Genes con efecto mayor sobre la fertilidad de ovejas. Revisión
Genes with major effect on fertility in sheep. Review
Carlos Luna Palomeraa, Rogelio Alejandro Alonso Moralesb
a División Académica de Ciencias Agropecuarias, Universidad Juárez Autónoma de Tabasco. Av. Universidad S/N, Zona de la Cultura, Col. Magisterial, Villahermosa, 86040 Tabasco, México. carlos.luna@ujat.mx. Correspondencia al primer autor.
b Departamento de Genética y Bioestadística. Facultad de Medicina Veterinaria y Zootecnia, Universidad Nacional Autónoma de México. México.
]]>Recibido el 21 de mayo de 2012.
Aceptado el 28 de septiembre de 2012.
Resumen
El estudio genético en razas de ovejas que naturalmente presentan altas tasas de ovulación (TO) y de prolificidad, ha permitido detectar la participación de varios genes. Entre ellos están los relacionados a la superfamilia del factor de crecimiento transformante β (BMPRIB, GDF9 y BMP-15), así como de otros genes con efecto mayor, tales como el "distal-less homeobox 3" (FecL), el receptor de estrógenos (ESR), el receptor de prolactina (PRLR) y el de las inhibinas (INHA y INHB). Sin embargo, las ovejas homocigotas para la mayoría de las variantes alélicas en BMP-15 o GDF9 son estériles, por lo que es importante el entendimiento de las bases genéticas y moleculares de estos polimorfismos, para su uso en programas racionales de mejoramiento genético con énfasis en la prolificidad. El polimorfismo en el gen de la melatonina (MTNR1) se ha asociado con la no estacionalidad reproductiva en ovejas. La selección en varias razas de ovejas para estas características ha reducido significativamente el anestro estacional, mostrando actividad reproductiva durante la primavera y el verano. Sin embargo, parece ser que estos genes en algunas razas pueden estar interactuando aditivamente, y una variante alélica en una raza puede no aparecer en otra, o bien dos al mismo tiempo, por lo que son variados los mecanismos genéticos que afectan la tasa ovulatoria y prolificidad.
Palabras clave: Genes de fecundidad, BMP-15, BMPR1B, ALK6, ESR, Inhibina, MTNR1, PRLR.
Abstract
The genetic study on sheep breeds with naturally high ovulation rate (OR) and prolificacy, has allowed to detect the participation of several genes. Among them there are the ones related to the transforming growth factor-beta superfamily (BMPRIB, GDF9 and BMP-15), as well as to other genes with major effect, such as the distal-less homeobox 3' (FecL), estrogen receptor (ESR), prolactin receptor (PRLR) and inhibin receptor (INHA and INHB). However, sheep homozygous for the majority of the allelic variants in BMP-15 or GDF9 are sterile, for which it is important the understanding of the genetic and molecular basis of these polymorphisms, for their use in rational programs of genetic improvement on prolificacy. Polymorphism in the melatonin gene (MTNR1) has been associated with non-reproductive seasonality in sheep. Selecting several sheep breeds for these characteristics has significantly reduced the seasonal anoestrus, showing reproductive activity during spring and summer. It seems that these genes in some breeds may be interacting additively and an allelic variant in a breed may not be present in other, or two be present at the same time; therefore, the understanding of the genetic mechanisms that affect ovulation rate and prolificacy are important for the selection of reproductive traits.
]]> Key words: Fecundity genes, BMP-15, BMPRIB, ALK6, ESR, Inhibin, MTNR1, PRLR.
INTRODUCCIÓN
La oveja es una especie diversa con gran variabilidad genética en características fisiológicas, fertilidad y el desarrollo muscular, y se ha constituido en un importante modelo experimental en el estudio de los genes que controlan estos mecanismos(1). Sin embargo, poblaciones no seleccionadas que incluyen a las locales y nativas, están amenazadas y sufren erosión genética(1). Algunas razas son portadoras de muchas variantes y es una buena razón para su conservación y estudio. La fertilidad en las hembras está influida en gran medida por la tasa ovulatoria (TO), definida por el número de ovocitos liberados durante un ciclo reproductivo; estos responden a su vez a un complejo intercambio de señales endocrinas entre la glándula pituitaria y el ovario, asociadas a señales paracrinas y autocrinas, así como a factores de señalización en las células foliculares, el ovocito y las células somáticas adyacentes controlados genéticamente(2-9).
Recientemente se han encontrado variaciones importantes en la tasa de ovulación. Estas variantes genéticas en las ovejas se nombran como Fec (de fecundidad) y a los diferentes alelos se les asigna adicionalmente una letra significativa. Así, FecB se refiere al alelo que aumenta la fertilidad en la raza Booroola. Se han identificado polimorfismos en genes específicos como en el gen GDF-9 (growth differentiation factor-9) llamado también FecG(10,11), en el gen BMP15 (bone morphogenic protein 15) o FecX (2,10,12,13), así como en el gen ALK-6 que produce la proteína activin-like kinase(6,14). Igualmente, se han identificado polimorfismos en el receptor tipo IB de la proteína morfogénica de hueso (BMPR-IB), conocido como FecB(14-18).
Este conjunto de genes tienen un efecto mayor en la regulación de la tasa de ovulación y fertilidad, y sus polimorfismos se han estudiado ampliamente en varias razas ovinas(3,7,19). El estudio de estos genes ha dado luz al conocimiento y a la exploración de las vías biológicas implicadas en el control de la ovulación en mamíferos en general(20).
Se han encontrado genes candidatos que pueden impactar positivamente la prolificidad y fertilidad en ovejas. El gen de la inhibina alfa (INHA) ha sido propuesto como gen con efecto mayor sobre la prolificidad, algunos alelos en cabras incrementan el tamaño de la camada(21,22). El gen de la inhibina beta (INHB), también se ha propuesto con efecto mayor sobre prolificidad e incrementa el tamaño de camada en algunas razas de ovejas(23).
Se ha encontrado variación en los genes para receptores de melatonina (MTNR1) y prolactina (PRLR), involucrados en el control de la estacionalidad reproductiva, lo que puede permitir la implementación de programas de selección intensiva ligada a marcadores moleculares fuera de épocas reproductivas(24,25). Probablemente estos genes pueden estar implicados en la variabilidad estacional reportada en ovinos de pelo en México.
El estudio en conjunto de estos genes y sus mecanismos genéticos a nivel molecular permitirá comprender la forma como participan en la regulación de la prolificidad y fertilidad.
El propósito de este trabajo es hacer un recuento de los principales genes identificados, que participan en el control de la tasa de ovulación, la prolificidad y la estacionalidad reproductiva en la oveja, así como describir los mecanismos genéticos que están actuando a nivel molecular.
]]> Control genético de la tasa ovulatoria y prolificidadLa variación en la fertilidad, prolificidad y tasa ovulatoria se ha observado en diferentes razas ovinas, lo cual está genéticamente regulado por un conjunto de genes con acción mayor o aditiva, los cuales pertenecen a la súper familia de los factores de crecimiento transformante beta (TGF-β)(2). Los cambios en las secuencias de los alelos en los genes, incluyen cambios en un nucleótido por otro, que llevan a su vez a cambios en la secuencia de aminoácidos. lo cual influye en la función de la proteína debido a cambios en sus actividades y plegamiento. Así mismo, se han descrito en estos genes indeles y mutaciones sin sentido(26,27). Otro tipo de cambios en la secuencia pueden afectar los niveles de expresión génica o la estabilidad del ARN(18,19).
Las principales mutaciones o polimorfismos que incrementan la tasa de ovulación y afectan la fecundidad se ubican en el gen BMP-15 (razas Inverdale, Hanna, Belclare, Cambridge y Lacaune), el gen del receptor BMPR-IB (raza Booroola) y el gen GDF-9; adicionalmente al parecer hay otros genes implicados que se abordarán en los apartados siguientes de esta revisión.
