
Isolation, molecular and fermentative characterization of a yeast used in ethanol production during mezcal elaboration
Aislamiento, caracterización molecular y fermentativa de una levadura usada en la producción de etanol durante la elaboración de mezcal
J.C. González-Hernández1*, E. Pérez1, R.M. Damián1 and M.C. Chávez-Parga2
1 Laboratorio de Bioquímica del Departamento de Ing. Bioquímica del Instituto Tecnológico de Morelia, Av. Tecnológico # 1500, Colonia Lomas de Santiaguito, C. P. 58120, Morelia, Michoacán, México. * Corresponding author. E-mail: jcgh1974@yahoo.com Tel. (+52-433) 3121570. Ext. 1240., Fax (+52-433) 3121570. Ext. 211.
]]> 2 Facultad de Ingeniería Química de la Universidad Michoacana de San Nicolás de Hidalgo. Morelia, Michoacán, México.
Received 6 of July 2012
Accepted 11 of September 2012
Abstract
Numerous ecological itudies have been conducted over the years to understand the dynamics, quantification, and composition of the microflora responsible for spontaneous fermentation. Among them, yeasts are microorganisms that exert key processes and are responsible for alcoholic fermentation. In the present study, we isolated and characterized molecularly two yeasts by RFLP (Restriction Fragment Length Polymorphisms) in which we compared the restriction patterns of different genomic regions corresponding to the ribosomal DNA. Experimental design (ED) was based on the Response Surface Methodology (MSR) used to design a suitable culture medium for the produ ction of Mezcal usmg as substrate an Agave cupreata extract juice and a LEVM yeast strain. We found by RFLP iwo Saccharomeces cerevisiae yeasts etrains (LEVM y LEVZ). in ED tor the LEVM the variables selected cuch as the pH, initial substrate concentration, and temperature were operating levels as close as possible to the original process, these preliminary results show the importance of using molecular techniques for the characterization of yeast strains used in the beverage indus try and the use of ED allowed establish the fermentation process conditions.
Keywords: yeast, RFLP, fermentation, experimental design.
Resumen
]]> Existen estudios dirigidos hacia la composición y cuantificación de la microflora responsable de las fermentaciones espontáneas. Dentro de ellos se ha encontrado que las levaduras son microorganismos clave durante la fermentación alcohólica. En el presente estudio se aislaron y caracterizaron molecularmente dos levaduras poa RFLP (Resfriction Fragment: Length Polymorphisms) en las cuales se comparan los patrones de restricción de diferentes regiones genómicas correspondientes al DNA ribosomal. Se realizó un diseño de experimentos (DE) basado en la metodología de superficie de respuesta (MSR) para diseñar un medio de cultivo en la producción de Mezcal, usando como sustrato jugo de Agave cupreata y la levadura LEVM. Encontrándose por RFLP dos levaduras Saccharomyces cerevisiae (LEVM y LEVZ). En el DE para la LEVM las variables analizadas fueron el pH, concentracion de sustrato y temperatura, para evaluar las variables y los niveles de operación lo mas cercanos posible al proceso original. Encontrando que la temperatura es la variable que tiene un efecto significativo sobre la producción de etanol., estos resultados preliminares muestran la importancia de usar técnicas moleculares para la caracterización de levaduras en la industria de las bebidas y el uso de DE permitio establecer las condiciones para llevar a cabo el proceso de fermentación.Palabras clave: levadura, RFLP, fermentación, diseño experimental.
1 Introduction
Mezcal is a traditional alcoholic beverage of México, which is made similarly to Tequila. The process begins with the harvesting of the agave after 8 years of cultivation; at this stage the plants are cut off from their base and most of their leaves are removed, obtaining the core of the plant called agave pineapple, which is cooked in ovens or autoclaves. At this stage, the polysaccharides, mainly from residues (fructans) are thermally hydrolyzed to fructose syrup, which then undergoes alcoholic fermentation with native yeasts. Finally a must with an approximate ethanol content of 3-6% v/v is distilled to obtain white or young Mezcal (Cedeno, 1995).
Tequila is only produced from Agave tequilana species (NOM-006-SCFI-1994), whereas in the case of Mezcal, the Mexican Official Standard NOM-070-SCFI-1994 indicates that a wide variety of Agaves can be used in its drafting, the most used are Agave angustiofolia, Agave esperrima, Agave potatorum, and Agave salmiana. Another important difference between Mezcal and Tequila is the elaboration process; in the first, a traditional process is usual, whereas for tequila a high tech process is applied. It is also important to note that the geographical areas that have a denomination of origin for Mezcal are more dispersed in the Mexican territory, whereas for tequila it is restricted to a smaller region, which adds a factor of variability to the production of Mezcal in each region (Molina et al., 2007).
