Grandes alcances de los RNAs pequeños RNA de interferencia y microRNA
Great potential of small RNAs: RNA interference and microRNA
Guelaguetza Vázquez–Ortiz,* Patricia Piña–Sánchez,* Mauricio Salcedo*
* Laboratorio de Oncología Genómica. Unidad de Investigación Médica en Enfermedades Oncológicas. Hospital de Oncología. Centro Médico Nacional Siglo XXI–IMSS.
]]> Reimpresos:
Recibido el 28 de septiembre de 2005. ]]> Aceptado el 7 de marzo de 2006.
ABSTRACT
Double–stranded RNA (dsRNA) induces a sequence–specific silencing in eukaryotic cells. This silencing process beggins when long dsRNA is cleaved to 21 to 26 long small RNA by means of the RNAse III–type enzyme Dicer. These small dsRNA are included into silencing effector complexes, that are targeted to complementary sequences. Small RNA dependent gene silencing can be achieved by distinct mechanisms based depending mainly on the nature of target sequences and on the proteins present in the effector complex. The route of interference RNA (RNAi) begins when Dicer yields small interference RNA (siR–NA) that bind to complementary mRNA for its degradation, forming the RISC complex. siRNA are naturally formed from transposons and dsRNA viruses during its replication, as well as from other bidirectional transcribed repetitive sequences. Some of the enzymes thar are part of the RNAi machinery, including Dicer, are encoded by multigene families in many species, that also play a role in other mechanisms of RND–dependent gene silencing. MicroRNA's (miRNA) are other small RNA's that can induce gene silencing at the mRNA level. These are formed in a general manner when Dicer process hairpin structures resulting from the transcription of non–coding sequences from plant and animal genomes. miRNA's are integrated into a RISC–like complex, after which, depending on their degree of complementarity with target mRNA, can either repress translation or induce mRNA degradation. miRNA–dependent silencing is essential for the development of multicellular organisms. Artificial RNAi induction by means of siRNA or miRNA is being used as a tool to inactivate gene expression in culture cells and in living organisms. This review focuses on the progress in the understanding of the mechanisms involved in gene regulation by RNA in animals and details some current efforts to apply theses phenomena as a tool in research and in the therapeutic of human diseases.
Key words. Double stranded RNA. Interference RNA. MicroRNA. Cancer.
RESUMEN
El RNA de doble cadena puede inducir un silenciamiento secuencia–específico en eucarionte. Este proceso de silenciamiento se inicia cuando el RNAdc largo es procesado a RNA pequeño de 21 a 26 nucleótidos mediante la enzima RNAsa III Dicer. Estos RNA pequeños se incorporan a complejos efectores de silenciamiento, que son guiados a secuencias complementarias blanco. Existen diferentes tipos de silenciamiento, cuyas diferencias se basan principalmente en la naturaleza de la secuencia blanco y en la composición proteica de los complejos efectores. La ruta del RNA de interferencia (RNAi) se inicia cuando Dicer genera pequeños RNA de interferencia (siRNA) que se unen por complementariedad al mRNA para su degradación, utilizando el complejo RISC. De manera natural, los siRNA se originan de transposones y virus que producen RNAdc durante su replicación, así como también de otras secuencias repetidas transcritas bidireccionalmente. Algunas de las enzimas que conforman la maquinaria del RNAi como Dicer, entre otras, son codificadas por familias multigénicas en varias especies y también participan en otros mecanismos de silenciamiento mediado por RNA. Los microRNA son otros RNA pequeños que pueden inducir silenciamiento al unirse al mRNA. Éstos se generan de manera general cuando Dicer procesa estructuras de horquilla compuestas de regiones no codificantes, en genomas de plantas y animales. Los miRNA se incorporan a un complejo similar a RISC y, dependiendo de su grado de complementariedad con el mRNA blanco, pueden tener represión traduccional o bien digerir el mRNA. El silenciamiento mediado por miRNA es esencial para el desarrollo de plantas y animales. La inducción artificial del RNAi mediante siRNA o miRNA ha sido adoptada como una herramienta para inactivar la expresión génica, tanto en células en cultivo como en organismos vivos. En esta revisión se muestra el gran progreso en el entendimiento de los mecanismos que participan en la regulación génica mediada por RNA en animales y detalla algunos esfuerzos actuales para encauzar a estos mecanismos como una herramienta en la investigación y como posible terapia en enfermedades.
