
Resistencia antibiótica de genotipos de cepas de Salmonella spp de cerdos sacrificados en rastros del Estado de México
Antibiotic resistance of Salmonella spp strain genotypes isolated from pigs slaughtered at abattoirs in the Estado de Mexico
Martín Talavera Rojas* Jorge Antonio Varela Guerrero* Nydia Edith Reyes Rodríguez* Salvador Lagunas Bernabé* Benjamín Valladares Carranza* María Uxua Alonso Fresan* Valente Velázquez Ordoñez*
* Centro de Investigación y Estudios Avanzados en Salud Animal, Facultad de Medicina Veterinaria y Zootecnia, Universidad Autónoma del Estado de México, Carretera Panamericana Toluca–Atlacomulco km 15.5, 50200, Toluca, México.
]]> Responsable del artículo:
Recibido el 27 de septiembre de 2010.
Aceptado el 26 de mayo de 2011.
Abstract
]]> Multiresistant Salmonella serovar Typhimurium strains are a worldwide problem in animal and human health. The aim of this study was to determine the frequency of some Salmonella spp resistance genes (cmlA/tetR, PSE–1, TEM, Sip B/C) in strains isolated from pigs slaughtered at abattoirs in the Estado de Mexico. Of 87 analyzed strains 22 (25.28%) had phenotypical resistance to chloramphenicol (30 μg), 15 (17.24%) to ampicillin (10 μg) and 54 (62.07%) to sulfamethoxazole (60 μg). The phenotypical and genotypical relation of the 87 strains was: of the 22 chloramphenicol resistant strains only 14 (63.63%) expressed the cmlA/tetR resistance gene, and of the 65 strains non–resistant to chloramphenicol only 36 (55.38%) expressed the cmlA/tetR resistance gene. Regarding the 15 ampicillin resistant strains only 2 (13.33%) were carriers of the PSE–1 gene and 7 (46.66%) presented the TEM gene; both genes confer genotypical ampicillin resistance. Of 72 non–resistant ampicillin strains, 11 (15.27%) carried the TEM gene which confers ampicillin resistance. Two Salmonella strains (2.28%) belonged to phagotype DT104. Strains not showing phenotypical resistance but carrying resistance genes have not been exposed to selection by competition, although they possess the mechanism to express such resistance.Key words: Salmonella Typhimurium, resistance, pigs, antibiotics.
Resumen
La aparición de cepas multirresistentes de Salmonella Typhimurium es un problema mundial, tanto en salud animal como en salud pública. El objetivo del presente trabajo fue determinar la frecuencia de algunos genes de resistencia (cmlA/tetR, PSE–1, TEM, Sip B/C) en cepas de Salmonella spp aisladas de cerdos en rastros del Estado de México. De las 87 cepas analizadas, 22/87 (25.28%) mostraron resistencia al cloranfenicol (30 μg), 15/87 (17.24%) a la ampicilina (10 μg) y 54/87 (62.07%) fueron resistentes al sulfametoxazol (60 μg). La relación fenotípica y genotípica de las 87 cepas analizadas fue: de las 22 cepas que presentaron resistencia fenotípica al cloranfenicol, sólo 14/22 (63.63%) expresaron el gen de resistencia cmlA/tetR, y de las 65 cepas que manifestaron sensibilidad al cloranfenicol, 36/65 (55.38%) expresaron el gen de resistencia cmlA/tetR. De las 15 cepas que expresaron resistencia a la ampicilina, sólo 2/15 (13.33%) mostraron el gen PSE–1, y 7/15 (46.66%) presentaron el gen TEM, ambos genes confieren resistencia genotípica a la ampicilina. De las 72 cepas que manifestaron sensibilidad a la ampicilina, 11 (15.27%) mostraron el gen TEM, el cual da resistencia a la ampicilina. De las 87 cepas de Salmonella sólo 2/87 (2.28%) expresaron el fagotipo DT104. Las cepas que son portadoras de genes de resistencia, pero no la manifiestan fenotípicamente, no han sido expuestas a una selección por competencia, por lo tanto, no expresan la resistencia fenotípica, pero cuentan con el mecanismo necesario para expresarla.
Palabras clave: Salmonella Typhimurium, resistencia, cerdos, antibióticos.
