Culture–independent characterization of the bacterioplankton community composition of a mesotrophic reservoir (Embalse Río III, Argentina)
Caracterización independiente de la composición de la comunidad del bacterioplancton en un embalse mesotrófico (embalse Río III, Argentina)
Daniela Polverino1*, Alejandro J. Mariñelarena2, Christina B. McCarthy1 and Rolando V. Rivera–Pomar1
1 Centro Regional de Estudios Genómicos, Universidad Nacional de La Plata (UNLP), Av. Calchaquí km 23.5 4° piso, (1888) Florencio Varela, Argentina. *polverino.d@gmail.com.
]]> 2 Instituto de Limnología "Dr. Raúl A. Ringuelet", Facultad de Ciencias Naturales y Museo (UNLP)–CONICET, Av. Calchaquí km 23.5 2° piso, (1888) Florencio Varela, Argentina.
Recibido: 26 julio 2011
Aceptado: 07 febrero 2012
Abstract
This study represents the first analysis of the bacterioplankton community structure from a freshwater reservoir of Argentina using amplification of the entire 16S rDNA gene. It includes the description and the phylogenetic relationships of the bacterioplankton community from the photic and aphotic layers of the Río III Reservoir in Córdoba, Argentina. The classical ecological approach indicated that the photic layer had greater diversity whereas the aphotic layer had a better distribution of species and higher abundance. Nevertheless, when the microbial communities in both layers were compared using phylogenetic information, this analysis indicated that both environments were similar and that neither was enriched for any particular lineage. The phyla present in the Río III reservoir were Acidobacteria, Actinobacteria, and Proteobacteria and the 2 dominant species in both layers were "Candidatus Planktophila sp." (class Actinobacteria) and Polynucleobacter sp. (class Betaproteobacteria).
Key words: Actinobacteria, phylogenetic analysis, Proteobacteria, 16S rDNA.
Resumen
]]> Este estudio constituye el primer análisis de la estructura de la comunidad del bacterioplancton en un reservorio de agua dulce de Argentina utilizando amplificación completa del gen ADNr 16S. Incluye las relaciones filogenéticas y una descripción del bacterioplacton de las zonas fótica y afótica del Embalse Río III, Córdoba, Argentina. El análisis ecológico clásico indicó que la zona fótica tenía mayor diversidad, mientras que la afótica tenía mayor abundancia y una distribución más uniforme de especies. Sin embargo, cuando se utilizó la información filogenética para comparar las comunidades microbianas de ambas zonas, este análisis indicó que ambos ambientes eran similares y que en ninguno predominaba algún linaje en particular. Los phyla identificados en el Embalse del Río III fueron Acidobacteria, Actinobacteria y Proteobacteria. Las especies dominantes en ambas zonas fueron "Candidatus Planktophila sp." (clase Actinobacteria) y Polynucleobacter sp. (clase Betaproteobacteria).Palabras clave: Actinobacteria, análisis filogenético, Proteobacteria, ADNr 16S.
Prokaryotes represent a significant biomass component in aquatic planktonic ecosystems and play a major role in biogeochemical processes. Recent studies have examined the bacterioplankton community composition (BCC) in the epilimnion of freshwater lakes and reservoirs. As a result, a core group of bacterial phylotypes common to freshwater has emerged (Zwart et al., 2002), which shows that rivers and lakes have a specific bacterioplankton community, distinct from bacteria in environments such as soil and sediments. The dominant divisions include Proteobacteria, Bacteroidetes, Verrucomicrobia, Cyanobacteria, Actinobacteria, and green nonsulfur bacteria.
The Río III Reservoir (32°11' S, 64°13' W) was built in 1936 over the Río III for hydroelectric supply. The Río III basin was strongly modified by damming for multipurpose uses including hydroelectricity, drinking water, and irrigation. Since then, water level fluctuations have been strongly reduced and the reservoir is usually oligo–mesotrophic. The reservoir area is 4.3 km2, and has a volume of 5.6 × 102 hm3, with a mean depth of 12.2 m and a maximum depth of 46.5 m. Previous reports (Mariñelarena et al., 2007, 2008) have focused on the planktonic abundance and biomass and bacterioplankton secondary production of this reservoir. Nevertheless, this study represents the first description of the bacterial community composition (BCC) of any freshwater inland water body in Argentina using molecular techniques and its results constitute the basis for future comparative studies.
