
Phylogenetic analysis of the envelope protein (domain lll) of dengue 4 viruses
Javier Mota, MSc,(1) José Ramos-Castañeda, MSc,(1) Rebeca Rico-Hesse, PhD,(2) Celso Ramos, PhD.(1)
Mota J, Ramos-Castañeda J, Rico-Hesse R, Ramos C.
Phylogenetic analysis of the envelope protein (domain lll) of dengue 4 viruses.
Salud Publica Mex 2002;44:228-236. ]]>
Abstract
Objective. To evaluate the genetic variability of domain III of envelope (E) protein and to estimate phylogenetic relationships of dengue 4 (Den-4) viruses isolated in Mexico and from other endemic areas of the world. Material and Methods. A phylogenetic study of domain III of envelope (E) protein of Den-4 viruses was conducted in 1998 using virus strains from Mexico and other parts of the world, isolated in different years. Specific primers were used to amplify by RT-PCR the domain III and to obtain nucleotide sequence. Based on nucleotide and deduced aminoacid sequence, genetic variability was estimated and a phylogenetic tree was generated. To make an easy genetic analysis of domain III region, a Restriction Fragment Length Polymorphism (RFLP) assay was performed, using six restriction enzymes. Results. Study results demonstrate that nucleotide and aminoacid sequence analysis of domain III are similar to those reported from the complete E protein gene. Based on the RFLP analysis of domain III using the restriction enzymes Nla III, Dde I and Cfo I, Den-4 viruses included in this study were clustered into genotypes 1 and 2 previously reported. Conclusions. Study results suggest that domain III may be used as a genetic marker for phylogenetic and molecular epidemiology studies of dengue viruses. The English version of this paper is available too at: http://www.insp.mx/salud/index.html
Key words: Den-4 virus; phylogenetic analysis; envelope protein
Mota J, Ramos-Castañeda J, Rico-Hesse R, Ramos C.
Análisis filogenético del dominio III de la proteína de envoltura del virus dengue 4.
Salud Publica Mex 2002;44:228-236.
El texto completo en inglés de este artículo también está disponible en: http://www.insp.mx/salud/index.html
Resumen ]]>
Objetivo. Evaluar la variabilidad genética del dominio III de la proteína de envoltura (E) y estimar la relación filogenética de los virus dengue 4 (Den-4) aislados en México y en otras regiones endémicas del mundo. Material y métodos. En el presente trabajo reportamos un estudio filogenético del dominio III de la proteína de envoltura (E) que se realizó en 1998 con virus Den-4 aislados en distintos años en México y en otras partes del mundo. Se usaron oligonucleótidos específicos para amplificar por RT-PCR la región del dominio III y para obtener la secuencia de nucleótidos. Mediante el análisis de la secuencia de nucleótidos y de la secuencia deducida de aminoácidos se estimó la variabilidad genética y se generó un árbol filogenético. Para facilitar el análisis genético del dominio III se usó la técnica basada en el polimorfismo de fragmentos generados con enzimas de restricción (PFER) utilizando seis enzimas de restricción. Resultados. Los datos demuestran que la información del análisis de la secuencia de nucleótidos y de aminoácidos de la región del dominio III es similar a la del gene completo de la proteína E. El análisis de PFER con las enzimas de restricción Nla III, Dde I y Cfo I, mostró que los virus Den-4 incluidos en este estudio se agruparon en los genotipos 1 y 2 reportados previamente. Conclusiones. Los resultados sugieren que el dominio III se puede utilizar como un marcador para estudios filogenéticos y de epidemiología molecular del virus Den-4. El texto completo en inglés de este artículo también está disponible en: http://www.insp.mx/salud/index.html
Palabras clave: virus Den-4; análisis filogenético; proteína E
Dengue viruses cause large epidemics of dengue fever (DF) and Dengue Hemorrhagic Fever/Dengue Shock Syndrome (DHF/DSS) in Asia, the Pacific Region, Africa and America.1 During the period of 1970-1994, dengue-1, 2 and 4 circulated in the American Region and dengue-3 recently reappeared in Central America and Mexico.2-4 Dengue-4 (Den-4) was introduced in the Americas in 1981 and subsequently spread throughout the Region.3,5 In Mexico, the four serotypes of dengue have been isolated during epidemics of DF and DHF. Even when it is known that Den-4 viruses have been associated with DF cases in Mexico,5 Den-4 has also been implicated in DHF epidemics in 1984,6 and recently it has been detected in the brain tissue of a DHF fatal case.7