Introducción
La aparición de cepas de Salmonella Typhimurium con resistencia cromosómica a la ampicilina, cloranfenicol, estreptomicina, sulfonamidas y tetraciclinas (tipo R ACSSuT) en los años 90 en el Reino Unido, y que posteriormente tuvieron una distribución mundial, ha causado una preocupación en la salud animal y la salud pública debido a que los tratamientos ya no tienen el efecto esperado.1 Estudios realizados entre 1997 y 2003 en Taiwán, demostraron la aparición de cepas multirresistentes de Salmonella;2 estas cepas de Salmonella Typhimurium fagotipo DT104 han mostrado una multirresistencia a antibióticos como la ampicilina, cloranfenicol y sulfonamidas.3,7 Los genes relacionados con dicha resistencia son: cmlA/tetR, el cual proporciona resistencia al cloranfenicol; el SipB/C, que confiere resistencia a las sulfonamidas; el PSE–1 y TEM, los cuales dan resistencia a la ampicilina.8,9
En la actualidad, la presencia de cepas de Salmonella spp multirresistentes en todo el mundo, ha causado un gran problema en la terapéutica de esta enfermedad debido a que los antibióticos utilizados comúnmente ya no funcionan o no se obtienen los resultados deseados.10
]]> El objetivo del presente trabajo fue identificar los genes cmlA/tetR, PSE–1, TEM, Sip B/C los cuales proporcionan resistencia al cloranfenicol, ampicilina y sulfonamidas en cepas de Salmonella spp aisladas de cerdos en el Estado de México.
Material y métodos
Preparación de los aislamientos de Salmonella
Las cepas utilizadas en este trabajo fueron obtenidas de cerdos para abasto provenientes de varios estados del país, sacrificados en rastros del Estado de México (Cuadro 1).
Resistencia fenotipica
La evaluación de la resistencia bacteriana a los antimicrobianos se realizará mediante el método Kirby–Bauer según lo establecen en la normativa del Clinical Laboratory Standars Institute (CLSI).11
Extracción del ADN bacteriano
Se obtuvieron 3 ml de crecimiento bacteriano, el cual fue centrífugado a 170,000 g. Posteriormente se decantó el sobrenadante y se resuspendió en 1 ml de PBS (Solución buffer fosfato salino); se lavó 4 veces y en el último lavado se resuspendió el comprimido con 1 ml de PBS (solución buffer de fosfatos), se le agregaron 3 μl de NaOH 1N y 22 μl de Tris–EDTA (Tris–HCl 10mM; EDTA 1mM) y se colocó en baño maría en seco a 90°C durante 25 min; por último se centrifugo brevemente y se conservó en congelación a –20°C, hasta el momento de su utilización.12
Preparación de las reacciones
]]> Las secuencias de los iniciadores que se utilizaron se describen en el Cuadro 2.La reacción consistió en 2.5 μl 10x de amortiguador de PCR, 1.5 μl de 25 mM MgCl2, 1μl de 200 mM DNTPs, 2 μl de cada primers (SipB/C, cmlA/tetR, PSE–1, TEM, DT 104), 1 μl (1 unit/μl) de Taq ADN polimerasa,* 3 μl de ADN bacteriano y 13 μl de agua inyectable, las reacciones se sometieron a una preincubación a 95°C por 5 minutos, posteriormente se realizaron 40 ciclos con una temperatura de desnaturalización a 95°C por 1 minuto, alineación de los primers a 48°C por 30 segundos y una extensión de ADN a 72°C por 30 segundos. Al finalizar los ciclos, las reacciones se incubaron por 3 minutos a 72°C y posteriormente se bajó la temperatura a 4°C para detener la reacción.13
Se tomaron 5 μl de cada reacción y se sometió a electroforesis en gel de agarosa al 3% por 45 minutos a 100 V con 1.5 μl de amortiguador de carga (TBE 1x). Los productos de PCR fueron visualizados con bromuro de etidio y fueron expuestos en un transluminador de UV.**13
Análisis estadístico
Las frecuencias de resistencia fenotípica–genotípica se evaluaron mediante medidas de asociación epidemiológica usando una tabla de contingencia de 2 × 2, con una significancia de P > 0.05, usando el riesgo relativo (RR) y razón de probabilidades (OR).14,15
Resultados
De las 87 cepas analizadas, 2/87 (2.29%) mostraron una banda de 102 pb, correspondiente al fagotipo DT104; 50/87 (57.47%) expresaron una banda de 260 pb, correspondiente al gen cmlA/tetR; 2/87 (2.29%) presentaron una banda de 132 pb correspondiente al gen PSE–1, y 18/87 (20.68%) mostraron una banda de 291 pb, correspondiente al gen TEM (Figuras 1 y 2). No se observó la banda de 232 pb que caracterizan al gen SipB/C en las cepas analizadas (Cuadro 3).