Triplicate integrated samples of the water column were taken by pumping at 5 regular depth intervals during the month of September 2008. The limit between the photic (PL) and aphotic (AL) layers was established at the depth where 1% of the photosynthetically active radiation was measured (PAR 1%). Samples were stored at 4 °C until they were processed. Three aliquots of 2 ml of water of each sample were processed according to Boström et al. (2004). The 16S rDNA gene was amplified using universal bacterial primers 8f (5' AGA GTT TGA TCC TGG CTC AG 3') and 1542r (5' AGA AAG GAG GTG ATC CAG CC 3') (Crump et al., 1999). The PCR amplified 16S rDNA sequences were cloned into TOPO–TA PCR Cloning Kit pCR 2.1–TOPO Vector® (Invitrogen, Carlsbald, California, USA) in accordance with the manufacturer's instructions and the ligation products were transformed into competent Escherichia coli DH5α. DNA was purified with Illustra plasmidPrep Mini Spin Kit (GE Healthcare, Little Chalfont, Buckinghamshire, UK) and sequenced by Macrogen Inc, Korea.
The full or partial length 16S rDNA gene sequences from the 63 clones generated in this study were deposited in GenBank under accession numbers HQ008536 – HQ008557 (from AL) and HQ008558 – HQ008598 (from PL). Sequences were compared against 2 ribosomal databases: Ribosomal Database Project (RDP; http://rdp.cme.msu.edu/; Cole et al., 2009) and EMBnet (http://www.ch.embnet.org/software/aBLAST.html). All sequences were checked for chimera using the http://www.bioinformatics–toolkit.org/Help/Topics/chimera.html online tool, edited with BioEdit Sequence Alignment Editor 7.0.9.0, and aligned with Clustal × 2.0.11. Bayesian analysis (see details in Figure 2) was performed to build the phylogenetic tree using only full length 16S rDNA sequences, an outgroup (AB523784), and reference clones from GenBank.
To determine bacterial abundance, a volume of 200 µl of water of each layer was filtered. Cells were stained with Live/Dead®BacLightTM Bacterial Viability kit L–13152 (Molecular Probes Inc., USA), according to the manufacturer's instructions. Fifteen photos of each sample were taken using a fluorescence binocular microscope (Leica DM 1000) with a built–in CCD camera (CoolSNAP–Pro cf color, Media Cybernetics). Cells were counted and their abundance was calculated using N= Y * A * D / a * v; where N is the number of bacteria per ml; Y is the average number of organisms per field; A is the effective filtration area; D is the dilution coefficient; a is the microscope's visual field area; v is the volume of filtered water sample. The SPSS 15.0 program was used to analyze and compare these values statistically.
The Shannon–Wienner and Pielou indexes (Shannon and Wiener, 1949; Pielou, 1966) were used to describe generic diversity (H') and evenness (J) distribution in PL and AL communities. Clones which could not be resolved at the genus level (HQ008551, HQ008579, HQ008587) were incorporated in an upper level in the analysis.
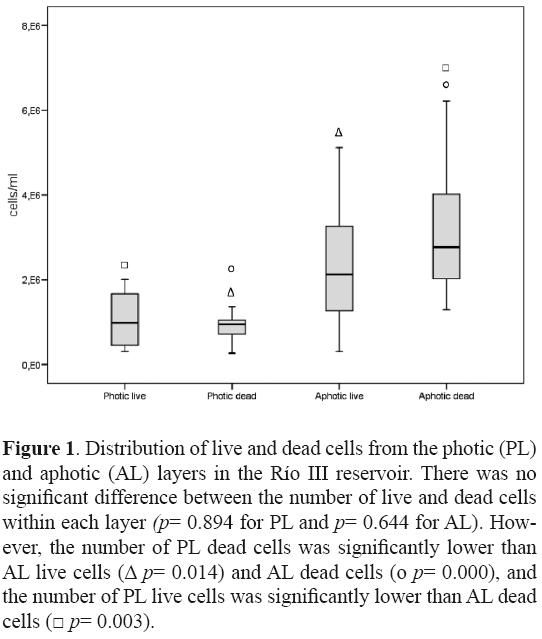
After confronting the sequences to free databases, results were comparable in 51.42% of the cases. Only 1 clone was classified as unidentified bacteria (HQ008579) and clone HQ008548 was reclassified as Polynucleobacter sp. (phylum Proteobacteria; class Betaproteobacteria) at the moment of writing this research note. From the total of 63 clones, 41 corresponded to PL and 22 to AL (see Table 1 and text below). Total bacteria abundance in the Río III reservoir (Fig. 1) was similar to others reports (Pedrós–Alió and Guerrero, 1994), even though there was a significant difference between PL and AL, 2.32 × 106 cells/ml and 5.85 × 106 cells/ml, respectively (P = 0.000). Furthermore, the Shannon–Wienner diversity index at genus level was 1.77 and 1.63 for PL and AL, respectively, whereas evenness was 0.77 and 0.84, respectively. This indicated that PL had a greater number of species but that the distribution of species in AL was more even, in concordance with previous studies (Markosova et al., 1990; Glockner et al., 1999).
