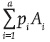 and
and 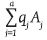 , then the genotypic array of the progeny produced by the cross between these two populations [GEA(PXQ)] is:
, then the genotypic array of the progeny produced by the cross between these two populations [GEA(PXQ)] is:
Genetic stability of synthetics derived from double-cross or three-way line hybrids
Estabilidad genética de sintéticos formados con cruzas dobles o trilineales
Jaime Sahagún-Castellanos
Instituto de Horticultura, Departamento de Fitotecnia, Universidad Autónoma Chapingo. Carretera México-Texcoco km 38.5, Chapingo, Estado de México. C.P. 56230. MÉXICO. Correo-e: jsahagunc@yahoo.com.mx
]]>Received: April 4, 2014.
Accepted: May 11, 2015.
Abstract
In the development of the theory of synthetic varieties (SVs) of crop species such as maize (Zea mays L.) and onion (Allium cepa L.), formed with single-cross hybrids, it has been shown that genetic erosion may occur during their formation. This erosion increases inbreeding and reduces yield. This gene loss is caused by the mating of heterozygous genotypes, the finite number of their progeny, and the randomness of the genetic mechanism. The main objective of this study was to determine the number of non-identical by descent (NIBD) genes lost during the development of the individuals that represent the double-cross (DC) or three-way (TW) line hybrids that in turn will be parents of a SV. The initial lines were assumed to be inbred and unrelated and each hybrid derived from them was represented by m individuals, and formulae for the mean, variance and number of lost NIBD genes per hybrid were derived. The number of lost NIBD genes of a DC (TW) was expressed as the number of NIBD genes in the 4 (3) initial lines minus the mean of NIBD genes forming the genotypes of the m individuals representing each DC (TW). It was found that each DC and TW loses on average 4/2m and 2/2m NIBD genes, respectively. The magnitudes of these losses reflect the effect of the sample size (m) and the number of loss sources (the single crosses) of the parents of the DCs (2) and TWs (1). The largest NIBD gene losses per parent were 2 (DC) and 1 (TW), which occur when m = 1. However, when m is large (m ≥ 12), as should occur in reality, the losses and the variance of the number of NIBD genes are nearly zero.
Keywords: Zea mays L., Allium cepa L., non-identical by descent genes, inbreeding coefficient, genotypic mean.
Resumen
En el desarrollo de teoría de las variedades sintéticas (VSs), de especies como el maíz (Zea mays L.) y la cebolla (Allium cepa L.), formadas con híbridos de cruza simple se ha evidenciado que durante su formación puede ocurrir erosión genética. Ésta incrementa la endogamia y reduce el rendimiento. Los causantes de esta pérdida de genes son el apareamiento de genotipos heterocigotos, el número finito de sus progenies y el azar del mecanismo genético. El principal objetivo de este trabajo fue determinar la pérdida de genes no idénticos por descendencia (NIPD), que ocurre en el desarrollo de los progenitores de VSs formadas con híbridos de cruza doble (CD) o triples (CT). Se consideró el uso de líneas puras no emparentadas para formar los híbridos; cada uno representado por m individuos, y se derivaron fórmulas para la media y la varianza del número de genes NIPD que se pierden. El número de genes NIPD perdidos en una CD (CT) se expresó como el número de genes NIPD en las 4 (3) líneas iniciales menos la media de genes NIPD de los m individuos que representan cada CD (CT). Se encontró que en cada CD y CT se pierden en promedio 4/2m y 2/2m genes NIPD, respectivamente. Las magnitudes de estas pérdidas reflejan el efecto del tamaño de muestra (m) y del número de fuentes de pérdida (las cruzas simples) de los progenitores de las CDs (2) y de las CTs (1). Las mayores pérdidas de genes NIPD por progenitor fueron 2 (CD) y 1 (CT), que ocurren cuando m = 1. Sin embargo, cuando m es grande (m ≥ 12), como debe suceder en la práctica, las pérdidas y la varianza del número de genes NIPD se reducen prácticamente a cero en ambos casos.
]]> Palabras clave: Zea mays L., Allium cepa L., genes no idénticos por descendencia, coeficiente de endogamia, media genotípica.