Gen de la proteína morfogénica de hueso (BMP-15)
También conocido como inverdale (Fec XI), fue uno de los primeros genes reportados con efecto mayor sobre la tasa de ovulación en ovejas Romney(2,28). El locus FecXI se encuentra localizado en una región a 10 centimorgans (cM) del centro del cromosoma X. La secuenciación del gen ovino muestra ser similar a la del humano (~80 %), ratón (~76 %) y rata (~75 %), siendo de una longitud de 1,179 pb contenida en dos exones(3).
La proteína BMP-15 actúa a través de una cascada de proteínas señalizadoras (vía Smad), que son responsables de un enorme rango de comportamientos fisiológicos a nivel celular, incluyendo el desarrollo y maduración de ovocitos (9,29-31). Los efectos biológicos de los BMPs son mediados por receptores celulares de superficie específicos tipo I y tipo II, estructuralmente similares; ambos poseen actividad serina/treonina cinasa intrínseca, cuya estimulación inicia cascadas de señalización intracelular que regulan eventos transcripcionales esenciales para la proliferación y la diferenciación celular(32). Existen dos receptores BMP tipo I, el BMPR-IA (o ALK-3) y BMPR-IB (o ALK-6), y de los receptores tipo II sólo se ha identificado el BMPR-II. Los receptores BMP tipo I y II se unen al ligando, pero la señal de transducción por el receptor BMP, requiere la formación de un complejo heterodimérico entre ambos receptores. Una vez formado el complejo BMPR-ligando, el receptor tipo II, el cual tiene actividad cinasa constitutiva, fosforila y activa al receptor tipo I, el cual en su momento dispara los eventos de señalización BMP. El ARN mensajero (ARNm) de BMPs y sus receptores han sido localizados en ovocitos de mamíferos(9).
En este proceso las moléculas Smad señalizadoras a nivel intracelular de BMP cumplen una función muy importante; en ratones Knockout las hembras doble condición Smad1 y Smad5 son infértiles y desarrollan tumores metastásicos en las células de la granulosa(33). En ovejas chinas de la raza Hu, la abundancia de ARNm de Smad4 en los folículos antrales es mayor en ovejas con alta prolificidad (HF) comparada con ovejas de baja prolificidad (LF), lo cual sugiere que la diferencia puede estar asociada con la tasa de ovulación, y atribuido al hecho de que Smad4 es un par común esencial tanto para la vía TGF-β/activina como para BMP(9,34).
Los cambios nucleotídicos en el gen BMP-15 producen un incremento de al menos 1.0 en la tasa ovulatoria y 0.6 en el tamaño de camada(2,33), asociado a una mayor sensibilidad de los folículos a la FSH; sin embargo, la BMP-15 se expresa en el ovocito de varias especies y es un gen candidato para el fenotipo asociado con el tamaño de camada en ovejas(3,35).
El cambio del alelo FecXH consiste en una transición de una citocina (C) por una timina (T) en el nucleótido 67 (C67T) de la región codificante para el péptido maduro (aminoácido 291 del péptido no procesado), que introduce un codón de paro prematuro en la secuencia (Cuadro 1), lo cual muy probablemente resulta en una pérdida de la función de la BMP-15 en los homocigotos para esta variante FecXH(3).
]]> El cambio de base en el gen FecXI consiste en una transversión de una sola T por adenina (A) en la posición 92 (T92A) del gen, resultando en la sustitución de una valina por un ácido aspártico en una región altamente conservada de la proteína(35). Este cambio parece debilitar la habilidad de la BMP-15 para formar dímeros, e interfiere con la acción biológica de la BMP-15 en ovejas homocigotas para esta variante FecXi(3).El cambio de base en el alelo FecXG es una transición de una C por T en el nucleótido 718 (C718T), e introduce un codón de paro prematuro en el aminoácido 239 de la proteína no procesada(7) (Cuadro 1), y presumiblemente resulta en pérdida completa de la función del BMP-15(35,36). La variante introduce un cambio de una arginina por una histidina, la cual sustituye un grupo polar cargado básico con otro, y ocurre en una posición antes del sitio de corte en el péptido maduro, e invariablemente afecta la actividad de la proteína madura(37).
El polimorfismo sencillo en el gen FecXB también es una transición de una G por T en el nucleótido 1100 (G1100T), sustituyendo una serina por una isoleucina en la posición 367 del aminoácido del péptido no procesado (aminoácido 99 de la proteína madura, Cuadro 1)(7).
El cambio de base en el gen FecXL afecta al gen BMP-15, y se ha demostrado que segrega en la población de ovejas Lecaune incrementando la tasa ovulatoria en cerca de 2.0. Este alelo (Cuadro 1) consiste en un cambio de un aminoácido cisteína por una tirosina en la posición 53, responsable de una alteración dramática en el procesamiento del péptido BMP-15(37).
En la raza Aragonesa(38), el alelo FecXR consiste en una deleción de 17 pares de bases (pb) que lleva a una alteración en la secuencia de aminoácidos, introduciendo un codón de paro prematuro, por lo que no se sintetiza el péptido maduro. Esta alteración incrementa la prolificidad en ovejas heterocigotas con 2.66 corderos por parto, comparada con 1.36 corderos por partos en hembras sin la mutación (Cuadro 2), pero en las ovejas homocigotas causa esterilidad debido a fallas ováricas primarias(13,38,39). Por lo tanto, la selección de hembras con este tipo de polimorfismos incrementa la tasa de partos gemelares o múltiples en heterocigotas, pero también incrementa la probabilidad de homocigotos, resultando en corderas freemartins y por tanto estériles(39-41). Las ovejas homocigotas presentan ovarios pequeños y aplanados con folículos que no se desarrollan hasta la etapa de folículos primarios(2,10,6,38), lo que resulta en completa infertilidad(10,13).
Por otra parte, ya se han iniciado estudios para evaluar las ventajas potenciales que ofrece el uso estratégico de ovejas con estas mutaciones en la producción de corderos, asociado a estrategias de manejo reproductivo y nutricionales(42,43), las cuales ofrecen beneficios potenciales en la producción de corderos.
Recientemente se han reportan alelos del gen BMP-15 en las raza Aragonesa en España, las denominadas cola grasa de Malasia (razas Afshari, Baluchi, Makui y Mehraban) y en las razas Banpalas y del valle de Kashmir en la India, asociados a mayor prolificidad en las ovejas heterocigotas(39,42-46).
Por tanto, la proteína madura BMP-15 es un factor clave en la determinación de la tasa de ovulación y fertilidad de mamíferos(47) y, en ovejas heterocigotas con cualquier tipo de polimorfismo del gen BMP-15 (FecX1, FecXH, FecXG, FecXB, FecXL y FecXR), se ha documentado existe una alta tasa de ovulación, comparadas con las del gen tipo silvestre(6,13,38-40).
Gen activin-like kinasa (ALK6)
]]> También conocido como receptor tipo IB de la proteína morfogénica de hueso (BMPRIB), y Booroola (FecB)(14-17,33). Fue uno de los primeros locus reportados con efecto mayor sobre la tasa ovulatoria en ovejas Merino(41), cuyo efecto es aditivo. El incremento en la tasa de ovulación en portadoras FecBB, fisiológicamente está asociado con mayores concentraciones FSH y con una maduración precoz de un gran número de folículos antrales, los cuales son de menor tamaño en comparación con el tipo silvestre(6).El alelo FecB se ha reportado en algunas de las razas más prolíficas en el mundo, tal como el Merino Booroola Australiano(16), Garole Indian(17) cuyo gen ha mostrado un alto grado de polimorfismo(48), Javanese Indonesia(17) y en ovejas Small-Tailed de las razas Han y Hu en China(18). El tamaño de camada y la tasa de ovulación en ovejas incrementan con el número de copias para el gen ALK6(7). Las ovejas que heredan una copia del gen Booroola de sus padres producen cerca de 1.5 óvulos más, y dan un cordero extra por parto (Cuadro 2). Las ovejas homocigotas producen cerca de tres óvulos extras, resultando en 1.5 corderos más por parto(7).