]]>
Based on the relative abundance of clones, the most abundant group in both layers was "Candidatus Planktophila sp." (class Actinobacteria), comprising 36.59% (15/41) of clones from PL and 40.91% (9/22) from AL. Polynucleobacter sp. followed in abundance constituting 26.83% (11/41) of the clones from PL and 22.73% (5/22) from AL. For the rest of the clones, the relative abundance was lower than 10%. The prevalence of Actinobacteria clade acI in freshwater could be related to its high UV resistance capacity (Warnecke et al., 2005). The high number of Betaproteobacteria that inhabit freshwater could be due to its diverse metabolic composition (Burket et al., 2003), as this would enable phylogenetically similar taxa to occupy different niches within the same physical space (Newton et al., 2006). Furthermore, Actinobacteria and Betaproteobacteria tend to co–occur and are very abundant in most lakes, suggesting continuous feeding with allochthonous bacteria from the catchment area (Barberán and Casamayor, 2010). "Candidatus Planktophila sp." and Polynucleobacer sp. were dominant in both layers, also in accordance with previous reports (Jezbera et al., 2009; Barberán and Casamayor, 2010; Hahn et al., 2010). Moreover, both species have a cosmopolitan distribution in freshwater and represent an important fraction of the plankton (Jezbera et al., 2009; Hahn et al., 2010).
Phylogenetic analysis (Fig. 2) revealed that the reservoir's bacterioplankton community comprised 3 phyla: Proteobacteria, Actinobacteria, and Acidobacteria. Furthermore, phyla Proteobacteria and Actinobacteria each formed 2 phylogenetically divergent groups (nodes II and VI for Actinobacteria and nodes VII and XI for Proteobacteria). The only representative of phylum Acidobacteria formed a branch (XV) phylogenetically more related to phylum Proteobacteria (node X). Phylogenetic clustering of Ca. Planktophila sp. (phylum Actinobacteria) (nodes III and VI) and Polynucleobacter sp. (phylum Proteobacteria) (nodes VII and XI), whose sequences dominated the BCC, indicated the existence of 2 evolutionarily divergent groups in each genera. More studies should be made to confirm and establish the significance of this divergence and to further characterize the natural bacterioplankton community and the way in which the environmental conditions determine the composition of this assemblage. In conclusion, this study represents the first analysis of bacterioplankton diversity from an Argentine freshwater reservoir using amplification of the entire 16S rDNA gene. The phyla found in the PL and AL of the Río III reservoir in Córdoba, province of Argentina, form part of the recently described core group of bacterial phylotypes common to freshwater (Zwart et al., 2002). Two genera, "Candidatus Planktophila sp." (class Actinobacteria) and Polynucleobacter sp. (class Betaproteobacteria), dominated the water column.
We thank Dr Miguel Di Siervi for his expertise and assistance with the microscope. This work was supported by ANPCYT, Ministerio de Ciencia, Tecnología e Innovación Productiva de la Nación, Argentina (grant PICT20366/07), and the Max Planck Society through the External Partner Laboratory Grant to R. V. R. P. (R. V. R. P. is a investigator of the Consejo Nacional de Ciencia y Tecnología of Argentina, CONICET).