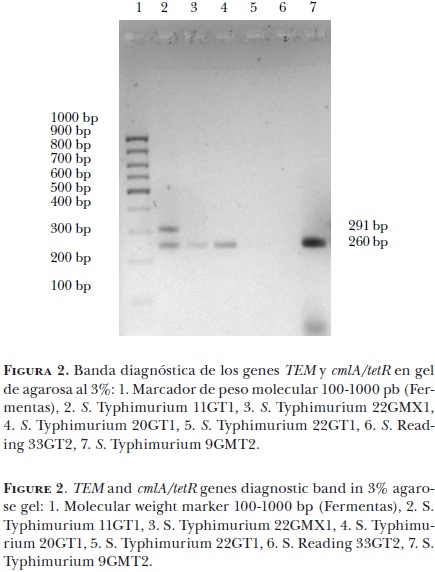
En cuanto a la relación fenotípica–genotípica realizada en el presente estudio, se encontró que de 22 cepas que mostraron resistencia fenotípica al cloranfenicol, 14 (63.63%) expresaron el gen de resistencia cmlA/tetR, y de 65 cepas que mostraron sensibilidad fenotípica al cloranfenicol, 36 (55.38%) mostraron el gen de resistencia (Cuadro 4).
De las 15 cepas que presentaron resistencia a la ampicilina, 2 (13.33%) mostraron el gen PSE–1 y 7 (46.66%) expresaron el gen TEM. De 72 cepas que mostraron sensibilidad fenotípica a la ampicilina in vitro, 11 (15.28%) presentaron el gen TEM (Cuadro 5).
Discusión
De 87 cepas analizadas, sólo 2 (2.98%) fueron fagotipo DT104, la presencia de dicho fagotipo en este trabajo fue muy baja, en comparación con otros trabajos realizados, 9,16–20 los cuales obtuvieron una frecuencia por arriba de 50%; sin embargo, se observa que la presencia de este fagotipo no se relaciona con la presencia de genes que confieren multirresistencia.3
Se observó que 57.41% de las cepas fueron positivas al gen cmlA/tetR, de las cuales 14 mostraron resistencia fenotípica al cloranfenicol; lo que indica una asociación positiva entre la presencia del gen de resistencia y la resistencia fenotípica (P < 0.05). Este gen es una fusión de dos genes: el cmlA y tetR, que le da una resistencia genética específica al cloranfenicol, según los trabajos realizados por Carlson et al.8 y Briggs y Fratamico,21 quienes encontraron este gen en un 10% y 97%, respectivamente; ambos trabajos se realizaron con cepas obtenidas de laboratorios de diagnóstico de algunos países, como Estados Unidos de América, Reino Unido y Alemania; sin embargo, en otro trabajo realizado en cepas aisladas de alimentos en Corea, no se presentó este gen.13 Actualmente, en México este fármaco se encuentra descontinuado, pero aún existe otro de la misma familia del cloranfenicol llamado florfenicol, el cual tiene un mecanismo de acción similar. En cuanto a las cepas que mostraron resistencia fenotípica al cloranfenicol, pero no manifestaron el gen (cmlA/tetR), probablemente la resistencia esté dada por otro gen llamado floR, mencionado en los trabajos de Weill et al.22 y Bolton et al.3 el cual confiere resistencia al florfenicol y al cloranfenicol.