INTRODUCTION
Sahagún-Castellanos and Villanueva-Verduzco (1997) conducted a theoretical study of synthetic varieties (SVs) derived from a set of single-cross (SC) hybrids. This way of forming synthetics, which can be applied to species such as maize (Zea mays L.) and onion (Allium cepa L.), was later extended to double-cross (DC) hybrids (Sahagún-Castellanos, Rodríguez-Pérez, & Peña-Lomeli, 2005; Márquez-Sánchez, 2008), three-way (TW) line hybrids (Márquez-Sánchez, 2010) and mixtures of different types of hybrids (Sahagún-Castellanos & Rodríguez- Pérez, 2011; Sahagún-Castellanos & Villanueva-Verduzco, 2012). The idea of forming synthetic varieties in this way was inspired by practices carried out by some farmers to avoid the high annual seed cost of hybrid maize varieties. These farmers grow advanced generation hybrids or mixtures thereof, which have been viewed as SVs that would be formed with their parental lines. It is expected that these lines have already successfully passed through a selection process due to their combining ability, among other desirable characteristics. In the case of double crosses already released, it is expected that their parental lines will have good general and specific combining ability, which should be expressed in the population generated by randomly mating them in the form of a synthetic variety with good yield in the farmer's field.
However, there is evidence that a SV formed with double crosses cannot be the same as a SV that would be formed with the single crosses that are parents of the double crosses, and from that derived from the parental lines of such single crosses (Sahagún-Castellanos & Villanueva-Verduzco, 2012). These differences include changes in the inbreeding coefficient and the genotypic array. However, when the parents are double crosses, it is not precisely known whether these changes are reflected in the frequencies or even in the loss of some parental genes. The objectives of this study were to determine the mean and variance of the number of non-identical by descent (NIBD) genes passing from initial lines to double-cross or three-way line hybrids that will form a SV, and derive the number of NIBD genes that every parent loses in this process.
MATERIALS AND METHODS
This study was based on the model of a locus of diploid species reproduced by random mating. The starting point to form DC and TW hybrids was a set of L unrelated inbred lines (completely homozygous).
It should be noted that in the formation of L/2 SCs, NIBD genes cannot be lost. This is because each individual of the progeny of the cross between two inbred lines always carries two NIBD genes, each from a parental line. However, in the formation of L/4 DCs this may be different since each gamete from a SC can carry the gene of either line. For this reason, the number of NIBD genes reaching the genotypes that will represent each DC is a random variable. Likewise, genotypes formed with TWs contain a number of NIBD genes, which is a variable that takes random values because one parent of each three- way line hybrid is a single cross.
As in a previous study (Sahagún-Castellanos, Rodríguez-Pérez, & Escalante-González, 2013), this work was based on the concepts of gametic and genotypic array. If in a population the frequencies of the gene Ai (i = 1, 2,..., a) and genotype AiAj are pi and pij, the gametic (GAA) and genotypic (GEA) arrays, respectively, are defined as follows:
]]>
If the gametic arrays of the two populations P and Q are 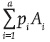 and
and 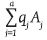 , then the genotypic array of the progeny produced by the cross between these two populations [GEA(PXQ)] is:
, then the genotypic array of the progeny produced by the cross between these two populations [GEA(PXQ)] is:
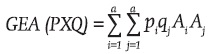
The initial number of candidate NIBD genes to be a part of the genotype of a hybrid parent of the SV is the number of lines which form it because they are inbred and unrelated. The number of NIBD genes lost by each parent of the SV was considered as the difference between the initial number and the average or expected number of NIBD genes carried by the genotypes of the m individuals representing this parent. If the number of NIBD genes which on average reach the m representatives of a DC and a TW are symbolized as E(Dm) and E(Tm), respectively, and the respective number of lost genes is expressed as (NIBD)pD and (NIBD)pT, then:

and

The m individuals representing a parent of the SV were considered as a random sample taken with replacement from the population of the genotypes forming the genotypic array of this parent (Sahagún et al., 2013).