Currently, the process used to produce Mezcal in most municipalities in the state of Michoacan is carried out by craftsmen in open containers. Among the problems facing the Mezcal Agave-chain, it can be mentioned that production is considered a seasonal activity that only takes place during the months of October through May, at the end of the rainy season. Most "Vinatas" (vineyard) are located near streams or rivers, some of them at the bottom of deep ravines. Besides, there is an overexploitation of wild populations of the agave for Mezcal in the different regions, and the marketing of the product is at small-scale and limited to the local level. There is no control in the process of preparing the drink to the detriment of its validity, despite the existence of standards, which are unknown to most producers (Gallardo et al., 2008). Particular strains of yeast are not used for fermentation, this is accomplished only with the yeast in the environment, and because it is insufficient, the process has low yields and a longer fermentation time, no care is taken for sterility in the production area, which can affect the characteristics of the final product, the fermentation scheme is inadequate, and there is no equipment developed for the Mezcal industry in the state of Michoacan to enable standardization of its products.
As a consequence of this set of problems, it is necessary to ensure the completion of alcoholic fermentation, as well as to attain a typical Mezcal that may be reproducible. The best strategy seems to be the inoculation with an indigenous strain that is better suited and can maintain the typical features of the area and to provide producers of Michoacan Mezcal with scientific and technical tools that will allow them to make a product that meets the specifications of the standards governing this drink without stifling its natural qualities and losing its authenticity and craftsmanship, which make the drink a unique product of superior quality 100% Mexican.
The yeasts responsible for fermentation can come from either the Agave (the main raw material for Mezcal production) or the environment of the Vinata or distilleries. Spontaneous fermentations are those that occur naturally, i.e., made from the Agave yeasts and material of the Vinata, without any external inoculation. Spontaneous fermentations are not products of the action of a single species or strain of yeast, but result from a succession of species and different yeast strains during fermentation (Kunkee and Amerine, 1970; Ribereau-Gayon et al., 1975, Lafon-Lafourcade, 1983; Zambonelli, 1988), all contributing to the transformation of sugars into ethanol, glycerol, organic acids, and volatile compounds, which have a direct influence on the flavor and aroma of distillates.
There have been few studies on the microbial ecology of fermented beverages and distilled products from Agave. Natural fermentation of Tequila and Mezcal from Agave include non-Saccharomyces and Saccharomyces cerevisiae strains, which are the main producers of ethanol. In the tequila industry, a common practice is the use of pure cultures of S. Cerevisiae as the initial inoculum. However, a growing number of non-Saccharomyces yeasts have been systematically investigated for their ability to improve the sensory characteristics and optimize the typical attributes of local fermented products, making it necessary to characterize the native yeasts to use them with S. cerevisiae as a mixed inoculum (Jacques-Hernandez et al., 2009).
]]> Although spontaneous fermentation occurs from a succession of genera and species of yeast, only a few strains of S. cerevisiae control most fermentation. This is the result of natural selection during spontaneous fermentation (Frezier and Dubourdieu, 1992; Vezinhet et al., 1992, Fleet and Heard, 1993; Versavaud et al., 1993).Traditionally, the methods used for the identification and characterization of yeast species and strains have been based on morphological and sexual characteristics, but these features are heavily influenced by culture conditions and can give inaccurate results (Kreger-Van Rij, 1984).
For the selection of strains, it is essential to establish their oenological properties. There are different criteria that can be divided into: favorable (ethanol tolerance, good performance in the transformation of sugars into ethanol, ability to grow at high sugar concentrations, etc.) and unfavorable (production of H2S, foaming or volatile acidity). However, there are some aspects that are usually considered favorable properties that can be included in a third group called neutral (Cuinier, 1985; Esteve-Zarzoso et al., 1999).
The RFLP technique allows differentiating various microorganisms by analyzing the band patterns resulting from the breaking of their DNAs. These patterns, known as DNA restriction patterns, are obtained through the activity of restriction endonucleases. The smaller the size of the nucleotide sequence, the greater the number of fragments generated. The fragments can be separated by agarose gel electrophoresis, resulting in characteristic restriction profiles. The profiles depend on the restriction enzyme and the DNA (nDNA or mtDNA) used, although the most used is mtDNA. Comparison of profiles allows differentiating various species from each other or even populations within a species (Salas and Arenas, 2001).
