of glutathione transferases rTs24GST and rTs26GST at pH 7.4 and 25 (C.
| Vmax(CDNB) ((mol min-1 mg-1) | Km(CDNB) (mM) | n* | Vmax(GSH) ((mol min-1 mg-1) | Km(GSH) (mM) | |
|---|---|---|---|---|---|
| rTs24GST | 3.13 ( 0.26 | 4.27 ( 0.27 | 3.1 ( 0.8 | 4.86 ( 0.15 | 2.05 ( 0.21 |
| rTs26GST† | 30.3 ( 2.9 | 9.4 ( 1.2 | 18.1 ( 0.63 | 0.80 ( 0.10 |
-
2J. Mol. Graph. Model, 2020
Comparing the kinetic parameters of rTs24GST and rTs26GST, it is evident how this last transferase is much more active than the rTs24GST for the conjugation of CDNB and binds GSH with more affinity. On the other hand, while the most active transferase shows a Michaelian kinetic behavior with both substrates [2], the kinetics of the less active GST presents a sigmoidal behavior (Fig. 3(A)) with the hydrophobic substrate CDNB in accordance with an homotropic activation. Given the low transferase activity of Ts24GST, its most relevant biological role may be another such as synthesizing prostaglandin D2, and when the detoxifying capacity of the other cytosolic GSTs is exceeded and its activity is required, it may be more efficient to activate it cooperatively through allosteric regulation.
-
2J. Mol. Graph. Model, 2020
Thermal unfolding
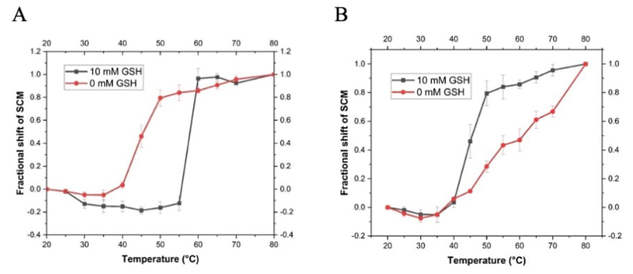
The results of intrinsic fluorescence measurements of both rTsGSTs at different temperatures, in the absence and presence of excess GSH ([GSH]/[TsGST] = 120) are presented in Fig. 4.
Thumbnail
Fig. 4
Thermal unfolding profiles of recombinant T. solium GSTs in the absence and presence of 10 mM GSH. Both enzymes (A) rTs26GST and (B) rTs24GST were used at 2.0 (g mL-1. Intrinsic fluorescence spectra were obtained at the different temperatures and their Spectral Center of Mass (SCM) were determined. Fractional shifts of the SCM were calculated at each temperature considering the maximum shift, observed at 80 (C, as the total change for each system.
Thermal unfolding profiles of recombinant T. solium GSTs in the absence and presence of 10 mM GSH. Both enzymes (A) rTs26GST and (B) rTs24GST were used at 2.0 (g mL-1. Intrinsic fluorescence spectra were obtained at the different temperatures and their Spectral Center of Mass (SCM) were determined. Fractional shifts of the SCM were calculated at each temperature considering the maximum shift, observed at 80 (C, as the total change for each system.
Ts24GST has only one Trp, W45, which is hidden from the solvent with an exposed area of only 11.9 % in the crystallographic structure. On the other hand, Ts26GST has 4 Trp (W9, W42, W202 and W207) which are also hidden inside the protein with an average exposure of 8.2 % in our structural model. In both proteins the intrinsic fluorescence emission shifted towards longer wavelengths (redshift) with increasing temperature, consistent with greater exposure of the fluorophores to the aqueous solvent caused by denaturation. It can be observed in Fig. 4(A) how the presence of GSH shifts the thermal unfolding profile of Ts26GST in more than 10 (C to higher temperatures, in an apparently two-state transition. On the other hand, the unfolding profiles of Ts24GST shown in Fig. 4(B) indicate that the presence of GSH facilitates the redshift of the protein's emission spectrum with temperature, making it less stable. It is not clear whether the unfolding process of Ts24GST in the absence of GSH is two-state or if there is some intermediate, but it is evident that the thermal transition becomes more cooperative with GSH binding.