Los BMPs pueden unirse al receptor tipo II (BMPRII) y activar al receptor BMP tipo I (BMPRIA y BMPRIB). Se ha reportado que estos receptores y sus ligandos están presentes en folículos de bovinos, porcinos y ovinos, y que juegan un extenso rol en el desarrollo y funcionalidad del folículo(49-51). Las moléculas señalizadoras participan sincronizadamente con otras para afectar el fenotipo de la tasa de ovulación vía BMP/Smad, y otras moléculas relacionadas (GDF9, TGF-βRI), han demostrado tener una cercana relación en la prolificidad de ovejas de la raza Hu(9).
El polimorfismo en el gen BMPRIB (cambio de adenina por guanina en la posición 746 de la secuencia nucleotídica del gen) está significativamente asociado con el fenotipo de hiperprolificidad en ovejas Booroola (FecB). Sin embargo, Xu et al(9) reportan ovejas de baja fertilidad (LF) con el polimorfismo BMPRIB, lo que sugiere que otros factores pueden participar inhibiendo la expresión del fenotipo del alelo BMPRIB, lo cual fue asociado a mayor abundancia de RNA mensajero (RNAm) de receptores BMPRIB y BMPRII en los folículos antrales de ovejas altamente fértiles (HF).
Gen del factor de diferenciación del crecimiento (GDF9)
También conocido como FecG, y se encuentra en el cromosoma 5 de la oveja(19). El punto de cambio en la secuencia nucleotídica identificado en la oveja Belclare (FecGH) resulta en un aminoácido no conservado en una región que se piensa interactúa con el receptor ligador de dominio tipo 1(6).
El gen autosomal GDF9 tiene un patrón de herencia sobre dominante, su rol esencial es controlando el crecimiento folicular por su influencia en la función de la célula de la granulosa(10,35). En los homocigotos con el alelo GDF9 el desarrollo ovárico desde la etapa preantral hasta la de crecimiento folicular es diferente al del tipo silvestre(6). Los animales homocigotos para este alelo no ovulan y son infértiles, mientras que en los animales heterocigotos la tasa de ovulación promedio es de 2 comparados con las de tipo silvestre (Cuadro 1), por lo que la ventaja en fertilidad, prolificidad y tasa ovulatoria es sólo para los heterocigotos(33,36).
Recientemente, Silva et al(11) en ovejas de raza Santa Inés con antecedentes de partos gemelares y trillizos, describen por primera vez el polimorfismo de nucleótido sencillo (SNP) del gen GDF9, denominado FecGE (Embrapa), que no produce esterilidad en las hembras homocigotas. Este gen incrementa la tasa ovulatoria en homocigotos en 82 % del número de cuerpos lúteos (CLs), promedio de 2.22 CLs por oveja, y 96.3 % en partos múltiples comparada con las heterocigotas (E/+) y el tipo silvestre (Cuadro 2). Este alelo marca un efecto aditivo diferente al de sobredominancia de los alelos FecGH y FecX descritos hasta hace poco, y con ovocitos fértiles en los genotipos E/E, marcando una nueva acción fisiológica para el gen GDF9.
Aunque en algunas ovejas hiperprolíficas no se reportan los polimorfismos en el gen GDF9, se ha encontrado que los productos genéticos de BMPRII y TGF-βRI están involucrados en el control negativo de la señalización del GDF9(52), con diferencias significativas en los niveles de ARNm de GDF9, BMPRII y TGF-βRI en los folículos antrales en ovejas HF y LF(9).
]]>Gen Lacaune (FecL)
Es un gen autosómico recesivo identificado en la raza Francesa Lacaune de línea cárnica y lechera. El locus FecL se encuentra en el cromosoma 11 ovino, entre un intervalo de 2.1 cM entre los marcadores DLX3 y BM17132(20), o entre los marcadores BM17132 y FAM117A del mismo cromosoma, lo cual corresponde a un bloque sinténico de 1.1 megabases presente también en el cromosoma 17 humano. Este segmento contiene 20 genes(53).
El cambio en la secuencia de ADN en este alelo resulta en un cambio del aminoácido cisteína por una tirosina en la posición 53 (Cuadro 1), responsable de una alteración asociada al procesamiento del péptido BMP15(37). A diferencia de todos los genes Fec identificados e implicados en la regulación de la TO pertenecientes al sistema de señalización BMP(7), ninguno de los 20 genes del locus FecL corresponden a ligandos conocidos, receptores, antagonistas o moléculas señalizadoras del sistema BMP o TGFβ(20).
Una estrategia de secuenciación masiva de fragmentos de PCR largos que cubren el locus FecLL en ovejas portadoras y no-portadoras definió un nuevo gene implicado en fecundidad llamado B4GALNT2, el cual codifica para una enzima de glicosilación que no está relacionada a la familia de los BMP. La elevada fertilidad de las ovejas Lacaune se explica por a sobreexpresión de B4GALNT2 en el ovario llevando a una glicosilación atípica de la inhibina(54).
La influencia de FecLL en la TO es aditiva, incrementando 1.5 óvulos con una copia y 3.0 óvulos con dos copias del alelo (Cuadro 2). Inicialmente se pensaba que en ovejas de carne la tasa de ovulación estaba bajo control genético de gen FecL(12), pero posteriormente se identificó el alelo FecXL, el cual afecta al gen BMP15, y que se demostró segregar en la misma población, incrementando la tasa ovulatoria en cerca de 2.0 óvulos.
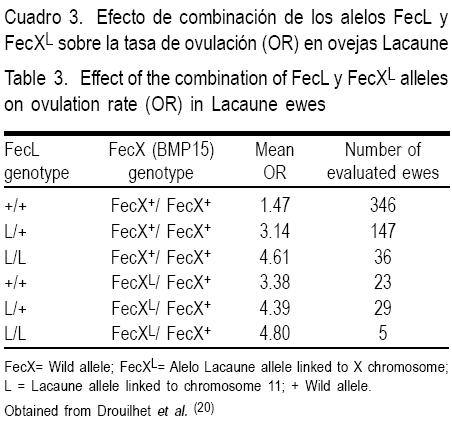
Estudios del efecto simultáneo de los locus FecXL y FecLL sobre la tasa ovulatoria en ovejas(20) muestran (Cuadro 3) que en la ausencia de la mutación FecXL, una copia del alelo FecLL incrementa la TO en cerca de 1.5 óvulos, y dos copias del FecLL incrementan la TO en 3.0. Este hallazgo confirma el efecto aditivo en la TO entre estos alelos. Por otra parte, una copia del alelo mutante FecXL incrementa la TO en 1 .9 óvulos. Cuando están presentes simultáneamente en estado doble heterocigoto, las dos mutaciones incrementan la TO en 2.9 óvulos. Por tanto, aquellas ovejas portadoras de la mutación en los loci FecL y FecX tienen mayor TO que aquéllas con una sola mutación(20).

Se han encontrado nuevas vías fisiológicas implicadas en la regulación de la tasa ovulatoria por el gen FecLL. Las diferencias más importantes entre ovejas homocigotas (L/L) y de tipo silvestre (+/+) radican en el fenotipo ovárico y perfiles endocrinos. Los estudios concluyen que: a) el número de folículos medianos (3.5 y 5.5 mm) y grandes (≥ 6 mm) fue mayor en ovejas L/L, y el tamaño del folículo dominante es de 1 mm menos en L/L; b) la relación entre la concentración de progesterona y testosterona en folículos grandes fue mayor en ovejas L/L, pero la relación entre progesterona y estrógenos fue menor; c) los niveles de ARNm del receptor de FSH incrementaron en las células de la granulosa de ovejas L/L(55).
]]>Gen del receptor para estrógenos (ESR)
El ESR es un miembro de la súper familia de receptores nucleares del factor de transcripción ligando - activado. Se encuentra ubicado en el cromosoma 17 en bovinos y ovejas, en el 8 en cerdos y en el 4 en humanos(19). Este gen está relacionado con el incremento en el tamaño de camada en cerdos(47), pero en ratones knockout el ESR muestra regulación caótica de la LH, anovulación e insensibilidad del útero a estrógenos(56).