The antigenic and genetic variability of dengue viruses has been reported by several groups.8-15 The analysis of the flavivirus genome provides information about virus evolution and the complex interactions between viruses and biological factors.16 Phylogenetic studies on Den-2 suggested that the recent introduction of the Asian genotype into the Americas, is associated with an increased risk for DHF;17 therefore, phylogenetic analysis of dengue viruses can be a useful tool for epidemiological surveillance and for the development of efficient vaccines against dengue.17,18 Moreover, a good example of the information provided by this type of analysis is the recent observation reported by Leitmeyer et al19 and Watts et al20 in which the analysis of dengue- 2 genotypes isolated from patients with dengue fever and DHF, suggested a correlation between the severity of the disease and the infectious genotype. However, the most important limitation of this type of analysis is the time-consuming nucleotide sequencing procedure. Nevertheless, Vorndam et al21 recognized Den-2 and Den-3 subgroups by comparing Restriction Fragment Length Polymorphism (RFLP) patterns, which corresponded to those previously determined by oligonucleotide fingerprinting or genome sequencing,10,11,22 and pointed out that RFLP differentiates the major genotypes of dengue and provides a simple and rapid method for monitoring the relationship of dengue virus populations.
Phylogenetic patterns of Den-4 viruses have been studied using nucleotide sequences from the entire E gene.15 In an attempt to improve the phylogenetic analysis of dengue viruses, we focused on the analysis of domain III of the E protein of Den-4 virus; since this region is presumably under immune selective pressure,23 we would expect higher rates of nucleotide substitution and probably reflects better evolutionary and epidemiological relationships. Domain III of E protein has been recently implicated in the emergence of urban dengue due to the interaction of this domain with cellular receptors for virus entry; Wang et al reported that six of the eight amino acid changes are in domain III.16 In fact, some experimental evidence shows that domain III is the most variable region of protein E of mosquito-borne flavivirus.24 In the present work we performed a phylogenetic and a RFLP approach for rapid molecular analysis using domain III of the E gene, which could provide a tool for the analysis of evolutionary relationships between Den-4 viruses without having to determine the sequence of the entire E gene.
Material and Methods
Viruses ]]>
Dengue viruses were isolated from human serum in the mosquito cell line C6/36 and serotyped by indirect immunofluorescence using monoclonal antibodies.25 Selected Den-4 isolates from dengue fever cases in Mexico and six from other parts of the world, were included (Table I). Strain 0028, isolated in Mexico in 1984 from a DHF case, and Den-4 prototype virus (strain H241), were also included. Isolates used for phylogenetic and RFLP analysis were three from Mexico: Guerrero (strain 0153), San Luis Potosi (strain SLP-01) and Puebla (strain 0047)), and six from other parts of the world: Senegal (strain DAKHD34460), Venezuela (strain 88609), China (strain BN-L8 TVO259), Malaysia (strain LN-72992), Dominican Republic (strain TVP2177), and India (strain 611319 TVP2395)).

RT-PCR assay
The oligonucleotides used as primers were designed using the Oligo 4.0 computer software (National Biosciences, Playmouth MN, USA), based on the published nucleotide sequence of Den-4 (strain 814669) from Dominica (GenBank Accession No. M14931). Total RNA was extracted from infected cells with guanidine thiocyanate and phenol-chloroform, using a previously described procedure.10 The RNA was resuspended in distilled water and the reverse transcriptase (RT) reaction was carried out at 37 ºC for 1 h in a reaction mixture (10 ml) containing 5 mM MgCl2, 500 mM KCl, 100 mM Tris-HCl, pH 8.3, 1U/ml of RNase inhibitor, 2.5 U/ml of MuLV reverse transcriptase, 400 mM of nucleotides (Perkin Elmer, Norwalk, CT) and 1.2 mM of the anti-sense primer DENEB-I (21-mer, 5'-GTTTCACCTAGAATGGCCATT-3'). One ml of RT reaction mixture was used for cDNA amplification by the Polymerase Chain Reaction (PCR). The PCR mixture reaction (100 ml) contained 500 mM KCl, 100 mM Tris-HCl, 200 mM of nucleotides, 1.5 mM MgCl2, 2.5 U DNA Polymerase (Amplitaq; Perkin Elmer, Norwalk CT) and 1.2 mM of anti-sense (DENEB-I) and sense primer (DENEB-II; 21-mer, 5'-AAGTGCAAAGTCCGTATGGAG-3'). PCR was performed by an initial DNA denaturation step at 94 oC followed by 35 cycles of denaturation (94 ºC, 1 min), annealing (56 ºC, 1 min) and extension (72 ºC, 3 min). A final extension step was done at 72 ºC for 10 min. DENEB-I and DENEB-II primers amplified a fragment of 404 bp, encompassing the domain III region. The primers were tested using RNA extracted from cells infected separately with the four serotypes, and a fragment of 404 bp was amplified by RT-PCR only for Den-4 virus (data not shown).