Literature cited
Barberán, A. and E. O. Casamayor. 2010. Global phylogenetic community structure and β–diversity patterns in surface bacterioplankton metacommunities. Aquatic Microbial Ecology 59:1–10. [ Links ]
Boström, K. H., K. Simu, A. Hagström and L. Riemann. 2004. Optimization of DNA extraction for quantitative marine bacterioplankton community analysis. Limnology Oceanography: Methods 2:365–373. [ Links ]
Burkert, U., F. Warnecke, D. Babenzien, E. Zwirnmann and J. Pernthaler. 2003. Members of a readily enriched beta–proteobacterial clade are common in surface waters of a humic lake. Applied and Environmental Microbiology 69:6550–6559. [ Links ]
Cole, J. R., Q. Wang, E. Cardenas, J. Fish, B. Chai, R. J. Farris, A. S. Kulam–Syed–Mohideen, D. M. McGarrell, T. Marsh, G. M. Garrity and J. M. Tiedje. 2009. The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Research 37:141–145. [ Links ]
Crump, B. C., E. V. Armbrust and J. A. Baross. 1999. Phylogenetic analysis of particle–attached and free–living bacterial communities in the Columbia River, its estuary, and the adjacent coastal ocean. Applied and Environmental Microbiology 65:3192–3204. [ Links ]
Glockner, F. O., B. M. Fuchs and R. Amann. 1999. Bacterioplankton compositions of lakes and oceans: a first comparison based on fluorescence in situ hybridization. Applied and Environmental Microbiology 65:3721–3726. [ Links ]
Hahn, M. W., E. Lang, U. Brandt, H. Lunsdorf, Q. L. Wu and E. Stackebrandt. 2010. Polynucleobacter cosmopolitanus sp. nov., free–living planktonic bacteria inhabiting freshwater lakes and rivers. International Journal of Systematic Evolutionary Microbiology 60:166–173. [ Links ]
Jezbera, J., A. K. Sharma, U. Brandt, W. F. Doolittle and M. W. Hahn. 2009. 'Candidatus Planktophila limnetica', an actinobacterium representing one of the most numerically important taxa in freshwater bacterioplankton. International Journal of Systematic Evolutionary Microbiology 59:2864–2869. [ Links ]
Mariñelarena, A., M. Casco, C. Claps, M. Di Siervi, J. Donadelli, M. Hechem, D. Colautti, M. Mac Donagh, L. Balboni and D. Ardohain. 2007. Estudio limnológico del Embalse del Río III, Córdoba. Informe final. Instituto de Limnología "Raúl A Ringuelet". Universidad Nacional de La Plata – CONICET, Florencio Varela. [ Links ]
Mariñelarena, A., M. Casco, C. Claps, M. Di Siervi, J. Donadelli, M. Hechem, D. Colautti, M. Mac Donagh, L. Balboni and D. Ardohain. 2008. Estudio limnológico del Embalse del Río III, Córdoba. Informe final. Instituto de Limnología "Raúl A Ringuelet". Universidad Nacional de La Plata – CONICET, Florencio Varela. [ Links ]
Markosova, R., M. Benediktova and A. Volkova. 1990. Time and vertical distribution of bacterioplankton in a shallow eutrophic reservoir. Water Research 24:1057–1067. [ Links ]
Newton, R. J., A. D. Kent, E. W. Triplett and K. D. Mcmahon. 2006. Microbial community dynamics in a humic lake: differential persistence of common freshwater phylotypes. Environmental Microbiology 8:956–970. [ Links ]
Pedrós–Alió, C. and R. Guerrero. 1994. Prokaryotology for the limnologist. In Limnology now: a paradigm of planetary problems, R. Margalef (ed.). Elsevier, Amsterdam. p. 37–57. [ Links ]
Pielou, E. C. 1966. Species–diversity and pattern–diversity in the study of ecological succession. Journal of Theoretical Biology 10:370–383. [ Links ]
Shannon, C. E. and W. Wiener. 1949. The mathematical theory of communication. Urbana: University of Illinois Press. 125 p. [ Links ]
Warnecke, F., R. Sommaruga, R. Sekar, J. S. Hofer and J. Pernthaler. 2005. Abundances, identity, and growth state of actinobacteria in mountain lakes of different UV transparency. Applied and Environmental Microbiology 71:5551–5559. [ Links ]
Zwart, G., B. C. Crump, M. P. Kamst–Van Agterveld, F. Hagen and S. K. Han. 2002. Typical freshwater bacteria: an analysis of available 16S rRNA gene sequences from plankton of lakes and rivers. Aquatic Microbial Ecology 28:141–155. [ Links ]
]]>