En ovejas este gen puede jugar un rol importante en la prolificidad(57), ya que Bi et al(58) estudiando el polimorfismo del gen del ESR en ovejas hiperprolíficas Small Tail en las razas Han, Hu y German Mutton Merino, encontraron tres genotipos (AA, AB y BB) con frecuencias génicas para el alelo A de 0.846, 0.672 y 0.786, y para el alelo B de 0.154, 0.328 y 0.214, respectivamente. El polimorfismo incluye un cambio de una cisteína por una guanina en la posición 363 de la secuencia nucleotídica (C363G) en el exón 1 del gen ESR en ovejas Small Tail Han, las cuales se asociaron con 0.5 y 0.7 corderos más en los genotipos AB y BB, respectivamente, comparadas con AA. Por lo anterior, el gen ESR tiene un efecto mayor ligado a la prolificidad de estas ovejas. Sin embargo, en ovejas de otras razas aunque se informa de genotipos AB y BB, no se ha encontrado polimorfismo o asociación de este gen con la prolificidad(57,59). Lo anterior lleva a concluir que en las razas de ovejas hiperprolíficas existen diferentes genes que controlan los mecanismos de desarrollo, maduración y ovulación que caracterizan a este fenotipo.
Gen de inhibina
La inhibina es una glicoproteína perteneciente a la súper familia del factor-β de crecimiento transformante, que suprime selectivamente la síntesis y secreción de la FSH(60,61). Está compuesta de dos subunidades, α y β, unidas por enlace disulfuro; se han identificado dos tipos de inhibinas, compartiendo una subunidad α común pero con diferentes subunidades β (βA o βB)(62), dando lugar a las inhibinas αβA (INHA) y αβB (iNHBB). La subunidad a se localiza en los cromosomas 2 en humanos, bovinos y ovinos, y en el 15 en cerdos; y la subunidad βA se localiza en los cromosomas 7 en humanos, 4 en bovinos y ovinos, y 18 en porcinos. La subunidad pB se localiza en los cromosomas 2 en humanos, bovinos y ovinos, y en el 12 en porcinos(19).
El gen INHA tiene efecto aditivo significativo en la prolificidad de ovejas, y es posible que INHA, INHBA e INHBB tengan un efecto importante en el tamaño de la camada(63-65). Por otra parte, un estudio demostró que el gen INHA muestra polimorfismo de segmentos de restricción (RFLP) detectado con la endonucleasa TaqI, siendo el alelo A asociado al tamaño de camada en ovejas(66).
Hasta ahora no se ha documentado polimorfismo para el gen INHA en ovejas; sin embargo, en poblaciones de cabras de las razas Matou, Nubi, Boer y Haimen en China(21) se encontró la participación del gen INHA en la prolificidad, identificando un cambio de base de una guanina por una adenina en la posición 284 (G284A), indicando además que el alelo G se hereda de forma dominante. Este estudio demostró que el efecto del genotipo de INHA en el tamaño de camada fue mayor para el genotipo GG, seguido por AG y AA, por lo que el polimorfismo asociado al gen INHA puede estar involucrado en el control del tamaño de camada en cabras.
En China, He et al(22) en cabras Haimen, Boer y Huanghuti encontraron un cambio silente de una cisteína por una timina en la posición 865 (C865T) en el exón del gen para INHA, pero que no induce cambio en el aminoácido. Esto preliminarmente demuestra que existe una asociación entre el genotipo BB y la presentación de estro durante todo el año comparado al estro estacional. Por tanto, el gen INHA puede ser considerado como un gen candidato para alta prolificidad en cabras.
]]> Se ha encontrado que el gen INHBB tiene un efecto significativo sobre el tamaño de camada en algunas razas de ovejas(63), y por su importante rol en la reproducción es considerado un posible gen candidato para la prolificidad en ovejas. Al respecto Chu et al(23) estudiaron su efecto sobre la prolificidad en ovejas Small Tail de las razas Han y Hu, detectando polimorfismo dentro del exón 2 del gen INHBB. En las ovejas Hu se encontró un cambio de base de una adenina por una guanina en la posición 276 (A276G) en el exón 2 en el genotipo BB comparada con el AA, la cual no causó cambio de aminoácido, ya que estaba dentro de la región 3' de la región no transcrita (UTR) del gen INHBB. Las ovejas con genotipo BB mostraron 0.58 corderos más que aquéllas con genotipo AA, lo cual preliminarmente muestra que el gen INHBB tiene un efecto significativo en el tamaño de camada en ovejas Hu, pero no en las Small Tail Han, lo que implica que existen bases genéticas moleculares diferentes para la hiperprolificidad entre razas.
Gen del receptor de prolactina (PRLR)
El gen de PRL se encuentra en el cromosoma 16 porcino y ovino(67,68). La prolactina es una hormona polipeptídica de la pituitaria anterior, involucrada en muchos procesos endocrinos esenciales para un adecuado comportamiento reproductivo(69). Su acción es mediada por su receptor (PRLR), cuyo gen es miembro de la familia del receptor de la hormona de crecimiento/prolactina, el cual contiene regiones con secuencias idénticas(70). Se ha encontrado que este gen tiene un efecto mayor sobre la alta prolificidad asociado al exón 10 en cabras Jining Grey(71) y en ovejas Small Tail de raza Han, o está ligado al gen responsable de tal fenotipo(34).
En estas ovejas el gen del PRLR es polimórfico, encontrándose los genotipos AA, AB y BB. El genotipo BB mostró dos cambios de bases de una timina por una citocina en la posición 84 (T84C), y de una timina por una guanina en la posición 174 del gen (T174G) comparado con el genotipo AA de un producto de 176 pb. El tamaño de camada para el genotipo BB fue de 0.64 a 0.75 y para AB de 0.44 a 0.54 más corderos en comparación con los genotipos AA, concluyendo que el alelo B está positivamente correlacionado a mayor tamaño de camada en ovejas Small Tail Han(34). En relación al gen PRL y la prolificidad en ovejas Small Tail Han, se encontró un cambio de base de una guanina por una timina en la posición 63 del gen (G63T), donde las ovejas del genotipo AB tuvieron 0.83 corderos más, comparada con las homocigotas AA, lo que prueba que el locus de PRL puede estar asociado con alta prolificidad en ovejas Small Tail Han, pero es necesario confirmar este efecto con otros estudios(34).
En el caso de las cabras Jining Grey, se detectaron polimorfismos en el gen de PRLR asociados a varios genotipos, alguno de las cuales producen cambios en aminoácidos, pero sin efecto sobre tamaño de camada. El cambio de una adenina por una guanina en el aminoácido 143 (A143G) de una de ellas, resultó en un cambio de aminoácido (metionina por valina), donde las cabras con genotipo FG tuvieron 0.76 más cabritos, comparada con las FF, lo cual sugiere que el gene PRLR también tiene un efecto mayor sobre la prolificidad en cabras(72).
Gen del receptor de melatonina A1 (MTNR1A)
La variación estacional en la fertilidad es un importante factor limitante de la eficiencia productiva en ovejas. Los patrones nocturnos de secreción de melatonina, una señal endocrina importante que controla la estacionalidad reproductiva en ovejas(73), es mediada por el MTNR1A involucrado en la regulación de la actividad reproductiva(74).
Se han registrado diferencias importantes entre razas respecto al tiempo y duración del anestro estacional(75). En la raza Merino y sus antecesores (Rambouillet y Dorset) muestran una relativa estacionalidad larga(76) y responden particularmente a la introducción del carnero (efecto macho) para estimular la ovulación y el estro(77). Razas europeas prolíficas tales como la Finish Landrace y Romanov muestran menor estacionalidad con buen comportamiento en verano, en contraste con razas Británicas como la Down, que muestran época corta de empadre y menor respuesta al efecto macho(75).