For cDNA sequencing
The PCR product (404 bp) was excised from agarose gel and purified using the Geneclean II kit (Bio 101, Inc) and used as a template in the fmol DNA Sequencing System (Promega Corporation) using the primers DENEB-I and DENEB-II, and  dATP (10 mCi/ml). The sequence of domain III (282 nucleotides) was obtained using overlapping data from forward and reverse primers and used for phylogenetic analysis.
dATP (10 mCi/ml). The sequence of domain III (282 nucleotides) was obtained using overlapping data from forward and reverse primers and used for phylogenetic analysis.
Phylogenetic analysis ]]>
Nucleotide and deduced amino acid sequences encoding domain III were obtained for 9 Den-4 viruses (Table I) and compared with 15 virus sequences from the GenBank database. Sequence alignments were performed using the Wisconsin Package of the Genetics Computer Group, Inc. Phylogenetic analysis was done using the PAUP (Phylogenetic Analysis Using Parsimony) program, with uniform character weights, using branch and bound and heuristic search algorithms for most parsimonious trees;26 sequences of serotypes 1, 2 and 3 were used to root the tree. The reliability of the inferred tree was estimated using the bootstrap method17,19 with 100 replications, as described by Felsenstein.27Restriction enzyme analysis
Based on available Den-4 virus sequences; a computer-based analysis using the GCG software (Wisconsin Package, Genetics Computer Group, Inc.) was performed to analyze polymorphism in domain III region of envelope protein and six restriction enzymes were chosen. PCR products were digested with the restriction enzymes Mae III, Alu I, Sac I, Nla III, Dde I and Cfo I. Digestion reactions were performed using 5-10 ml of PCR product, 2.5 ml of appropriate buffer and 3 U of restriction enzyme in a final volume of 25 ml. The reaction was performed at 37 ºC for 1 h for all enzymes, except for Mae III (55 ºC, 1 h). Digestion products were separated by electrophoresis in a 3% agarose gel, stained with ethidium bromide and visualized under ultraviolet (UV) light.
Results
For phylogenetic analysis, nucleotide sequences encompassing domain III (282 bp, 301-394 aa of E protein) of 24 Den-4 virus strains were compared; sequences for 15 viruses were obtained from the GenBank (accession numbers: U18425-U18428, U18430-U18436, U18438-U18440, U18442). To root the tree the homologous domain III sequences for serotypes 1, 2 and 3 were included: Nauru74 strain (M32904), New Guinea44 (M29095), and Philippines56 strain (L11423), respectively. Only 9 nucleotide sequences (Table I) of Den-4 viruses used in this study were included in the phylogenetic analysis. Three of them represent viral isolates from dengue outbreaks occurring in Mexico (strains: 0153/Guerrero87, 0047/Puebla91 and SLP-01/SLP95) and from other parts of the world (strains: DAKHD34460/Senegal88, 88609/Venezuela90, BN-L8TV0259/China90, LN72992/Malaysia90, TVP2177/Dominican Republic89 and 611319/India90).
Mexican virus samples included in the phylogenetic analysis are representative of DF outbreaks in time and space. The nucleotide sequences were submitted to the GenBank (accession numbers: AF051104 to AF051112).