Deriving results
]]> Double-cross hybridsThe formation of single crosses (SCs) such as (AA) X (BB) and (CC) X (DD) was considered for each of the L/4 four- line subsets. In addition, the genotypic array of the DC produced by crossing these two SCs [(GEA)DC] is as follows:

Each genotype comprises two NIBD genes in Equation 3. Therefore, if m = 1 the number of NIBD genes in the genotype sampled must be two. However, if m = 2 the number of NIBD genes in both genotypes of the sample is a variable because when the sample captures the same genotype twice the number of NIBD genes is two, but when the genotypes captured are different, the number of NIBD genes carried can be three (with AC and AD, for example) or four (with AC and BD, or with AD and BC).
According to Equation 3, the sampling results, necessary to derive the mean of the number of NIBD genes in the m genotypes comprising the sample of a double cross, are:
1) The same genotype m times. This result can occur in four different ways, but always with the contribution of two NIBD genes.
2) Two different genotypes that contain three NIBD genes. The pairs of possible genotypes are: (AC and AD), (AC and BC), (AD and BD) and (BC and BD). To find the number of ways in which any of these pairs can occur, for example AC and AD, consider that if one of them (e.g. AC) occurs n times, the other (AD) must occur the remaining m-n times (n = 1, 2, 3,..., m – 1). The number of forms of occurrence of AC n times is the number of combinations
. Moreover, as AC can appear in the sample 1, 2, 3,..., and up to m – 1 times, the number of times in which AC and AD occurs in the sample (NAC'AD) is as follows:
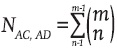
Since
]]>
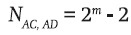
Thus, since each of the four pairs of genotypes carries three NIBD genes, the total number of ways in which two genotypes carrying three NIBD genes can occur in the sample is four (2m – 2).
3) Of the four genotypes of the double cross (Equation 3), six different pairs can be formed: the four mentioned in the previous paragraph and the remaining two: (AC and BD) and (AD and BC). Each one of the last two carries four NIBD genes. In addition, in the case of four pairs of two genotypes with three NIBD genes, the number of ways in which the remaining two pairs can occur is 2(2m – 2).
4) To complete the derivation it should be considered that the total number of equally possible and mutually exclusive outcomes produced by the random sampling of size m with replacement is 4m. Of these, those not yet discussed are 4m – 4(2m – 2) – 2(2m – 2) – 4, each of which corresponds to a sample of genotypes carrying four NIBD genes.
Considering the four possible results of the sampling, the mean of NIBD genes in samples of size m [E(Dm)] is:

Since each double cross is based on four lines which together carry four NIBD genes, the number of NIBD genes lost by every double cross [(NIBD)pD], according to Equations 1 and 4, is:
]]>
Equation 5 means that in the extreme case where m = 1 two NIBD genes are always lost. This is because a genotype of a single cross is formed by two NIBD genes, and the formation of the genotype of an individual generated by the double cross also requires two NIBD genes; of the four NIBD genes from this DC, the remaining two will always be lost. When m = 2, randomness becomes a factor regarding the number of NIBD genes. Even though one gene on average is lost (Equation 5), two genes are lost when the sample "captures" the same genotype both times.
Regarding the genetic erosion of each of the L/2 single crosses that are parents of a synthetic variety, Sahagún et al. (2013) found no loss of NIBD genes. Instead, according to Equation 5 in the case of (L/4) DCs a total loss of (L/4)(4/2m) = L(1/2)m NIBD genes is expected. This means that in terms of the cost of forming a synthetic with L/4 double crosses instead of one with L lines the loss of L(1/2)m NIBD genes should be included.
Given the information used to derive the mean of the number of NIBD genes of the m representatives of the double-cross parents of a SV (Equation 4), the variance [Var(Dm)] is:

The most evident is: with m = 1 there is no variability in the number of NIBD genes in the sample; consistent with the previous observation, according to which if m = 1, the number of NIBD genes is always two. Moreover, although when m is great the variance is zero, it is noteworthy that the greatest variance occurs when m = 2.