]]> En México, Arroyo(77) ha documentado extensamente la ausencia o baja estacionalidad de las ovejas de pelo principalmente Pelibuey, característica que seguramente está asociada a factores genéticos, pero que hasta el momento no se han estudiado a nivel molecular, para confirmar si los genes involucrados y sus frecuencias corresponden a las encontradas por otros autores(78,79) y en otras latitudes(80).Se han hecho esfuerzos para la selección de líneas con baja estacionalidad reproductiva, generalmente basadas en la fertilidad en épocas de empadre en primavera y verano. Sin embargo, bajo los sistemas de mejoramiento genético convencionales, es difícil debido a aspectos relacionados con la heredabilidad, tiempo, sistema de manejo y limitación del sexo(25,75).
A fin de estudiar las ventajas comparativas que pueden ofrecer en rebaños con la presencia de genotipos asociados a la no estacionalidad, el Tecnológico de Virginia trabajó con líneas seleccionadas fuera de la estación de empadre (mayo y junio) y establecidas con cruzas 50 % Dorset, 25 % Rambuillet y 25 % Finish Landrace por una generación de apareamientos ínter se, encontrando resultados interesantes. Los valores de crías estimados (EBV) por año para fertilidad fueron de 1.98 vs 0.61 % año-1 de la línea seleccionada sobre la control (E). Hubo un claro efecto de la edad de las ovejas, resultando en una mayor respuesta de las ovejas adultas sobre las ovejas jóvenes(75). Las ovejas en la línea seleccionada mostraron tener menores niveles nocturnos de melatonina circulante y altos niveles de PRL circulante comparadas con las ovejas testigo(81,82). El incremento en los EBV en fertilidad estuvieron asociados con declinación en la melatonina circulante (-2.23 pg ml-1 e incremento en la PRL circulante (1.23 pg ml-1%-1).
Por lo anterior, el conocimiento de los genes involucrados en el control de la estacionalidad reproductiva ligada a marcadores genéticos, permite implementar programas de selección intensiva y eficiente fuera de épocas reproductivas(25) e incluir esta ventaja en rebaños comerciales.
La caracterización molecular de variantes de secuencias del MTNR1A ovino encontradas por Messer et al(82) y Barret et al(83), consiste en una variante con ocho cambios de bases, tres de las cuales resultaron en substituciones de aminoácidos en el receptor. Posteriormente se confirmó que ambas formas del gen estaban presentes en ovejas Greyface x Suffolk, Greyface x Dorset y carneros Soay.
Notter et al(84) estimaron la frecuencia alélica en 0.42 y 0.58 para polimorfismo usando la enzima de restricción Mlnl, y de 0.34 y 0.66 para polimorfismo con la enzima de restricción Rsal en ovejas de las líneas no estacionales del Tecnológico de Virginia (Cuadro 4), demostrando que los polimorfismos no eran independientes de aquel rebaño. Los resultados referidos revelan una diversidad alélica sustancial en el sitio de restricción Mlnl en MTNR1A en varias razas seleccionadas para mínima o no estacionalidad(75).
Por otra parte, Migaud et al(85) identificaron seis polimorfismos dentro del gen MTNR1A en cabras estacionales de la raza Alpina en Francia, y en cabras criollas de Guadalupe con mínima estacionalidad. Se identificó un cambio de aminoácido en el receptor, pero no se identificaron diferencias en las frecuencias alélicas entre estas razas.
En un estudio reciente realizado por Mateescu et al(25) en las que incluyeron ovejas ¾ Dorset x ¼ East Frisian provenientes de un sistema de empadre acelerado (STAR), evaluaron el gen MTNR1A, amplificándolo y digiriéndolo con la endonucleasa Mlnl, lo que permitió identificar a dos alelos (M y m). En forma similar, se evaluó la presencia de corte con la enzima Rsal (alelos R y r). La frecuencia génica fue de 0.64 y 0.36 para el polimorfismo con la enzima Mlnl (alelos M y m, respectivamente). La frecuencia genotípica fueron de 0.43, 0.44, y 0.13 para los genotipos MM, Mm y mm en el polimorfismo identificado con la enzima Mlnl, y 0.13, 0.43 y 0.44 para los genotipos RR, Rr y rr en el polimorfismo identificado con la enzima Rsal.
Los individuos con los genotipos mmRR, MmRr y MMrr tuvieron mayores frecuencias que las esperadas bajo equilibrio Hardy-Weimberg (H-w), por lo que la población considerada no se encontró en equilibrio H-W considerando ambos polimorfismos. Otro aspecto derivado de este estudio fue la relación entre días abiertos y fertilidad mostrada por las ovejas con genotipos Mm o MM, las cuales necesitaron 136 días menos para el primer parto y 124 días menos entre el primero y segundo parto en comparación con las de genotipo mm. Sin embargo, esa diferencia no se observó entre ovejas bajo los genotipos determinados con la enzima Rsal. La conclusión que se deriva de las observaciones mencionadas son que las ovejas portadoras del alelo M conciben a edades más tempranas, y muestran intervalos más cortos entre el primero y segundo parto.
Por otra parte, las ovejas adultas no estacionales del Tecnológico de Virginia, mostraron una fertilidad media entre genotipos de 65.5 a 85.3 %. Las ovejas con al menos una copia del alelo (+) en el sitio de restricción con Mlnl fueron 11.2 % más fértiles en primavera que las ovejas homocigotas para el alelo (-). Sin embargo, no se observó efecto del genotipo MTNR1A sobre el tamaño de la camada(25). La descomposición de la variancia genotípica en aditiva y de dominancia en ovejas adultas para el locus MTNR1A fueron de 35.7 y 13.9 %(2), respectivamente. La varianza aditiva para el marcador MTNR1A encontrada para fertilidad fue de 23.8 % y para el efecto de dominancia en este locus fue de 9.3 %. La heredabilidad para fertilidad en primavera fue de 11 % en adultos(25), la cual se considera baja; por lo que existen también factores medioambientales asociados a esta característica.
]]> Chu et al(86) encontraron efecto de marcador MNTR1A en tamaño de camada en ovejas Small Tailed Han en China con actividad ovulatoria no estacional. Estas ovejas también mostraron tener alta frecuencia para la variante FecB. Las ovejas que fueron heterocigotas para la ausencia del sitio de restricción MnlI tuvieron camadas grandes en el segundo parto comparadas con las homocigotas para la presencia del sitio de restricción (3.19 vs 2.25 corderos camada-1), y camadas grandes comparada con las ovejas heterocigotas tanto en primero como en el segundo parto.Se ha demostrado que el gen MTNR1A es un candidato promisorio para ser usado en la selección asistida por marcadores a fin de mejorar la fertilidad de los rebaños fuera de época de empadre. No obstante, el mecanismo mediante el cual el polimorfismo en el gen MTNR1A influye sobre la reproducción fuera de época aún no se ha establecido, ya que de acuerdo con Mateescu et al(25) parece ser que el polimorfismo que da lugar a la diferencia fenotípica no se debe al cambio de aminoácidos. Es posible que el efecto sea debido a secuencias reguladoras u otros genes cercanos ligados al gen MTNR1A, y también es posible que el efecto de la melatonina sea mediado vía otros mensajeros bioquímicos u hormonales.
CONCLUSIONES
La identificación de poblaciones con variantes alélicas que favorecen la prolificidad, tasa ovulatoria y fertilidad en general, está permitiendo usar estas ventajas reproductivas a nivel comercial, así como el empleo de estas ovejas como modelos en el estudio de los procesos biológicos y fisiológicos en el campo de la reproducción. Esto está permitiendo conocer mejor los fenómenos del control ovárico de la foliculogénesis y la tasa de ovulación. La identificación de los alelos que participan en el aumento de la fertilidad por métodos moleculares permite incrementar de forma dirigida la prolificidad y obtener mayor número de corderos al destete, al mismo tiempo que permite planear los cruzamientos de una forma más racional para disminuir los casos de infertilidad en ovejas homocigotas portadoras.