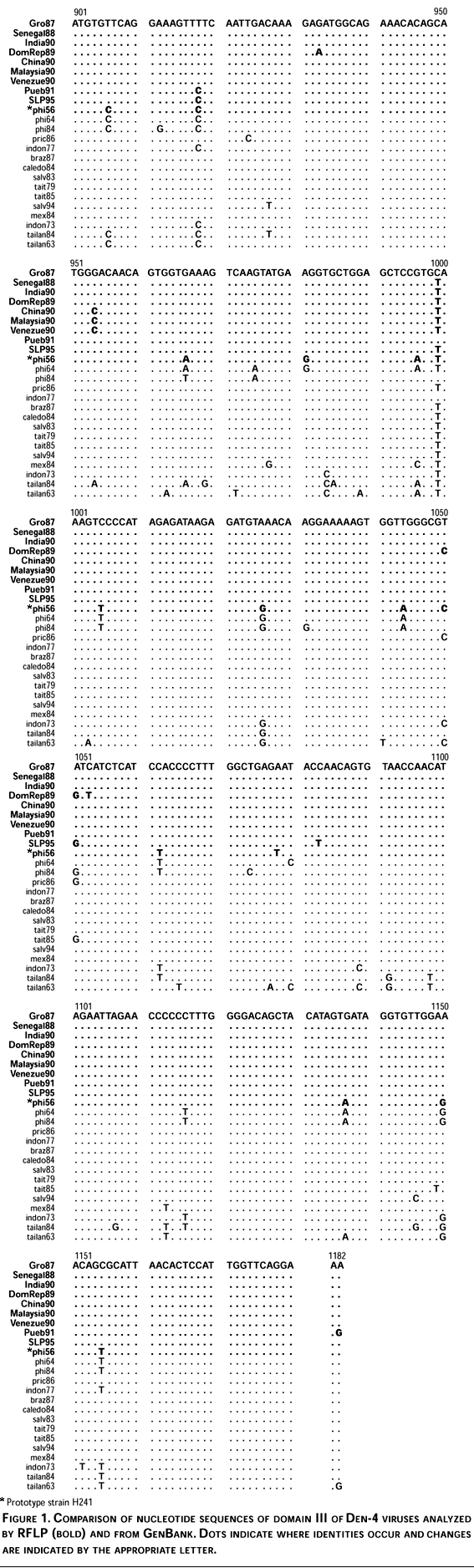
Nucleotide sequence comparisons among all Den-4 viruses included in this study showed a maximum divergence of 9.3% within Den-4 viruses. Nucleotide differences along domain III region (282 nt) occurred at 52 positions (18.4%) and most of them were silent. The region of maximum variability occurred at nucleotide sequence 961-1080 with 29 of the 52 changes, 55.8% (Figure 1). The deduced aminoacid sequence of domain III analysis showed eight changes (8.6%) and occurred at positions: Cys-Tyr,326 Thr-Ala,329 Glu-Gln,330 ILe-Val,335 Arg-Lys,344 Val-Ile,351 Tyr-Asn360 and Asp-Asn384 of E protein; most of these non-silent changes occurred in the above mentioned region (961-1080 nt).
]]>
Genetic variability and phylogenetic relationships among sequenced viruses (Table I) and virus sequences obtained from GenBank, were estimated and graphically represented using the PAUP program (Figure 2). The resulting phylogenetic tree showed two distinct groups: one including viruses only from Southeast Asia (Philippines 1956,1964,1984; Thailand 1963,1984), and the other including viruses from Asian countries, Africa, Central America, and South America.
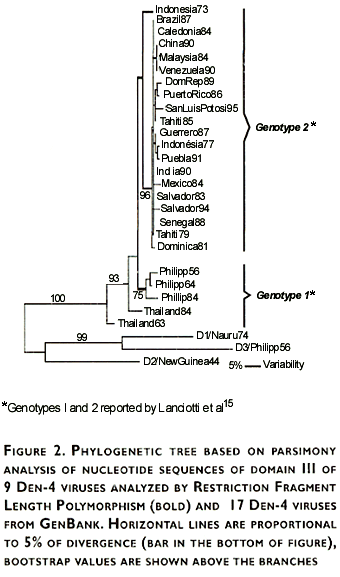
Even the short size of domain III and low variability of E gene of Den-4 viruses in comparison with the other serotypes15, phylogenetic analysis of Den-4 using sequences of domain III provides information that makes it a suitable target for molecular analysis using a different methodology instead of nucleotide sequencing. Our rationale was that domain III of Den-4 viruses contains enough variability, therefore genotypes could be evidenced by RFLP analysis. As described in materials and methods, a computer-based analysis for all restriction sites on domain III was performed and six enzymes were chosen for RFLP analysis (Mae III, Alu I,Sac I, Nla III, Dde I and Cfo I).