Thee-way line hybrids
With three inbred lines with genotypes AA, BB and CC, a three-way line hybrid can be, without loss of generality, (AAxBB)xCC. This cross generates the progeny whose genotypic array [(GEA)TL] is:

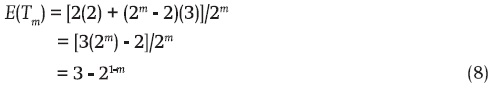
Furthermore, in terms of Equation 8, the variance of the number of NIBD genes in the sample [Var(Tm)] is:

Evidently then, E(Tm), the mean of the number of NIBD genes in the sample (Equation 8), is always equal to two when m = 1. In this case there is no variability, which is consistent with the value equal to zero of the variance (Equation 9). If m = 2, 3, 4,..., the respective average numbers of NIBD genes captured by the samples are: 2.5, 2.75, 2.875,..., respectively (Equation 8). For their part, the variances of the number of NIBD genes for these sample sizes are, respectively: 1/4, 3/16, 7/64,... These series, and Equations 8 and 9, mean that when the sample size is large the mean and variance of the number of NIBD genes in the sample representing a three-way line hybrid are 3 and 0, respectively.
Regarding the number of NIBD genes lost per each three-way line hybrid [(NIBD)pT], according to the previous result and Equation 8, we have the following:

In Equation 10, losses of NIBD genes of each three- way line hybrid are 1, 1/2, 1/4, ..., when m = 1, 2, 3, ..., respectively. These losses are half of those occurring in a double cross (Equation 5).
DISCUSSION
]]> To compare gene losses of double-cross and three-way line hybrids, consider that when m = 1, 2, 3,..., in a double-cross hybrid, 2,1,1/2,..., NIBD genes (Equation 5), respectively, are lost, while from a three-way line hybrid, NIBD gene losses (Equation 10) are, in the same order, 1, 1/2, 1/4,.... According to these results and Equations 5 and 10, the number of NIBD genes lost with a double cross is twice that lost with a three-way line hybrid. This is because, whereas in double crosses each of the two parental single crosses (due to their heterozygous condition) is a source of NIBD gene loss in the formation of m representatives, in the case of a three-way line hybrid only one parent has those characteristics; the other, an inbreed line, has no possibility of contributing to the loss of NIBD genes since all the gametes it produces have the same gene.The behavior of the variances of the number of NIBD genes in the samples of double-cross and three-way line hybrids (Equations 6 and 9) is, in general, very similar. When m = 1 both a double-cross and a three- way line hybrid always produce a genotype formed by two NIBD genes; that is, in both cases the variance is zero. With m = 2, however, both variances reach their peak as evidenced very clearly by Equations 6 and 9. From m = 3 the variances begin their descent to zero, faster in the case of three-way line hybrids. From m = 12, stability is almost reached and NIBD gene loss is about zero (Equations 5 and 10). These results are consistent with those of Butrón, Tarrío, Revilla, Malvar, and Ordás (2003) who, in order to save time and effort in developing a maize synthetic, proposed the formation of DCs and, based on a molecular evaluation, concluded that the method is appropriate, as it assures that the lines have equal contribution. They used m = 40.
According to Equations 6 and 9, the variance of the NIBD genes of a double cross is twice that of a TW. This is because the sources of variation of the number of NIBD genes are the two parental single crosses of each double cross and that of each TW. Every single cross produces the same variability and these sources of variability are independent.
Regarding inbreeding, which is inversely related to the genotypic mean, Sahagún-Castellanos et al. (2013) found that synthetics made with mixtures of lines and single crosses have inbreeding coefficients greater than those of synthetics whose parents are either lines or single crosses; this is due to the fact that while in these two cases the gene frequencies are balanced, this does not happen in the mixtures. In a TW there is also an imbalance of gene frequencies, and although there is no imbalance in the case of double crosses, they suffer the greatest loss of NIBD genes, which is directly related to inbreeding (Sahagún-Castellanos & Villanueva-Verduzco, 2012), and this with a reduction in the genotypic mean (Busbice, 1970). The inbreeding coefficients of the SVs derived from three- way line hybrids reported by Sahagún-Castellanos and Villanueva-Verduzco (2012) are slightly lower than those of the SVs of the four three-way line hybrids calculated with the formula reported by Márquez-Sánchez (2010). These numbers of double and single crosses, due to the line numbers involved, are consistent with those expected to be optimal for the derivation of a synthetic (Kutka & Smith, 2007).