The optimization of culture media for industrial purposes in most cases has been made by empirical procedures of trial and error, not only in developing the culture medium but also regarding operating conditions. In either way it is likely that the original culture medium can be optimized by changing the percentage of medium components and raw materials used, being feasible in many cases to optimize the environmental compounds so that the process is not only more productive but also of the same or less than the original cost, all which requires the use of various optimization methods. One of the most efficient techniques for process optimization is the Response Surface Methodology (MSR), its main objective is to determine the optimum operating conditions for a system, or to determine the region of space where factors are met to satisfy operating conditions. MSR is used successfully in the chemical industry and in recent years has been used in microbiological culture media formulation based on a set of mathematical and statistical techniques, through which we can model and analyze problems in which a response of interest is determined by several variables, with the goal of optimizing the response itself. This methodology is unique in determining the influence and importance of the parameters studied and the interactions between these a minimal of assays. Such designs can be of considerable value when it is important to reduce the number of runs as much as possible (Montgomery, 2004).
Using the Response Surface Methodology (MSR), it is possible to formulate a suitable culture medium to maximize the production of ethanol in the production of Mezcal, using yeasts isolated from spontaneous fermentations of a Mezcal producing region, aimed at designing culture media to optimize variables that maximize ethanol yield in the production of Mezcal, using isolated yeast ferments and as substrate Agave cupreata juice. In addition the MSR may help to characterize fermentatively the yeasts isolated from the Mezcal ferments, establish the ideal conditions for maximum ethanol production and cell growth at a flask level, and identify the volatile compounds present in the final product.
2 Materials and methods
2.1 Source of substrate and yeast isolation
Extract of Agave cupreata previously hydrolyzed. From fermented juice sample, it was realize the yeasts isolation. The plates were incubated at 32°C for 48 h for colony development. The various colony types were counted, and representative colonies of each type were isolated and subcultured in YPD (yeast extract 10 gL-1; peptone 20 g/L-1; dextrose 20 g/L-1; agar 20 g/L-1) for subsequent identification.
]]>2.2 Microorganisms
We used a yeast strain isolated from a spontaneous fermentation of a Mezcal producing region of the state of Michoacan (LEVM), and the producing region of the state of Zacatecas (LEVZ) and S. cerevisiae 288C yeast control.
2.3 Molecular characterization
The molecular characterization was performed by RFLP (Restriction Fragment Length Polymorphisms), which is a comparative analysis of restriction patterns of the different genomic regions corresponding to ribosomal DNA. This technique involves the combined use of RFLP and PCR (Polymerase Chain Reaction). In this way, specific DNA fragments are amplified by PCR and subsequently treated with selected restriction enzymes (Salas and Arenas, 2001). Cells were directly collected from a fresh yeast colony using yellow tips and suspended in 100 μl PCR reaction mix containing 0.5 μM primer ITS1 (5' TCCGTAGGTGAACCTGCGG 3'), 0.5 μM primer ITS4 (5' TCCTCCGCTTATTGATATGC 3'), 10 μM deoxynucleotides, 1.5 mM MgCl2 and 1X buffer (MAD-GEN). The suspension was heated at 95 °C for 15 min in a Progene (Techne) thermocycler. One unit of DNA Polymerase SuperTherm (MAD-GEN) was then added to each tube. PCR conditions were as follows: initial denaturation at 95 °C for 5 min; 35 cycles of denaturing at 94 °C for 1 min, annealing at 55.5 °C for 2 min, and extension at 72 °C for 2 min; and a final extension at 72 °C for 10 min. PCR products (10 μl or approximately 0.5-1.0 μg) were digested without further purification with the restriction endonucleases CfoI, HaeIII and HinfI (Boehringer Mannheim). The PCR products and their restriction fragments were separated on 1.4% and 3% agarose gels, respectively, with 1X TAE buffer. After electrophoresis, gels were stained with ethidium bromide, visualized under UV light and photographed (Image Master, Pharmacia). Sizes were estimated by comparison against a DNA length standard (100 bp ladder, Gibco-BRL) (Esteve-Zarzoso et al., 1999).
2.4 Formulation of the inoculum
For the formulation of the inoculum, 100 ml of Agave juice previously filtered were placed in a 250-ml Erlenmeyer flask; the concentration of sugars were adjusted to 12 º Brix using an ABBE refractometer, with a concentration of 1% (NH4)2HPO4. A potentiometer (Hanna Instruments) was used to adjust the pH to 4.5 (with HCl). The medium was subjected to sterilization, leaving it to cool to room temperature. Through a culture loop two samples of fresh colonies were taken to study the strain present in the Petri dish, which were then inoculated into the flask under sterile conditions. After the inoculation, the flask was placed in the incubator at a temperature of 28 ° C for 48 hours with agitation at 150 rpm.