Molecular dynamics simulations
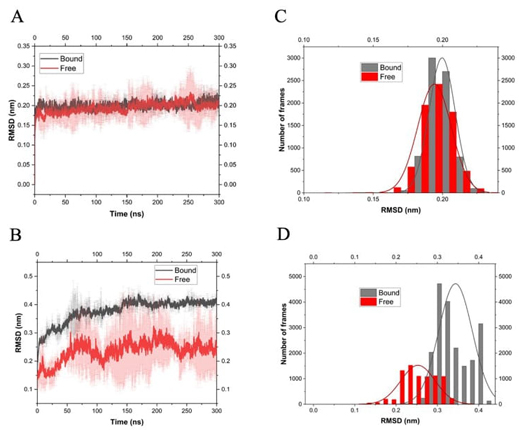
The mean Root Mean Square Deviation (RMSD) of three independent MD simulations in explicit solvent of free Ts26GST and bound to GSH at 310 K, as well as the corresponding histogram are presented in Fig. 5(A) and 5(C), respectively. As can be seen, both systems were equilibrated in the first tens of nanoseconds and behaved similarly throughout the simulation. On the other hand, the effect produced by bound GSH on the conformational dynamics of Ts24GST was much different (Fig. 5(B) and 5(D)). Simulations of the free enzyme showed greater variability than those performed with bound GSH. However, the final conformations of the complex presented a higher RMSD than that of the free protein, indicating that the presence of GSH causes a greater change in the structure of the protein. This change may be due to an expansion given the simultaneous loss of intramolecular hydrogen bonds (see Fig. 6(B)) and an increase in the radius of gyration of the macromolecule, as can be seen in Fig. 6(D). This result is consistent with the lower thermal stability of the complex rTs24GST-GSH observed in Fig. 4(B).
Thumbnail
Fig. 5
Root mean-square deviations (RMSD) of atomic position of C( in the trajectory of MD simulations performed using OPLS force field at 310 K. (A) Ts26GST in its free and bound state with GSH. (B) Ts24GST free and complexed with GSH. (C) and (D) show the histograms associated with (A) and (B), respectively.
Root mean-square deviations (RMSD) of atomic position of C( in the trajectory of MD simulations performed using OPLS force field at 310 K. (A) Ts26GST in its free and bound state with GSH. (B) Ts24GST free and complexed with GSH. (C) and (D) show the histograms associated with (A) and (B), respectively.
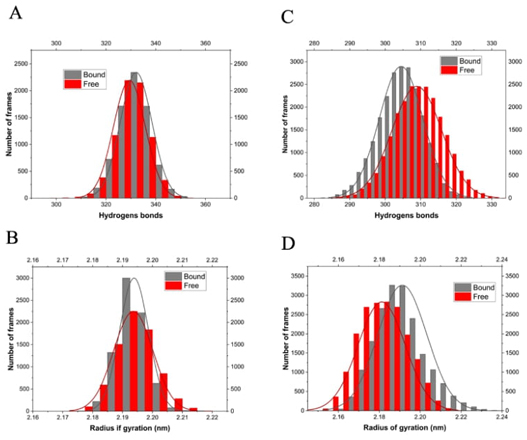
For the Ts26GST system, the presence of GSH in its binding site resulted in a small increase in the number of intramolecular hydrogen bonds, as shown in Fig. 6(A), which is consistent with its higher thermal stability (see Fig. 4(A)). The radius of gyration of the complex did not show significant changes compared with free Ts26GST (Fig. 6(C)).
Thumbnail
Fig. 6
Histograms of intramolecular hydrogen bonds present during the trajectory of the MD simulations of the free and bound enzyme (A-B) shown in Fig. 5. Histograms of the radius of gyration (C-D) for the same trajectories. (A and C) Ts26GST, (B and D) Ts24GST.
Histograms of intramolecular hydrogen bonds present during the trajectory of the MD simulations of the free and bound enzyme (A-B) shown in Fig. 5. Histograms of the radius of gyration (C-D) for the same trajectories. (A and C) Ts26GST, (B and D) Ts24GST.
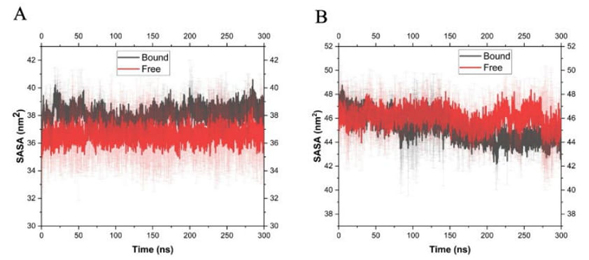
The difference observed in the Ca positions of the structures of Ts26GST and Ts24GST with an RMSD of 7.4 Å, (Fig. 2(C)) reflects the different way in which the secondary structures of these proteins are assembled to form the substrate binding sites. The hydrophobic substrate binding pocket (H site) of Ts24GST is wider and deeper than the corresponding site of Ts26GST. During the MD simulation, the solvent accessible surface area (SASA) of the H site of Ts24GST ranged around 46 nm2, while for the H site of Ts26GST the SASA changed around 35 nm2, which is 31 % more than binding surface in the Ts24GST (see Fig. 7). This agrees with the report on the comparative structural analysis of human (-, (-, and (-GST classes, where the H site for the latter is also the largest [18].
-
18Drug Metab. Rev, 2017
Thumbnail
Fig. 7
Variation of the solvent accessible surface area of the H sites of Ts26GST (A) and Ts24GST (B) during the MD simulations referenced in Fig. 5.
Variation of the solvent accessible surface area of the H sites of Ts26GST (A) and Ts24GST (B) during the MD simulations referenced in Fig. 5.