Total RNA of all strains listed in Table I was extracted from serum or cell lysates and used for RT-PCR amplification and restriction enzyme digestion, however, representative RFLP results for only nine isolates are presented. A similar pattern was obtained in all samples treated with Mae III, Alu I and Sac I (data not shown). However, for the Nla III restriction enzyme, the expected sizes (305 nt and 99 nt) were observed in all samples, except for Den-4 prototype H241 strain (Philippines, 1956) (Figure 3a). In the case of enzyme Dde I, all samples digested, except the prototype, showed the same fragments (223 nt and 181 nt) (Figure 3b). To confirm experimental results, nucleotide sequence of strain H241 (GenBank accession no. L11423) was analyzed for all possible Nla III and Dde I sites. Analysis showed 3 sites for Nla III (expected fragment sizes of 259, 85, 46 and 14 nt) that agree with our results. For Dde I two sites were found (expected fragment sizes of 181, 167 and 56 nt), as in the Nla III case, the expected sizes agree with our results. Finally, the digestion with Cfo I showed the expected fragments (304 nt and 100 nt), except for the prototype H241 and Dominican Republic strain (Figure 3c). Both prototype H241 and Dominican Republic viruses showed 3 different size fragments between them (197, 176 and 31 nt/ 197, 107 and 100 nt, respectively). This finding was confirmed by sequence analysis of PCR fragments; The Puerto Rican strain of Den-4 (GenBank accession no. U18436) had a similar pattern with Cfo I when it was analyzed by a computer program. In some cases, the expected small fragments (e.g. the 31 nt fragment for prototype H241 digested with Cfo I) were not visualized in the agarose gel, however the restriction enzyme cleavage sites were confirmed by nucleotide sequence analysis.



Discussion
Genetic analysis of dengue viruses has been a useful tool for monitoring the transmission of virus genotypes associated with the appearance and/or the increase of severe cases of the disease.17 Sequence variation and phylogenetic relationships among Den-4 viruses was first reported by Chungue et al,14 showing a maximum divergence of 4.9% in nucleotide sequences of the 5' terminus of the nucleotides encoding the E protein; however, genotypes could not be distinguished. More recently, Lanciotti et al15 reported a maximum nucleotide divergence of about 8% when E gene of Den-4 viruses from diverse geographic regions were compared, leading to their clustering in two genotypes.
In order to analyze a genome region that can provide phylogenetic information for Den-4 viruses to differentiate genotypes, we used the domain III of the envelope protein. Our results showed a maximum divergence of 9.3 % that is greater than the 8% reported by Lanciotti et al,15 considering that domain III region is less that 20% of the complete E gene. Our results support those reported by Gritsun et al,24 who suggested that variability in envelope protein of flaviviruses is not randomly distributed along the primary sequence but is characterized by distinct variable domains. This is a significant finding because the E protein is the target for neutralizing antibodies and T-cell responses and recently domain III of E protein has been identified as a key region during evolution of dengue viruses and its adaptation to mosquitoes and human hosts,16 since several aminoacid changes involved in this evolution and adaptation process are mapped into domain III. ]]>
The phylogenetic tree generated using the domain III sequences of Den-4 viruses (282 nt) showed two genetic groups; all Den-4 virus included in this work except prototype H241 strain (genotype 1), were clustered in genotype 2, according to Lanciotti et al.15 These results indicated that a shorter sequence than the entire E gene provide phylogenetic information, which allows to distinguish Den-4 viruses genotypes 1 and 2.Sequencing of viral genes is a time-consuming and expensive methodology that is impractical for the analysis of a large number of samples. In the present work we analyzed domain III of E protein by the RFLP approach using 6 different restriction enzymes. Den-4 viruses from Mexico and those from other parts of the world (Senegal88, Venezuela90, China90, Malaysia90, Dominican Republic89 and India90), showed similar restriction patterns, except for Den-4 prototype H241 (Philippines56) and Dominican Republic89 virus. The prototype H241 virus had a different restriction pattern with 3 out of 6 enzymes (Nla III, Dde I and Cfo I) and the Dominican Republic strain showed a different pattern only with Cfo I, which differs from that of prototype H241 (Figure 3). Although RFLP has some limitations in its interpretation, since results do not directly correlate with more precise sequence analysis, these results allowed us to rapidly classify viruses into genotypes that correlate with classifications suggested by others.21 This approach has been used in other viral studies, to quickly address epidemiological questions.28,29 Using information obtained from restriction enzyme patterns of Nla III, Dde I and Cfo I, we were able to cluster Den-4 viruses into genotypes 1 and 2;15 however, more virus isolates from genotype 1 of Den-4 are required to support the usefulness of the RFLP analysis. In spite of few viruses analyzed belonging to the Den-4 strains of genotype 1 with the enzymes described above, a computer-based analysis of the available GeneBank sequences of genotype 1 viruses, demonstrated that the same restriction sites were present as in the prototype strain H241 analized here (data not shown).