]]> 2.5 Formulation of culture media of the different treatments
For the preparation of these media, hydrolyzed Agave juice was filtered, the sugar concentration was adjusted by refractometry on the Brix scale according to the classical methodology of the sugar industry, through ABBE refractometer adding distilled water; salts of (NH4)2HPO4 were added at 0.1% concentration. This was carried out under the conditions stated in the experimental design for each flask. Finally, 100 mL of medium were placed in a 250-mL Erlenmeyer flask that was sealed with a cotton plug to prevent contamination; then, it was sterilized for 20 minutes at 15 lbs (121 °C). The medium was allowed to cool at room temperature and was inoculated at a concentration of 3 x 106 celmL-1.
2.6 Determination of cell growth
Cell growth was determined in the samples taken every four hours during fermentation by optical density measurements using a spectrophotometer (UNICO model 1000), the measurement was performed at a wavelength of 540 nm for which 100 μL of the fermented must were placed in 900 μL of distilled water (dilution 1:10), this mixture was then placed in a reading-cell and the optical density was measured.
2.7 Quantification of substrate consumption
One hundred microliter of appropriately diluted sample were placed in a tube (with screw-on cap), adding 100 μL of DNS reagent: after replacing the stopper, the tube was stirred and placed for 5 minutes in a water bath at 95-100 °C. The mixture was cooled in an ice bath and 1 mL of distilled water was added, finally the optical density of the sample was read at 540 nm in a spectrophotometer (UNICO model 1000). To obtain the value in grams per liter (gL-1) of total reducing sugars, a calibration curve with xylose, fructose and glucose was prepared to interpolate data.
2.8 Quantification of ethanol by an enzymatic method
]]> In a plastic covered cell, we placed 2 mL of distilled water, 0.10 ml of the sample, 0.20 mL pyrophosphate buffer solution (pH 9.0), 0.2 ml of NAD+ and 0.02 mL of aldehyde dehydrogenase solution (167 UmL-1); the same amount of reagents, except for the sample were placed in another cell, mixed and the absorption of both cells was read (340 nm) after about 2 minutes (A1). To each cell 0.02 mL of the alcohol dehydrogenase solution was added, and absorbance was measured after about 5 minutes (A2). We proceeded to perform the necessary calculations to obtain the concentration of ethanol in the sample (test procedure K-ETOH 11/05, Megazyme).
2.9 Determination of pH variation
To determine the pH, a sample was taken from the fermentation medium and placed in a 50 mL beaker, and the pH variation during fermentation was assessed with a pHmeter (Hanna Instruments).
2.10 Analysis of the variables for ethanol production at flask level
Variables were established as A: pH (4.5-5.5), B: initial substrate (12-14 ° Brix) and C: temperature (28-32 °C), levels of operation were established based on previous studies. Once selected variables and levels of operation were established, a Box-Behnken design was performed. The flasks with culture medium were inoculated with the pure strain of yeast previously isolated and characterized. In all experimental trials, the initial inoculum concentration and volume of culture medium were kept constant. Both experimental designs and statistical analysis were performed with the software STATGRAPHICS Plus (MR).
2.11 Box-Behnken design
Box-Behnken design is applied for three or more factors and this is often efficient in the number of runs. On the other hand, factorial designs involve two or more factors, each of which has different values or levels, and whose experimental units cover all possible combinations of these levels across all factors. Such experiments allow the study of the effect of each factor on the response variable, and the effect of interactions among factors on this variable (Gutierrez and de la Vara, 2008).
]]>3 Results and discussion
3.1 Molecular characterization
For the studies of cultures in YPD enriched solid medium, the strain was preserved in liquid YPD medium and glycerol at -20°C. The molecular characterization was performed by RFLP technique. Results are shown in Fig. 1. The LEVM (I) and LEVZ (II) (yeasts strains isolated from a spontaneous fermentation of a Mezcal producing region of the state of Michoacan and Zacatecas, respectively) yeasts that had provided an 880 bp amplicon was digested with enzyme Cfol (lane A) which provided three restriction patterns of 365 bp, 325 bp, and 150 bp sizes. With the enzyme Hae III (lane B), the generated patterns were of 320 bp, 230 pb, 180 bp, and 150 bp, the enzyme Hinf I (lane C) generated a 365 bp profile and one of 155 bp. Comparing these restriction profiles with those created in the control yeast S. cerevisiae 288C (II., lanes A, B, C respectively), it can be observed that they are similar, hence the LEVM and LEVZ yeasts belongs to the genus S. cerevisiae.