El estudio y desarrollo de poblaciones o líneas con estacionalidad reducida es claramente posible a través de la selección. La identificación de QTL y marcadores moleculares puede incrementar sustancialmente la respuesta a la selección.
El conocimiento de la presencia de alelos con efecto en el aumento de la fertilidad en diferentes razas, abre la posibilidad de poder establecer poblaciones para el estudio de las interacciones entre los diferentes locus, así como reservorio para la introgresión de estos alelos a poblaciones comerciales.
El empleo de herramientas moleculares para la identificación de polimorfismos en genes candidatos, es una estrategia promisoria para identificar las bases genéticas que pueden estar asociados a los niveles de la tasa ovulatoria, prolificidad y estacionalidad, en las razas de pelo que son empleadas de forma sobresaliente en condiciones tropicales, y que representan un segmento económico importante en diferentes países ecuatoriales.
AGRADECIMIENTOS
]]> Esta revisión es fruto de los apoyos al primer autor bajo el marco de Estancias Posdoctorales Vinculadas al Fortalecimiento de los Posgrados Calidad del CONACYT (96724), y de la Secretaria de Educación Pública a través del Programa de Mejoramiento del Profesorado (PROMEP).
LITERATURA CITADA
1. FAO. The state of the world animal genetic resources for food and agriculture. Edited by Barbara Rischkowsky and Daffyd Pilling. Roma. 2007; 524 p. Disponible en: http://www.fao.org/docrep/010/a1250e/a 1250e00.htm. [ Links ]
2. Galloway SM, Mcnatty KP, Cambridge LM, Laitinen MP, Jennifer JL, Jokiranta S, Mclaren RJ, et al. Mutations in an oocyte-derived growth factor gene (BMP15) cause increased ovulation rate and infertility in a dosage-sensitive manner. Nat Genet 2000;25:279-283. [ Links ]
3. Montgomery GW, Galloway SM, Davis GH, Mcnatty KP. Genes controlling ovulation rate in sheep. Reproduction 2001;121:843-852. [ Links ]
4. Knight PG, Glister C. Local roles of TGF-a superfamily members in the control of ovarian follicular development. Anim Reprod Sci 2003;78:165-183. [ Links ]
5. Shimasaki S, Moore RK, Otsuka F, Erickson GF. The bone morphogenetic protein system in mammalian reproduction. Endocr Rev 2004;25:72-101. [ Links ]
6. McNatty KP, Smith P, Moore LG, Reader K, Lun S, Hanrahan JP, et al. Oocyte-expressed genes affecting ovulation rate. Mol Cell Endocrinol 2005;234(1-2):57-66. [ Links ]
7. Fabre S, Pierre A, Mulsant P, Bodin L, Pasquale ED, Persani L, Monget P, Monniaux D. Regulation of ovulation rate in mammals: contribution of sheep genetic models. Reprod Biol Endocr 2006;4:20. [ Links ]
8. Guo X, Wang XF. Signaling cross-talk between TGF-[beta]/ BMP and other pathways. Cell Res 2009;19:71-88. [ Links ]
9. Xu E, Li E, Han Y, Chen L, Xie Z. Differential expression of mRNAs encoding BMP/Smad pathway molecules in antral follicles of high-and low-fecundity Hu sheep. Anim Reprod Sci 2010;120:47-55. [ Links ]
10. Hanrahan PJ, Gregan SM, Mulsant P, Mullen M, Davis GH, Powell R, Galloway SM. Mutations in the genes for oocyte-derived growth factors GDF9 and BMP15 are associated with both increased ovulation rate and sterility in Cambridge and Belclare sheep (Ovis aries). Biol Reprod 2004;70:900-909. [ Links ]
11. Silva BDM, Castro EA, Souza CJH, Paiva SR, Sartori R, Franco MM, Azevedo HC, et al. A new polymorphism in the Growth and Differentiation Factor 9 (GDF9) gene is associated with increased ovulation rate and prolificacy in homozygous sheep. Anim Genet 2011;42(1):89-92. [ Links ]
12. Bodin L, San Cristobal M, Lecerf F, Mulsant P, Bibe B, Lajous D, Belloc JP, Eychenne F, Amigues Y, Elsen JM. Segregation of a major gene influencing ovulation in progeny of Lacaune meat sheep. Genet Selec Evol 2002;34:447-464. [ Links ]
13. Monteagudo LV, Ponz R, Tejedor MT, Lavina A, Sierra I. A 17 bp deletion in the Bone Morphogenetic Protein 15 (BMP15) gene is associated to increased prolificacy in the Rasa Aragonesa sheep breed. Anim Reprod Sci 2009;110:139-146. [ Links ]
14. Wilson T, Wu XY, Juengel JL, Ross IK, Lumsden JM, Lord EA, Dodds KG, Walling GA, Mcewan JC, O'Connell AR, McNatty KP, Montgomery GW. Highly prolific Booroola sheep have a mutation in the intracellular kinase domain of bone morphogenetic protein IB receptor (ALK-6) that is expressed in both oocytes and granulosa cells. Biol Reprod 2001;64:1225-1235. [ Links ]
15. Mulsant P, Lecerf F, Fabre S, Schilber L, Monget P, Lanneluc I, Pisselet C, et al. Mutation in bone morphogenetic protein receptor-IB is associates with increased ovulation rate in Booroola Merino ewes. Proc Natl Acad Sci 2001;98:5104-5109. [ Links ]
16. Souza CJ, Macdugall C, Campbell BK, Mcneilly AS, Baird DT. The Booroola (FecB) phenotype is associated with a mutation in the bone morphogenetic receptor type 1 B (BMPR1B) gene. J Endocrinol 2001;169:R1-R6. [ Links ]
17. Davis GH, Galloway SM, Ross IK, Gregan SM, Ward J, Nimbkar BV, Ghalsasi PM, et al. DNA tests in prolific sheep from eight countries provide new evidence on origin of the Booroola (FecB) Mutation. Biol Reprod 2002;66:1869-1874. [ Links ]
18. Davis GH, Balkrishnan L, Ross IK, Wilson T, Galloway SM, Lumsden BM, Hanrahan JP, et al. Investigation of the Booroola (Fec B) and Inverdale (Fec X I) mutation in 21 prolific breeds and strains of sheep samples in 13 countries. Anim Reprod Sci 2006;92:87-96. [ Links ]
19. Montgomery GW. Genome mapping in ruminants and map locations for genes influencing reproduction. Rev Reprod 2000;5(1):25-37. [ Links ]
20. Drouilhet L, Lecerf F, Bodin L, Fabre S, Mulsant P. Fine mapping of the FecL locus influencing prolificacy in Lacaune sheep. Anim Genet 2009;40(6):804-12. [ Links ]
21. Hua GH, Chen Sl, Yao HG, Wu WS, Shen Z, Chen QK, Chen L, et al. RFLP of INHA and its relationship to goat litter size. Yi Chuan 2007;29(8):972-6. [ Links ]
22. He Y, Ma X, Liu X, Zhang C, Li J. Candidate genes polymorphism and its association to prolificacy in Chinese goats. J Agric Sci 2010;2(1):88-92. [ Links ]
23. Chu MX, Zhuang H, Zhang Y, Jin M, Li X, Di R, Cao G, et al. Polymorphism of inhibin äB gene and its relationship with litter size in sheep. J Anim Sci 2011;82:57-61.61 [ Links ]
24. Notter DR. Genetic aspect of reproduction in sheep. Reprod Dom Anim 2008;43 (Suppl 2):122-128. [ Links ]
]]>25. Mateescu RG, Lunsford AK, Thonney ML. Association between melatonin receptor 1A gene polymorphism and reproductive performance in Dorset ewes. J Anim Sci 2009;87:2485-2488. [ Links ]