To assess the relevance of the presence of both Den-4 genotypes and their possible association with severe manifestations of the disease, large molecular epidemiological studies are needed. However, the lack of epidemiological studies of Den-4 epidemics in Mexico makes it difficult to have a clear distribution of both genotypes, therefore this work provides an alternative methodology for Den-4 virus analysis.
The transmission and human infection by multiple dengue virus serotypes increases the risk for severe disease and recently it has been shown that genotypes of the same serotype could play an important role in the outcome of the disease.17,20 Since different environmental factors seem to increase the risk of introducing new dengue virus genotypes into areas where they did not previously exist, it is important to screen multiple samples rapidly, to detect their introduction into endemic areas. The RFLP analysis described here could be an important research tool to improve the detection of genetic variants of Den-4 virus. This information could also be used to correlate the transmission of specific virus variants and the risk for severe dengue outbreaks.
Acknowledgements
We thank Dr. Carlos Arias and Dr. Miguel Angel Torres from the Instituto de Biotecnología/Universidad Nacional Autónoma de México, for their support in nucleotide sequencing.
References
1. Gubler DJ, Clark GG. Dengue/dengue haemorrhagic fever: The emergence of a global health problem. Emerg Infect Dis 1995;1:55-57. [ Links ]
2. Monath TP. Dengue: The risk to developed and developing countries. Proc Nat Acad Sci USA 1994;91:2395-2400. [ Links ]
3. Gubler DJ, Trent DW. Emergence of epidemic dengue/dengue hemorrhagic fever as a public health problem in the Americas. Infect Agents Dis 1994;2:383-393. [ Links ]
4. Briceño-García B, Gómez-Dantés H, Argott-Ramírez E, Montesano R, Vázquez-Martínez AL, Ibáñez-Bernal S et al. Potential risk for dengue hemorrhagic fever : The isolation of serotype dengue-3 in Mexico. Emerg Infect Dis 1996;2:133-135. [ Links ]
5. Monath TP, Heinz FX. Flaviviruses. in: Virology. 3rd Ed. Fields B N, Knipe DM, Holay PM, ed. Filadelfia Lippincott-Raven (PA)1996: 1002. [ Links ]
6. Loroño PM, Farfán AJ, Rosado PE, Kuno G, Gubler DJ. Epidemic of dengue in the Yucatán, México. Rev Inst Med Trop Sao Paulo 1993;35: 449-455. [ Links ]
7. Ramos C, Sánchez G, Hernández-Pando R, Baquera J, Hernández D, Mota J et al E. Dengue virus in the brain of a fatal case of hemorrhagic dengue fever. J Neurovirol 1998;4:465-468. [ Links ]
8. Henchal EA, Repik PM, McCown JM, Brandt WE. Identification of an antigenic and genetic variant of dengue-4 virus from the Caribbean. Am J Trop Med Hyg 1986;35:393-400. [ Links ]
9. Blok J, Samuel S, Gibbs AJ, Vitarana UT. Variation of the nucleotide and encoded aminoacid sequences of the envelope gene from eight dengue 2 viruses. Arch Virol 1989;105:39-53. [ Links ]
10. Rico-Hesse R. Molecular evolution and distribution of dengue viruses type 1 and 2 in nature. Virology 1990;174:001-0015. [ Links ]
11. Lewis JA, Chang JG, Lanciotti RS, Kinney RK, Mayer LW, Trent DW. Phylogenetic relationships of dengue-2 viruses. Virology 1993;197:216-224. [ Links ]
12. Chungue E, Cassar O, Drouet MT, Guzmán MG, Laille M, Rosen L et al. Molecular epidemiology of dengue 3 viruses and genetic relatedness among dengue 3 strains isolated from patients with mild or severe form of dengue fever in French Polynesia. J Gen Virol 1993;74:2765-2770. [ Links ]