In the few works dealing with the characterization of the microbiota involved in the fermentation process of the different Agave spirits, the role played by non-Saccharomyces and Saccharomyces yeasts has been determined. In Mezcal from Oaxaca, Andrade-Meneses and Ruiz Teran (2004) isolated Candida, Hanseniaspora, Rhodothorula, and S. cerevisiae species. From a natural fermentation of Agave fourcroydes must, Lappe et al. (2004) reported a great diversity of yeasts (Candida spp., C. parapsilosis, C. lusitaniae, Debaryomyces hansenii, K. marxianus, Ogataea siamensis, Pichia angusta, Pichia caribbica, P. guilliermondii, Rhodotorula mucilaginosa, Rhodotorula spp., and T. delbrueckii) and a population of 3.9 x 105 cells mL-1 at the beginning of the fermentation, which increased to 1.3 x 108 cells mL-1 after 24 h. Fermented must underwent a dramatic reduction in yeast heterogeneity and the population diminished to 1.4 x 107 cells mL-1 after 48 h, with K. marxianus and S. cerevisiae being the predominant species. Escalante-Minakata et al. (2008) identified yeast and bacteria present in A. salmiana fermentations, where the microbial diversity was dominated by Z. mobilis ssp. Mobilis. Regarding yeast species, only C. lusitaniae, K. marxianus, and Pichia fermentans were identified. In these few papers published on the mezcal mycobiota, it appears that non-Saccharomyces yeasts play an important role in the initial fermentation stages and influence the generation of volatile compounds involved in the aromatic profile of the final product (Escalante-Minakata et al., 2008). Flores Berrios et al. (2005) used amplified fragment length polymorphism (AFLP) to detect DNA polymorphism, genotype identification, and genetic diversity between S. cerevisiae, Candida spp., and Hanseniaspora spp. Strains isolated from different Agave species, sotol (Dasylirion spp.), and grape musts. A direct correlation between the genetic profile, origin, and fermentation process was found particularly in Agave must strains. Little information is available on the evolution of yeast populations during the fermentative process. In the case of tequila, the population of S. cerevisiae reached 1.8 - 2.0 x 108 cells mL-1 after 7 h of cultivation, when the inoculum was developed under optimal conditions (sugar concentration between 50 and 80 g L-1, continuous aeration, temperature of 30 °C, and addition of a nitrogen source). During fermentation, with an initial population of 2.0 - 2.5 x 107 cells mL-1 and an initial concentration of sugar 140 g L-1, the fermentative process took 24 h; the yeast population reached 1.1 - 1.2 x 108 cells mL-1, with an alcohol production between 50 and 60 g L-1. The yeast population remained high throughout the process. In Mezcal from Oaxaca, the native yeast population, mainly non-Saccharomyces, reached 1.5 - 4.0 x 107 cells mL-1 and declined during the fermentative process. This reduction could be associated with the lower alcohol tolerance of these kinds of yeasts or some nutritional limitation; also, 50 g L-1 of ethanol was obtained after 58 days of fermentation with an initial 150 g L-1 of sugar concentration (Gschaedler et al., 2004).
3.2 Experimental design
Our aimed is to maximize the production of ethanol in the production of Mezcal, so the variables as cell growth response and yield of ethanol were established. It is important to understand the kinetic behavior of the strain, and cell growth is also considered as response variable. The experimental variables that were used to build a Box-Behnken experimental design were A: pH (4.5-5.5), B: initial substrate (12-14 ° Brix) and C: temperature (28-32 ° C).
Variables were established as cell growth response and yield of ethanol, as we aimed to maximize the production of this metabolite in the production of Mezcal. It is important to understand the kinetic behavior of the strain, therefore cell growth is also considered as response variable.
Table 1 shows the data matrix of 15 treatments performed and experimental results of cell growth and ethanol Our aimed is to maximize the production of ethanol in the production of Mezcal, so the variables as cell growth response and yield of ethanol were established. It is important to understand the kinetic behavior of the strain, and cell growth is also considered as response variable.
]]>
The results obtained in each of the treatments show that not all combinations tested resulted in the same amount of cells and ethanol; being treatments No.7 and 9 the most outstanding in the amount produced and No. 15 for the percentage of ethanol: 7.92% v / v.