It should be considered that the derivation of a synthetic variety is not over until the formation of the representatives of each parent, and that the rest of the process may involve an additional loss of NIBD genes. In individual representatives of parents, when these are hybrids, a necessary condition always occurs for gene loss: presence of heterozygous genotypes. However, when synthetic parents are inbred lines, there can be no loss of genes because each of these lines always transmits the same gene to their descendants.
CONCLUSIONS
The loss of non-identical by descent (NIBD) genes, which occurs during the derivation of the m individuals which will represent each parental DC of a synthetic variety, is twice that of one TW. The peak loss occurs in both cases when m = 1; in this case each DC always loses two NIBD genes and a TW loses one. Also, the loss of a DC is always equal to twice the loss of a TW, because the only sources of erosion are the SCs; two in one DC and one in a TW. However, when m = 12, the losses in both cases are approximately equal to zero. This means that in practice the risk of the genetic erosion discussed in this study should not be a problem with large numbers of m.
LITERATURE CITATION
]]>Busbice, T. H. (1970). Predicting yield of synthetic varieties. Crop science, 10(3), 265-269. doi:10.2135/cropsci1970.0011183X001000030017x [ Links ]
Butrón, A., Tarrío, R., Revilla, P., Malvar, R., & Ordás, A. (2003). Molecular evaluation of two methods for developing maize synthetic varieties. Molecular Breeding, 12(4), 329-333. doi: 10.1023/B:MOLB.0000006718.11324.4f [ Links ]
Kutka, F. J., & Smith, M. E. (2007). How many parents give the highest yield in predicted synthetic and composite populations of maize? Crop science, 47(5), 1905-1913. doi: 10.2135/cropsci2006.12.0802sc [ Links ]
Márquez-Sánchez, F. (2008). Endogamia y predicción de sintéticos de maíz de cruzas dobles. Revista Fitotecnia Mexicana, 31(3), 1-4. Recuperado de http://www.redalyc.org/articulo.oa?id=61009701 [ Links ]
Márquez-Sánchez, F. (2010). Inbreeding coefficient and mean prediction of maize synthetics of three-way lines hybrids. Maydica, 55(3), 227-229. Recuperado de http://www.maydica.org/articles/55_227.pdf [ Links ]
Sahagún-Castellanos, J., Rodríguez-Pérez, J. E., & Escalante-González, J. L. (2013). Yield prediction and inbreeding of maize synthetics generated with lines and single crosses. Classic probability. Revista de la Facultad de Ciencias Agrarias. Universidad Nacional de Cuyo, 45(2), 75-84. Recuperado de http://revista.fca.uncu.edu.ar/images/stories/pdfs/2013-02/T45_2_06_Sahagun-Castellanos.pdf [ Links ]
Sahagún-Castellanos, J., Rodríguez-Pérez, J. E., & Peña-Lomeli, A. (2005). Predicting yield of synthetics derived from double crosses. Maydica, 50(2), 129-136. Recuperado de http://www.maydica.org/articles/50_129.pdf [ Links ]
Sahagún-Castellanos, J., & Rodríguez-Pérez, J. E. (2011). Inbreeding of synthetic varieties derived from lines and single crosses. Revista Chapingo Serie Horticultura, 17(3), 107-115. doi:10.5154/r.rchsh.2011.17.022 [ Links ]
Sahagún-Castellanos, J., & Villanueva-Verduzco, C. (1997). Teoría de las variedades sintéticas formadas con híbridos de cruza simple. Revista Fitotecnia Mexicana, 20 (1). [ Links ]
]]>Sahagún-Castellanos, J., & Villanueva-Verduzco, C. (2012). ¿Variedades sintéticas derivadas de cruzas simples o de cruzas dobles? Revista Chapingo Serie Horticultura, 18(3), 279-289. doi: 10.5154/r.rchsh.2012.06.031 [ Links ] ]]>