13. Lanciotti RS, Lewis JG, Gubler DJ, Trent DW. Molecular evolution and epidemiology of dengue-3 viruses. J Virol 1994;75:65-75. [ Links ]
14. Chungue E, Cassar O, Drouet MT, Guzmán MG, Laille M, Rosen L et al. Molecular epidemiology of dengue-1 and dengue-4 viruses. J Gen Virol 1995; 76:1877-1884. [ Links ]
15. Lanciotti RS, Gubler DJ, Trent DW. Molecular evolution and phylogeny of dengue-4 viruses. J Gen Virol 1997;78:2279-2286. [ Links ]
16. Wang E, Ni H, Xu R, Barrett AD, Watowich SJ, Gubler DJ et al. Evolutionary relationships of endemic/epidemic and selvatic dengue viruses. J Virol 2000;74:3227-3234. [ Links ]
17. Rico-Hesse R, Harrison LM, Salas RA, Tovar D, Nisalak A, Ramos C et al. Origins of dengue type 2 viruses associated with increased pathogenicity in the Americas. Virology 1997; 230:244-251. [ Links ]
18. Beaty BJ. Virus variation and evolution mechanisms and epidemiological significance. En: Monath TP, ed. The arboviruses: Epidemiology and ecology. Boca Raton (FL): CRC Press, 1988: 59-85. [ Links ]
19. Leitmeyer KC, Vaughn DW, Watts DM, Salas R, Chacón IV, Ramos C et al. Dengue virus structural differences that correlate with pathogenesis. J Virol 1999;73:4738-4747. [ Links ]
20. Watts DM, Porter KR, Putvatana P, Vásquez B, Calampa C, Hayes CG et al. Failure of secondary infection with American genotype dengue 2 to cause dengue haemorrhagic fever. Lancet 1999;354:1431-1434. [ Links ]
21. Vorndam V, Kuno G, Rosado N. A PCR-restriction enzyme technique for determining dengue virus subgroups within serotypes. J Virol Methods 1994; 48:237-244. [ Links ]
22. Trent DW, Grant JA, Rosen L, Monath TP. Genetic variation among dengue 2 viruses of different geographic origin. Virology 1983;128:271-284. [ Links ]
23. Rey FA, Heinz FX, Mandl C, Kunz C, Harrison SC. The envelope glycoprotein from tick-borne encephalitis virus at 2 Å resolution. Nature 1995;375:291-298. [ Links ]
24. Gritsun TS, Holmes EC, Gould E. Analysis of flavivirus envelope proteins reveals variable domains that reflect their antigenicity and may determine their pathogenesis. Virus Res 1995;35:307-321. [ Links ]
25.Gubler DJ, Kuno G, Sather GE, Vélez M, Oliver A. Mosquito cell cultures and specific monoclonal antibodies in surveillance for dengue viruses. Am J Trop Med Hyg 1984;33:158-165. [ Links ]
26. Swofford D. AUP: Phylogenetic analysis using parsimony. Version 3.1.1 Champaign (IL): Illinois Natural History Survey, 1993. [ Links ]
27. Felsenstein J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985;39:783-791. [ Links ]
28. Vonsover A, Hansher R, Neuman M, Guillot S, Balanant J, Rudich H et al. Molecular epidemiology of type 1 Polioviruses isolated in Israel and defined by restriction fragment length polymorphism assay. J Infect Dis 1993; 167:199-203. [ Links ]
29. Wu JC, Chiang TY, Sheen IJ. Characterization and phylogenetic analysis of a novel hepatitis D virus strain discovered by restriction fragment length polymorphism analysis. J Gen Virol 1998;79:1105-1113. [ Links ]
]]> (1) Departamento de Arbovirus, Centro de Investigación sobre Enfermedades Infecciosas, Instituto Nacional de Salud Pública, Cuernavaca, Morelos, México.
(2) Department of Virology and Immunology, Southwest Foundation for Biomedical Research, San Antonio, Texas 78245, USA.
Received on: April 17, 2001 · Accepted on: October 30, 2001
Address reprint requests to: Dr. Celso Ramos. Area de Enfermedades Transmitidas por Vectores, CISEI/ INSP. Av. Universidad No. 655, Colonia Santa María Ahuacatitlán, 62508 Cuernavaca, Morelos, México.
E-mail: cramos@correo.insp.mx ]]>