The analysis of experimental design using response surface methodology is shown in the Pareto chart (Fig. 2); this type of analysis allows studying the influence of variables on the response (production of biomass and ethanol) and their interactions. Figures 3 and 4, as well as the Pareto diagram reveal which experimental factor is most influential in terms of the output variable; in addition it allows estimating the range of values in which range of values of each factor is possible to obtain a more favorable result.


]]>

It is observed that for the production of biomass the most influential factor is temperature, being ideal the use of an intermediate temperature (30 °C) to produce more cell growth (Fig. 3). For the production of ethanol it is observed that the most influential factor was also the temperature, being more favorable to use low temperatures (28 °C).
The above diagrams suggest the temperature change that is required and its effect on product performance. The effect is best visualized as described above in the response surface curves (Fig. 4). In them we see that biomass production is greater when using intermediate temperatures (30 °C), noting that neither pH nor initial substrate concentration has significant effects in terms of cell growth. To obtain larger quantities of ethanol, the process should approach higher temperatures (30-32 °C), high pH (5.0-5.5) and high initial concentrations of substrate (14 °Brix) noting also that the last two are not as significant factors for production of this metabolite.
Figure 5 shows the contour plot response surface, demonstrating, like the response surface diagrams, optimal points of the process to obtain better yields of the product.

In Table 2 we can see the analysis of variance for cell growth, which indicates which of the experimental factors and interactions between them are significant for the process. We can see that the temperature (°C) and the interaction between them (CC) are the most significant with a P value < 0.05.
]]>
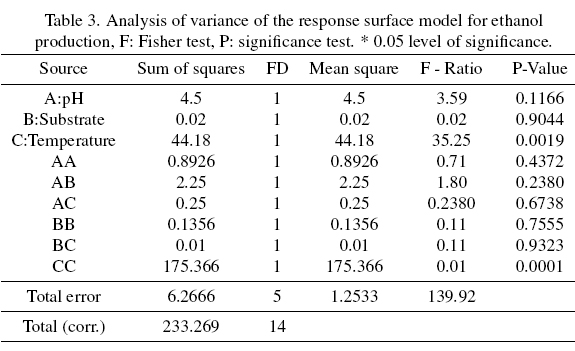
The analysis of variance of Table 3 shows that the effect of temperature (C) and temperature-temperature interaction (CC) are significant experimental factors for the process. The analysis yields an R2 of 97.31% and 94.30%, respectively, indicating their percentage significance in our process. Figure 6 shows the confirmatory kinetics of cell growth, substrate consumption, pH and ethanol production of yeast LEVM (Assay 15), which presented an ethanol yield of 12.96% v/v (Test Procedure K-ETOH 11 / 05, Megazyme). Cell growth increased from an initial load of 3x106 cells mL-1 to approximately 1.355x 108 cells mL-1. The initial substrate for this test was 132.82 g L-1 (14 ° Brix), reaching a final amount of 7.28 g L-1. Finally, the behavior of the pH varied from an initial pH of 5.0 to 3.7.
De León-Rodríguez et al. (2008) optimized the fermentation conditions for the production of Agave salmiana Mezcal with the native microbiota. The highest ethanol production (37.7 g L-1) was obtained in must with 105 g L-1 of sugars, and 1 g L-1 of ammonium sulfate, fermented 15 h at 28 °C. At the end of the fermentation the biomass (yeasts and bacteria) concentration reached 1.04 g L-1. Arrizon et al. (2006) compared the behavior of yeasts of different origins during fermentation of A. tequilana Weber var. azul and grape musts.
In comparison with Agave yeasts (C. magnoliae, Issatchenkia orientalis, H. uvarum, and S. cerevisiae) grape yeasts (H. uvarum and S. cerevisiae) exhibited a reduced fermentation performance in Agave musts with a high sugar concentration, while both groups of yeasts showed similar fermentation behavior in grape must. The presence of toxic compounds like furfural and vanillin and the high concentration of fructose in the Agave most could explain the poor fermentation performance of the wine yeasts. Fiore et al. (2005) demonstrated that non-Saccharomyces Agave yeast strains (Candida krusei, C. magnolia and H. vineae) possess a high sulfite and ethanol (10-12%) tolerance in controlled fermentations under laboratory conditions. These experimental results on ethanol tolerance contradict what was found in the traditional Mezcal process where different yeast strains and different fermentation conditions prevail; however, it highlights an important characteristic that must be studied more thoroughly.
]]> Conclusions
Our study shows that a series of conditions can lead to an improvement in the production of Mezcal by simultaneously analyzing different variables involved in the alcoholic fermentation and establishing the influence of each one on the amount of ethanol produced.