26. Palmer JS, Zhao ZZ, Hoekstra C, Hayward NK, Webb PM, Whiteman DC, Martin NG, et al. Novel variants in growth differentiation factor 9 in mothers of dizygotic twins. J Clin Endocr Metab 2006;91:4713-4716. [ Links ]
27. Davis GH, Mcewan JC, Fennessy PF, Dodds KG, McNatty KP. Infertility due to bilateral ovarian hypoplasia in sheep homozygous (FecXI FecXI) for the Inverdale prolificacy gene located on the X chromosome. Biol Reprod 1992;46:636-640. [ Links ]
28. McNatty KP, Juengel JL, Reader KL, Lun S, Myllymaa S, Lawrence SB, Western A, et al. Bone morphogenetic protein 15 and growth differentiation factor 9 cooperate to regulate granulosa cell function. Reproduction 2005;129:473-480. [ Links ]
29. Juengel Jl, Reader Kl, Bibby AH, Lun S, Ross I, Haydon LJ, McNatty KP. The role of bone morphogenetic proteins 2, 4, 6 and 7 during ovarian follicular development in sheep: contrast to rat. Reproduction 2006;131(3):501-513. [ Links ]
]]>30. Yoshino O, Mcmahon HE, Sharma S, Shimasaki S. A unique preovulatory expression pattern plays a key role in the physiological functions of BMP-15 in the mouse. Proc Natl Acad Sci 2006;103(28):10678-10683. [ Links ]
31. Shimasaki S, Zachow RJ, Li D, Kim H, Iemura SI, Ueno N, Sampath K, Chang RJ, Erickson GF. A functional bone morphogenetic protein system in the ovary. Proc Natl Acad Sci 1999;96:7282-7287. [ Links ]
32. Pangas SA, Li X, Umans L, Zwijsen A, Huylebroeck D, Gutierrez C, Wang D, Martin JF, et al. Conditional deletion of Smadl and Smad5 in somatic cells of male and female gonads leads to metastatic tumor development in mice. Mol Cell Biol 2008;28(1):248-257. [ Links ]
33. Davis GH. Major genes affecting ovulation rate in sheep. Genet Sel Evol 2005;37:S11-S23. [ Links ]
34. Chu MX, Mu YL, Fang L, Ye SC, Sun SH. Prolactin receptor as a candidate gene for prolificacy of Small Tail Han sheep. Animal Biotechnol 2007;18(1):65-73. [ Links ]
]]>35. Galloway SM, Gregan SM, Wilson T, McNatty KP, Juengel JL, Ritvos O, Davis GH. BMP15 mutations and ovarian function. Mol Cell Endocrinol 2002;191:15-18. [ Links ]
36. Barzegari A, Atashpaz S, Ghabili K, Nemati Z, Rustaei M, Azarbaijani R. Polymorphisms in GDF9 and BMP15 associated with fertility and ovulation rate in Moghani and Ghezel sheep in Iran. Reprod Dom Anim 2010;45:666-669. [ Links ]
37. Bodin L, Di Pasquale E, Fabre S, Bontoux M, Monget P, Ersani L, Mulsant P. A novel mutation in the bone morphogenetic protein 15 gene causing defective protein secretion is associated with both increased ovulation rate and sterility in Lacaune sheep. Endocrinol 2007;148:393-400. [ Links ]
38. Martínez-Royo A, Jurado JJ, Smulders JP, Martí JI, Alabart JL, Roche A, Fantova E, et al. A deletion in the bone morphogenetic protein 15 gene causes sterility and increased prolificacy in Rasa Aragonesa sheep. Anim Genet 2008;39(3):294-297. [ Links ]
39. Martínez-Royo A, Dervishi E, Alabart JL, Jurado JJ, Folch J, Calvo JH. Freemartinism and FecXR allele determination in replacement ewes of the Rasa Aragonesa sheep breed by duplex PCR. Theriogenology 2009;72(8):1148-52. [ Links ]
]]>40. Hashimoto O, Moore RK, Shimasaki S. Posttranslational processing of mouse and human BMP-15: potential implication in the determination of ovulation quota. Proc Natl Acad Sci USA 2005;102(15):5426-5431. [ Links ]
41. Davis GH, Montgomery GW, Allison AJ, Kelly RW, Bray AR. Segregation of a major gene influencing fecundity in progeny of Booroola sheep. NZ J Agric Res 1982;25:525-529. [ Links ]
42. Lahoz B, Alabart JL, Jurado JJ, Calvo JH, Martínez-Royo A, Fantova E, Folch J. Effect of the FecX(R) polymorphism in the bone morphogenetic protein 15 gene on natural or equine chorionic gonadotropin-induced ovulation rate and litter size in Rasa Aragonesa ewes and implications for on-farm application. J Anim Sci 2011;89(11):3522-30. [ Links ]
43. Demmers KJ, Smaill B, Davis GH, Dodds KG, Juengel JL. Heterozygous Inverdale ewes show increased ovulation rate sensitivity to pre-mating nutrition. Reprod Fertil Dev 2011;23(7):866-75. [ Links ]
44. Javanmard A, Azadzadeh N, Esmailizadeh AK. Mutations in bone morphogenetic protein 15 and growth differentiation factor 9 genes are associated with increased litter size in fat-tailed sheep breeds. Vet Res Commun 2011;35(3):157-67. [ Links ]
]]>45. Roy J, Polley S, De S, Mukherjee A, Batabyal S, Pan S, Brahma B, Datta TK, et al. Polymorphism of fecundity genes (FecB, FecX, and FecG) in the Indian Bonpala sheep. Anim Biotechnol 2011;22(3):151-62. [ Links ]
46. Shabir M, Ganai TA. Nucleotide sequencing and DNA polymorphism studies of BMP 15 gene in Corriedale and local Kashmir valley sheep (Ovis aries). Gene 2012; 499(1):231-235. [ Links ]
47. Polley S, De S, Brahma B, Mukherje A, Vinesh PV, Batabyal S, Arora JS, Pan S, Samanta AK, Datta TK, Goswami SL. Polymorphism of BMPR1B, BMP15 and GDF9 fecundity genes in prolific Garole sheep. Trop Anim Health Prod 2010;42:985-993. [ Links ]
49. Fatehi AN, Van Den Hurk R, Colenbrander B, Daemen AJ, Van Tol HT, Monteiro RM, Roelen BA, Bevers MM. Expression of bone morphogenetic protein2 (BMP2), BMP4 and BMP receptors in the bovine ovary but absence of effects of BMP2 and BMP4 during IVM on bovine oocyte nuclear maturation and subsequent embryo development. Theriogenology 2005;63(3):872-89. [ Links ]
50. Edwards SJ, Reader KL, Lun S, Western A, Lawrence S, McNatty KP, Juengel JL. The cooperative effect of growth and differentiation factor-9 and bone morphogenetic protein (BMP)-15 on granulose cell function is modulated primarily through BMP receptor II. Endocrinol 2008;149(3):1026-1030. [ Links ]
]]>51. Zhu G, Guo B, Pan D, Mu Y, Feng S. Expression of bone morphogenetic proteins and receptors in porcine cumulus-oocyte complexes during in vitro maturation. Anim Reprod Sci 2008;104:275-283. [ Links ]
52. Mazerbourg S, Klein C, Roh J, Kaivo-Oja N, Mottershead DG, Korchynskyi O, Ritvos O, Hsueh AJ. Growth differentiation factor-9 signaling is mediated by the type I receptor, activin receptor-like kinase 5. Mol Endocrinol 2004;18(3):653-65. [ Links ]
53. Drouilhet L, Taragnat C, Fontaine J, Duittoz A, Mulsant P, Bodin L, Fabre S. Endocrine characterization of the reproductive axis in highly prolific Lacaune sheep homozygous for the FecLL mutation. Biol Reprod 2010;82:815-824. [ Links ]
54. Drouilhet L, Mansanet C, Sarry J, Tabet K, Bardou P, et al. The Highly Prolific Phenotype of Lacaune Sheep Is Associated with an Ectopic .Expression of the B4GALNT2 Gene within the Ovary. PLoS Genet 2013;9(9):e1003809. doi:10.1371/journal.pgen.1003809. [ Links ]