Quince cepas mostraron resistencia fenotípica a la ampicilina, de las cuales 13.33% expresaron el gen PSE–1, y 46.66% el gen TEM, de las 72 cepas que mostraron sensibilidad a la ampicilina, 15.27% presentaron el gen TEM. Ambos genes confieren resistencia a la ampicilina.8,13 Aunque otros autores23,24 mencionan que el gen (PSE–1) confiere resistencia a los ß–lactamicos. La presencia de resistencia genotípica a la ampicilina en cepas aisladas de cerdos de rastros del Valle de Toluca es baja, pero va en incremento en comparación con países más desarrollados.23,24 La presencia del gen PSE–1 fue muy baja en comparación con los resultados obtenidos en cepas aisladas en Corea a partir de alimentos (26.7%),13 ambos resultados fueron mucho menores al observar los resultados obtenidos en Francia en el periodo 1993–2003 (70%) en cepas aisladas de humanos.22
El gen SipB/C, que confiere resistencia a las sulfas, no estuvo presente en las cepas de Salmonella spp analizadas en el presente estudio; por lo tanto, se puede decir que la resistencia de estas cepas (62.07%) no está ligada a la presencia de este gen.
Por los resultados mostrados en la relación genotipo–fenotipo se puede afirmar que las cepas sensibles fenotípicamente y que mostraron resistencia genotípica, probablemente no han sido expuestas al fármaco, pero es importante señalar que estas cepas ya cuentan con el gen, es decir, que existe la posibilidad de llegar a expresar la resistencia fenotípica, por algún mecanismo codificado por estos genes en caso de una exposición.25
]]>References
1. THRELFALL JE. Epidemic Salmonella Typhimurium DT104– a truly international multiresistant clone. J Antimicrob Chemother 2000; 46: 7–10. [ Links ]
2. CHIU CH, SU LH, CHU CH, WANG MH, YEH CM, WELLI FX et al. Detection of Multidru–Resistant Salmonella enterica Serovar Typhimurium Phage Type DT102, DT104, and U302 by Multiplex PCR. J Clin Microbiol 2006; 44: 2354–2358. [ Links ]
3. BOLTON FL, KELLEY CL, LEE DM, FEDORKA JP, MAURER JJ. Detection of multidrug–resistant Salmonella enterica Serotype Typhimurium DT104 based on a gene which confers cross–resistance to florfenicol and chloramphenicol. J Clin Microbiol 1999; 37: 1348– 1351. [ Links ]
4. RABATSKY–Ehr T, WHICHARD J, ROSSITER S, HOLLAND B, STAMEY K, HEADRICK ML et al. 276 Multidrug & – resistant strains of Salmonella enterica Typhimurium, United States, 1997–1998. Emerg Infect Dis 2004; 10: 795–801. [ Links ]
]]>5. HELMS M, ETHELBERG S, MOLBAK K. International Salmonella Typhimurium DT104 Infections, 1992–2001. Emerg Infect Dis 2005; 11: 859–867. [ Links ]
6. RIBOT EM, WIERZBA RK, ÂNGULO FJ, BARRETT TJ. Salmonella enterica serotype Typhimurium DT104 isolated from humans, United States 1985,1990, and 1995. Emerg Infect Dis 2002; 8:387–391. [ Links ]
7. YOKOYAMA E, MARUYAMA S, KABEYA H, HARA S, SATA S, KUROKI T et al. Prevalence and Genetic Properties of Salmonella enterica Serovar Typhimurium Definitive phage Type 104 Isolates from Rattus norvegicus and Rattus rattus House Rats in Yokohama City, Japan. Appl Envi Microbiol 2007; 73: 2624–2630. [ Links ]
8. CARLSON SA, BOLTON LF, BRIGGS CE, HURD HS, SHARMA V, FEDORKA PJ et al. Detection of multiresistant Salmonella Typhimurium DT104 using multiplex and fluorogenic PCR. Mol Cell Probes 1999; 13: 213–222. [ Links ]
9. ALVAREZ J, SOTA M, VIVANCO AB, PERALES P, CISTERNA R, REMENTERIA A et al. Development of a Multiplex PCR Technique for Detection and Epidemiological Typing of Salmonella in Human Clinical Simples. J Clin Microbiol 2004; 42: 1734–1738. [ Links ]
]]>10. LEE KE, LEE Y. Isolation of Multidrug–Resistant Salmonella Typhimurium DT104 from Swine in Korea. J Microbiol 2007; 45:590–592. [ Links ]