The results of the RFLP technique used for the molecular characterization of the isolated yeast LEVM suggests that the isolated yeast strain belongs to the Saccharomyces cerevisiae genus showing restriction patterns similar to those obtained with yeast belonging to that characterized genus (S. cerevisiae 288C).
By the response surface methodology, it was possible to find the formulation of a medium that will improve the production of ethanol (12.96% v/v) using the LEVM isolated strain, for which the process should take place at temperatures between 30-32 ºC, pH values of 5.0-5.5, and initial substrate concentrations between 12-14 °Brix.
The great variety of Agaves and their multiple uses have played an important role in the cultural identification of México. They have been exploited in many ways for over 10 000 years, and one of these applications is the production of alcoholic and distilled beverages. Until today, the microbiota that participates in the fermentation and its biochemical role in this process remain largely unknown; therefore, it is essential to carry out more studies on the traditional processes that are still in use because they are the source of important microbial consortia that could disappear with the introduction of new technologies. A detailed phenotypical and genotypical characterization of the microbiota must be carried out in order to conserve this specific biodiversity and subsequently evaluate its potential as starter cultures and in the production of different chemical compounds of biotechnological importance. In addition, it was shown to be a powerful tool for demonstrating the relationship between molecular profile, strain origin and fermentation process. Even though in future an extensive characterization must be performed with other wine and Mexican beverage strains, these preliminary results show the importance of using molecular techniques for the characterization of yeast strains used in the beverage industry.
Acknowledgements
This work was partially supported by Grant, PROMEP 103.5/12/3679, DGEST and CIC 20.22 (UMSNH).
References
]]>Andrade-Meneses, O. E. and Ruiz-Teran, F. (2004). Study of yeast populations in a mezcal fermentation. Presentation PF-17. 15-20 August. Rio de Janeiro, Brazil. 11th International Congress on Yeasts. Yeasts in Science and Technology, The quest for Sustainable Development. [ Links ]
Arrizon, J., Fiore, C., Acosta, G., Romano, P. and Gschaedler, A. (2006). Fermentation behavior and volatile compounds production by agave and grape must yeasts in high sugar Agave tequilana and grape must fermentations. Antonie van Leeuwenhoek 8, 181-189. [ Links ]
Cedeño, M. C. (1995). Tequila Production. Critical Reviews in Biotechnology 15, 1-11. [ Links ]
Cuinier, C. (1985). Le levurage specifique. Viticulture. Tourangelle. 215, 15-18. [ Links ]
De León-Rodríguez, A., Escalante-Minakata, P., Barba de la Rosa, A. and Blaschek, H.P. (2008). Optimization of fermentation conditions for the production of the mezcal from Agave salmiana using response surface methodology. Chemical Engineering Progress 47, 76-82. [ Links ]
]]>Escalante-Minakata, P., Blaschek, H.P., Barba de la Rosa, A. P., Santos, L. and De León-Rodríguez, A. (2008). Identification of yeast and bacteria involved in the mezcal fermentation of Agave salmiana. Letters in Applied Microbiology 46, 629-638. [ Links ]
Esteve-Zarzoso, B., Belloch, C., Uruburu, F. and Querol, A. (1999). Identification of yeast by RFLP analysis of the 5.8S rRNA gene and the two ribosomal internal transcribed spacers. International Journal of Systematic Bacteriology 49, 329-337. [ Links ]
Fiore, C., Arrizon, J., Gschaedler, A., Flores, J. and Romano, P. (2005). Comparison between yeasts from grape and agave musts from traits of technological interest. World Journal Microbiology and Biotechnology 21, 1141-1147. [ Links ]
Fleet, G.H. and Heard, G.M. (1993). Yeast growth during fermentation. In: Wine Microbiology and Biotechnology, (Ed. G.H. Fleet), Pp. 27-54. Harwood Academic Publishers, Switzerland. [ Links ]