55. Rothschild M, Jacobson C, Vaske D, Tuggle C, Wang L, Short T, Eckardt G, et al. The estrogen receptor locus is associated with a major gene influencing litter size in pigs. Proc Natl Academy Sci. USA 1996;93:201-205. [ Links ]
]]>56. Hewitt SC, Korach KS. Oestrogen receptor knockout mice: roles for oestrogen receptors alpha and beta in reproductive tissues. Reproduction 2003;125(2):143-149. [ Links ]
57. Shi H, Bai J, Niu Z, Muniresha, Fen L. Jia B. Study on candidate gene for fecundity traits in Xingjiang Cele black sheep. African J Biotechnol 2010;9(49):8498-8505. [ Links ]
58. Bi XD, Chu MX, Jin HG, Fang L, Ye SC. Estrogen receptor as a candidate gene for prolificacy of small tail Han sheep. Yi Chuan Xue Bao 2005;32(10):1060-1065. [ Links ]
59. Di R, Jia LH, Chu MX, Chen HQ, Fang L. Polymorphism Analysis of Exon 4 of Estrogen Receptor Gene in Sheep. China Anim Husb Vet Med 2008;12. [on line]: http://en.cnki.com.cn/Article_en/CJFDTOTAL-GWXK200812024.htm [ Links ]
60. Robertson DM, Foulds LM, Leversha L, Morgan FJ, Hearn MT, Burger HG, Wettenhall RE, et al. Isolation of inhibin from bovine follicular fluid. Biochem and Bioph Res Comm 1985;126:220-226. [ Links ]
61. Woodruff TK, Besecke LM, Groome N, Draper LB, Schwartz NB, Weiss J. Inhibin A and inhibin B are inversely correlated to follicle-stimulating hormone, yet are discordant during the follicular phase of the rat estrous cycle, and inhibin A is expressed in a sexually dimorphic manner. Endocrinol 1996;137:5463-5467. [ Links ]
62. Mason AJ, Hayflick JS, Ling N, Esch F, Ueno N, Ying SY, Guillemin R, et al. Complementary DNA sequences of ovarian follicular fluid inhibin show precursor structure and homology with transforming growth factor-beta. Nature 1985;318(6047):659-63. [ Links ]
63. Jaeger C, Hiendleder S. Cosmid cloning and characterization of the coding regions and regulatory elements of the ovine a-(INHA), âA-(INHBA) and äB-inhibin (INHBB) genes. Anim Genet 1994;25(Suppl 2):33. [ Links ]
64. Hiendleder S, Lewalski H, Jaeger C, Plante Y. Genomic cloning and comparative sequence analysis of different alleles of the ovine âA-inhibin/activin (INHBA) genes as a potential QTL for litter size. Anim Genet 1996;27(Suppl 2):119. [ Links ]
66. Hiendleder S, Lewalshi H, Jaeger C, Pracht P. Nucleotide sequence of ovine ä-inhibin (INHA) genes and evaluation of RFLP marker effects on reproductive performance. Anim Genet 2002;33(3)247-248. [ Links ]
66. Leyhe B, Hiendleder S, Jaeger C, Wassmuth R. Pronounced differences in the frequency Taq I äA inhibin alleles between sheep breeds with different reproductive performance. Anim Genet 1994;25(1):41-43. [ Links ]
67. Vincent AL, Wang L, Tuggle CK, Robic A, Rothschild MF. Prolactin receptor maps to pig chromosome 16. Mammal Genom 1997;8(10):793-794. [ Links ]
68. Jenkins ZA, Henry HM, Sise JA, Montgomery GW. Follistatin (FST), growth hormone receptor (GHR), and prolactin receptor (PRLR) genes map to the same region of sheep chromosome 16 [abstract]. Anim Genet 2000;31(4):280. [ Links ]
69. Chu MX, Wang XC, Jin M, Di R, Chen HQ, Zhu GQ, Fang L, Ma YH, Li K. DNA polymorphism of 5' flanking region of prolactin gene and its association with litter size in sheep. J Anim Breed Genet 2009;126(1):63-68. [ Links ]
70. Kelly PA, Djiane J, Postel-Vinay MC, Edery M. The prolactin/ growth hormone receptor family. Endocr Rev 1991;12(3):235-251. [ Links ]
71. Zhang GX, Chu MX, Wang JY, Fang L, Ye SC. Polymorphism of exon 10 of prolactin receptor gene and its relationship with prolificacy of Jining Grey goats. Yi Chuan 2007;29(3):329-36. [ Links ]
72. Bittman EL, Dempsey RJ, Karsch FJ. Pineal melatonin secretion drives the reproductive response to daylength in the ewe. Endocrinol 1983;113:2276-2283. [ Links ]
73. Weaver DR, Liu C, Reppert SM. Nature's knockout: The Mel1b receptor is not necessary for reproductive and circadian responses to melatonin in Siberian hamsters. Mol Endocrinol 1996;10:1478-1487. [ Links ]
74. Notter DR, Cockett NE. Opportunities for detection and use of QTL influencing seasonal reproduction in sheep: a review. Genet Sel Evol 2005;37(Suppl 1):S39-S53. [ Links ]
75. Notter DR. Genetic improvement of out-of-season breeding through selection, in: Proc Out-of-Season Breeding Symp. Ames, Iowa 1992:55-81. [ Links ]
76. Pearce DT, Oldham CM. The ram effect, its mechanism and application to the management of sheep, in: Lindsay DR, Pearce DT editors. Reproduction in sheep. Aust Academy Sci. Canberra 1984:26-34. [ Links ]
77. Arroyo J. Reproductive seasonality of sheep in Mexico. Trop Subtrop Agros 2011;14:829-845. [ Links ]
78. Hinojosa-Cuellar JA, Oliva-Hernández J. Distribución de partos por estación en ovejas de razas de pelo y cruces en un ambiente tropical húmedo. Rev Cient FCV-Luz 2009;XIX:288-294. [ Links ]
79. Luna PC, Berumen AAC, Aguilar CJA, Cansino AGR. Fertility of hair ewes supplemented with African palm kernel. Livest Res Rural Develop 2010; [on line]: http://www.lrrd.org/lrrd22/10/luna22178.htm [ Links ]
80. Wildeus S. Hair sheep genetic resources and their contribution to diversified small ruminant production in the United States. J Anim Sci 1997;75:630-640. [ Links ]
81. Notter DR, Chemineau P. Nocturnal melatonin and prolactin plasma concentrations in sheep selected for fertility in autumn lambing. J Anim Sci 2001;79:2895-2901. [ Links ]
]]>82. Messer LA, Wang L, Tuggle CK, Yerle M, Chardon P, Pomp D, Womack JE, Barendse W, Crawford AM, Notter DR, Rothschild MF. Mapping of the melatonin receptor 1a (MTNR1A) gene in pigs, sheep, and cattle. Mamm Genome 1997;8:368-370. [ Links ]
83. Barrett P, Conway S, Jockers R, Strosberg AD, Guardiola-Lemaitre B, Delagrange P, Morgan PJ. Cloning and functional analysis of a polymorphic variant of the ovine Mel 1a melatonin receptor. Biochem Biophys Acta 1997;1356:299-307. [ Links ]
84. Notter DR, Cockett NE, Hadfield TS. Evaluation of melatonin receptor 1a as a candidate gene influencing reproduction in a fall-lambing sheep flock. J Anim Sci 2003;81:912-917. [ Links ]
85. Migaud M, Gavet S, Pelletier J. Partial cloning and polymorphism of the melatonin1a (Mel1a) receptor gene in two breeds of goat with different reproductive seasonality. Reprod 2002;124:59-64. [ Links ]
86. Chu MX, Ji CL, Chen GH. Association between PCR-RFLP of melatonin receptor 1a gene and high prolificacy in Small Tail Han sheep, Asian-Aust J Anim Sci 2003;16:1701-1704. [ Links ]
]]>