Conclusion
The binding of GSH to Ts26GST has effects on its structural dynamics, such as an increase in the number of intramolecular hydrogen bonds and a decrease in the radius of gyration observed by MD simulations, and on its thermal stability reflected in an increase of almost 10 °C in its Tm; while the binding of GSH to Ts24GST causes a decrease in the number of intramolecular hydrogen bonds, an increase in the radius of gyration and a decrease in its thermal stability. Furthermore, the low relative transferase activity and the larger size of the H site of Ts24GST suggest that the conjugation activity could be a secondary function and that there are other larger substrates that can be modified by GSH.
Acknowledgements
The authors acknowledge the financial support provided by the Consejo Nacional de Humanidades, Ciencias y Tecnologías (CONAHCYT-Frontier Science CF19-7397), the scholarship granted for the execution of the LlCSP postdoctoral stay (CVU 467703), and the doctoral scholarships for CSJ (CVU 937683) and RFL (CVU 763380). RMB received a postdoctoral scholarship from DGAPA-UNAM.
References
-
1.Torres-Rivera, A.; Landa, A. Acta Trop. 2008, 105, 99e102. Links
-
2. García-Gutiérrez, P.; Zubillaga R. A.; Téllez-Plancarte, A.; Flores-López, R.; Camarillo-Cadena, M.; Landa, A. J. Mol. Graph. Model. 2020, 100, 107707. Links
-
3. Miranda-Blancas, R.; Rodríguez-Lima, O.; García-Gutiérrez, P. FEBS Open Bio. 2024, 14, 726-739. DOI: https://doi.org/10.1002/2211-5463.13795. Links
-
4.Higgins, L. G.; Hayes, J. D. Drug Metab. Rev. 2011, 43, 92-137. DOI: https://doi.org/10.3109/03602532.2011.567391. PMID: 21495793. Links
-
5. Arbildi, P.; La-Rocca, S.; Kun, A.; Lorenzatto, K. R.; Monteiro, K. M.; Zaha, A.; Mourglia-Ettlin, G.; Ferreira, H. B.; Fernández, V. Acta Trop . 2021, 221, 105991. DOI: https://doi.org/10.1016/j.actatropica.2021.105991. Links
-
6. Madeira, F.; Madhusoodanan, N.; Lee, J.; Eusebi, A.; Niewielska, A.; Tivey, A. R. N.; Lopez, R.; Butcher, S. Nucleic Acids Res. 2024, 52, W521-W525. DOI: https://doi.org/10.1093/nar/gkae241. Links
-
7. Sánchez Pérez L. C.; Zubillaga, R. A.; García-Gutiérrez, P.; Landa, A. Trop. Med. Infect. Dis. 2024, 9, 85. Links
-
8. Habig, W. H.; Jakoby, W. B. Methods Enzymol. 1981, 77, 398-405. Links
-
9.Abraham, M. J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J. C.; Hess, B.; Lindahl, E. SoftwareX. 2015, 1, 19-25. Links
-
10.Dodda, L. S.; Cabeza de Vaca, I.; Tirado-Rives, J.; Jorgensen, W. L. Nucleic Acids Res . 2017, 45, W331-W336. Links
-
11.Molecular Operating Environment (MOE). Program for Molecular Modelling. Chemical Computing Group ULC; 910-1010 Sherbooke St. West, Montreal, Quebec, Canada, 2024; https://www.chemcomp.com/ Links
-
12.https://server.poissonboltzmann.org/pdb2pqr , accessed in May 2024 Links
-
13.Fuentes-Azcatl, R.; Alejandre, J. J. Phys. Chem. B. 2014, 118, 1263-1272. Links
-
14.Pollock, E. L.; Glosli, J. Comput. Phys. Commun. 1996, 85, 93. Links
-
15.Parrinello, M.; Rahman, A. J. Appl. Phys. 1981, 52, 7182-7190. Links
-
16. Hess, B.; Bekker, H.; Berendsen, H. J. C.; Fraaije, J. G. E. M. J. Comp. Chem. 1997, 18, 1463-1472. Links
-
17.Fraczkiewicz, R; Braun, W. J. Comp. Chem . 1998, 19, 319-333. Links
-
18.Mohana, K.; Achary, A. Drug Metab. Rev . 2017, 49, 318-337. DOI: https://doi.org/10.1080/03602532.2017.1343343. Links

![Amino acid sequence alignment of Ts24GST((( class) and Ts26GST ((/(
class) classes. The alignment was performed with Clustal Omega [6]. Amino acid residues that contact
the GSH molecule (3.5 Å or less away) are highlighted in blue (G
site).](/img/revistas/jmcs/v69n1//1870-249X-jmcs-69-01-39-gf1.jpg)
![Structures of Ts26GST and Ts24GST. (A) Modeled structure
of Ts26GST previously used to find inhibitors by virtual screening
[2]. (B)
Crystallographic structure of Ts24GST. Glutathione molecules are shown
in stick representation.](/img/revistas/jmcs/v69n1//1870-249X-jmcs-69-01-39-gf2.jpg)