Flores-Berrios, E.P., Alba-González, J.F., Arrizon-Gaviüo, J.P., Romano P., Capece, A. and Gschaedler-Mathis, A. (2005). The uses of AFLP for detecting DNA polymorphism, genotype identification and genetic diversity between yeasts isolated from Mexican agave-distilled beverages and from grape musts. Letters in Applied Microbiology 41, 147-152. [ Links ]
]]>Frezier, V. and Dubourdieu, D. (1992). Ecology of yeast strain Saccharomyces cerevisiae during spontaneous fermentation in a Bordeaux winery. American Society for Enology and Viticulture 43, 375-380. [ Links ]
Gallardo, V. J., Gschaedler, M. A. C., Chazaro, B. M., Tapia, C. E., Villanueva, R. S., Salado, P. J. H., Villegas, G. E., Medina, N. R., Aguirre, O. M. and Vallejo, P. M. (2008). La producción de mezcal en el estado de Michoacán. Gobierno del Estado de Michoacán y Centro de Investigación y Asistencia Tecnológica y Diseño del Estado de Jalisco. Michoacan México. [ Links ]
Grupo de estudios ambientales. (2002). Informe de mercadeo maguey-mezcal. Disponible en: http://www.dfid.gov.uk/r4d/PDF/Outputs/Forestry/R7925f_Maguey-mezcal.pdf . Accesado: 4 Mayo 2012. [ Links ]
Gschaedler, M., A., Ramírez, C. J., Diaz M., D., Herrera L., H., Arellano, P., M., Arrizon G., J. and Pinal, Z. L. (2004). Fermentación: etapa clave en la elaboración Perspectivas (Centro de Investigación y Asistencia Tecnológica y Diseño del Estado de Jalisco, ed.), Pp. 32-120. CIATEJ, Guadalajara, México. [ Links ]
Gutiérrez, P. H., De la Vara, S. R. (2008). Análisis y Diseño de Experimentos. Editorial McGraw-Hill. México. [ Links ]
]]>Jacques-Hernandez, C., Soto-Cruz, O.N., Rutiaga-Quiñones, O. M., Sifuentes-Rincón, A.M., Taillandier, P., Ramón-Portugal, F., (2009). Ecología de levaduras del mezcal San Carlos Tamaulipas: Presentación 0X-08. 21-26 Junio. Acapulco, Guerrero, México: XIII Congreso Nacional de Biotecnología y Bioingeniería y VII Simposio Internacional de Producción de Alcoholes y Levaduras. [ Links ]
Kreger-Van Rij, N.J.W. (1984). The yeast, a taxonomic study. Elsevier, Amsterdam. [ Links ]
Kunkee, R.E. and Amerine, M. (1970). Yeasts in winemaking. In: The Yeasts, 3: Yeast Technology. (A.H. Rose y J.S. Harrison eds.). Pp. 5-72. Academic Press, London. [ Links ]
Lafon-Lafourcade, S. (1983). Wine and brandy. In: Biotechnology, vol. 5: Food and Feed Production with Microorganisms, (H.J. Rehm y G. Reed eds.), Pp. 81-163. Verlag Chemie, Weinheim. [ Links ]
Lappe, P. and Ulloa, M. (1993). Microbiología del pulque. Alimentos fermentados indígenas de México (Wacher, M.C. and Lappe, P., eds.), Pp. 75-80. Universidad Nacional Autónoma de México, México. [ Links ]
]]>Molina, G. J. A., Botello, A. J. E., Estrada, B. A., Navarrete, B. J. L., Jimeínez, I. H., Cárdenas, M. M. and Rico, M. R. (2007). Compuestos volaítiles en el mezcal. Revista Mexicana de Ingeniería Química 6, 41-50. [ Links ]
Montgomery, D. (2004). Diseño y Análisis de Experimentos. Editorial Limusa. México. [ Links ]
Ribéreau-Gayon, P., Glories, Y., Maujean, A., Dubourdieu. (1998). Traité d'oenologie. vol. 2. Chimie du vin, Stabilisation and traitements. Editorial Dunod, París. [ Links ]
Salas, E. and Arenas, R. (2001). Biología molecular en micología medica. Dermatología Venezolana 39, 7-10. [ Links ]
Torija, M. (2002). Ecología de levaduras: Selección y adaptación a fermentaciones vínicas. Tesis de Doctorado en Bioquímica, Universidad Pública de Tarragona, España. [ Links ]
]]>Vezinhet, F., Hallet, J., Valade, M. and Poulard, A. (1992). Ecological survey of wine yeast strains by methods of identification. American Society for Enology and Viticulture 43, 83-86. [ Links ]
Versavaud, A., Dulau, L. and Hallet, J.N. (1993). Ecological study of the yeast microflora spontaneous Charentes vineyards and molecular approach of intraspecific diversity in Saccharomyces cerevisiae. Revue Francaise d'Oenologie 142, 20-28. [ Links ]
Zambonelli, C. (1988). I lieviti selezionati. Editorial Microbiologia e biotecnologia dei vini. Edagricole, Bologna. [ Links ]
]]>