11. NCCLS. National Committee for Clinical Laboratory Standards. Performance standards for antimicrobial susceptibility testing; Nineteenth Informational Supplement. 10 ed. Vol. 29, No. 3. Approved standard M100–S19. Wayne, PA: National Committee for Clinical Laboratory Standards, 2009. [ Links ]
12. CHIU HC, OU TJ. Rapid Identification of Salmonella serovars in Feces by Specific Detection of Virulence Genes, invA and spvC, by an Enrichment Broth Culture–Multiplex PCR Combination Assay. J Clin Microbiol 1996; 34: 2619–2622. [ Links ]
13. YANG SJ, PARK KY, SEOK KS, YOO HS, NOH KM, KIM SH et al. Multidrug– resistant Salmonella Typhimurium and Salmonella enteritidis identified by multiplex PCR from animals. Vet Sci 2001; 2: 181 188. [ Links ]
14. DANIEL WW. Bioestadística (bases para el análisis de las ciencias de la salud). 4ª ed. México DF: Limusa Wiley, 2008. [ Links ]
]]>15. HIRD D, HERNÁNDEZ AJ, WILLIAMS JJ, RODRÍGUEZ BJC, SEGURA CJC. Medidas de asociación epidemiológica. Memorias del V Curso Internacional Teórico–Práctico en Epidemiología; 2000 noviembre 27 & – diciembre 1; Mérida (Yucatán) México. Mérida (Yucatán) México: Facultad de Medicina Veterinaria y Zootecnia. Universidad Autónoma de Yucatán, 2000: 19–27. [ Links ]
16. YU CY, CHOU JS, YEH MC, CHAO RC, HUANG CK, CHANG FY et al. Prevalence and Characterization of Multidrug–Resistant (Type ACSSu) Salmonella enterica Serovar Typhimurium Strains in Isolates from Four Gosling Farms and a Hatchery Farm. J Clin Microbiol 2008; 46: 522–526. [ Links ]
17. GRAZIANI C, BUSANI L, DIONISI A, LUCARELLI C, OWCZAREK S, RICCI A et al. Antimicrobial resistance in Salmonella enterica serovar Typhimurium from human and animal sources in Italy. Vet Microbiol 2008; 128: 414–418. [ Links ]
18. FRAZAN A, FRIENDSHIP RM, POPPE C, MARTIN L, DEWEY CE, FUNK J. Molecular epidemiology and antimicrobial resistance of Salmonella Typhimurium DT104 on Ontario swine farms. Can J Vet Res 2008; 72:188–194. [ Links ]
19. MIKO A, PRIES K, SCHROETER A, HELMUTH R. Molecular mechanisms of resistance in multidrug–resistant serovars of Salmonella enterica isolated from foods in Germany. J Antimicrob Chemother 2005; 56: 1025–1033. [ Links ]
]]>20. RANDALL LP, COOLES SW, OSBORN MK, PIDDOCK LJV, WOODWARD MJ. Antibiotic resistance genes, integrons and multiple antibiotic resistances in thirty–five serotypes of Salmonella enterica isolated from humans and animals in the UK. J Antimicrob Chemother 2004; 53: 208–216. [ Links ]
21. BRIGGS CE, FRATAMICO PM. Molecular Characterization of Antibiotic Resistance Gene Cluster of Salmonella Typhimurium DT104. Antimicrob Agents Chemother 1999; 43:846–849. [ Links ]
22. WEILL FX, GUESNIER F, GUIBERT V, TIMINOUUNI M, DEMARTIN M, POLOMACK L et al. Multidrug resistance in Salmonella enterica serotype Typhimurium from humans in France (1993–2003). J Clin Microbiol 2006; 44: 700–708. [ Links ]
23. KHAN AA, NAWAZ SM, KHAN AS, CERNINGLIA EC. Detection of multidrug–resistant Salmonella Typhimurium DT104 by multiplex polymerase chain reaction. FEMS Microbiol Letters 2000; 182: 355–360. [ Links ]
24. WALSH C, DUFFY G, NALLY P, O'MAHONY R, MCDOWELL DA, FANNING S. Transfer of ampicillin resistance for Salmonella Typhimurium DT104 to Escherichia coli K12 in food. Lett Appl Microbiol 2007; 46: 210–215. [ Links ]
]]>25. TALAVERA RM, VAZQUEZ CHJC, FLORES BR, ROBLES GF, LAGUNAS BS, ALONSO FMU. GyrA gene mutations and fluoroquinolone resistance in Salmonella isolates from pigs in central Mexico. Vet Rec 2007; 160: 630–632. [ Links ]
* Invitrogen, USA.
**Transilluminator (transluminador) modelo M–20E. Marca Upland, CA91786, USA.
]]>