Palabras clave. RNA doble cadena. RNA de interferencia. MicroRNA. Cáncer.
]]> INTRODUCCIÓN
El estudio de la regulación génica ha cobrado gran importancia; esto confirma que la expresión génica tiene influencia directa sobre una gran cantidad de actividades celulares. Aunque es conocida la actividad de las proteínas que intervienen en este proceso, cada día hay más evidencia de que cientos de transcritos pequeños pueden ser factores muy importantes en la regulación genética.1 Estas especies de transcritos pequeños (RNA) salieron a escena al final de la era genómica, como RNA de expresión ubicua y como represores extraordinariamente versátiles de la expresión génica en plantas, animales y hongos.2 Estos RNA diminutos, de aproximadamente 21–26 nt, dependen de la maquinaria del RNA de interferencia (RNAi) para efectuar su acción más conocida, es decir, el silenciamiento de la expresión génica al interactuar con secuencias complementarias.3 Estos RNA se clasifican como a) RNA pequeños de interferencia, siRNA (o "short interfering RNA", por su nombre en inglés) y b) microRNAs (miRNA). Ambos son componentes de un mecanismo de regulación de la expresión génica basado en transcritos, que funciona principalmente en eucariontes.4,5 Dentro de las nuevas líneas de investigación, estos RNA pequeños se están utilizando con dos propósitos fundamentales: 1) inhibir la actividad viral y de transposones para identificar nuevos métodos de defensa del hospedero contra estos microorganismos, y 2) el silenciamiento de la expresión de genes codificantes de diversas proteínas de interés funcional.6–8 Otros tipos de RNA pequeños poco estudiados son los RNA temporales pequeños (o stRNA por su nombre en inglés: "small temporal RNAs"), y siRNA heterocromáticos. Se ha observado que los siRNA heterocromáticos pueden intervenir en la organización cromosómica y en las modificaciones epigenéticas de regiones específicas dentro del genoma.9
Tomando en cuenta que recientemente el RNAi se ha convertido en uno de los métodos más utilizados para analizar a las células de los mamíferos, y que además tiene el potencial de servir como tratamiento terapéutico de algunas enfermedades esporádicas y hereditarias, en esta revisión se muestra la biogénesis, mecanismos de regulación genómica y aplicaciones de dos de las especies de RNA pequeños más estudiados hasta ahora: los siRNA y miRNA.
RNA DE DOBLE CADENA Y RNAi
La capacidad del RNA de doble cadena (RNAdc) de interferir en la expresión génica de algunos eucariontes es conocida desde hace varios años, ya que el RNA largo de doble cadena genera una respuesta de interferón inhibiendo la traducción de proteínas a través de la acción de PKR. Sin embargo, el silenciamiento de genes mediante RNA pequeño en eucariontes también se identificó desde hace aproximadamente 20 años, pero los mecanismos para llevar a cabo el silenciamiento están apenas siendo descritos. Este fenómeno recibió el nombre de RNA de interferencia (RNAi).10
Antes se pensaba que los RNAdc eran agentes silenciadores inespecíficos, incapaces de disociarse y aparearse específicamente y capaces de inducir destrucción general de distintos RNA mensajeros (RNAm) y, por lo tanto, la supresión total de la traducción de proteínas.
Uno de los primeros indicios de que el RNAi era un fenómeno biológico aún no descrito fue la potencia de su efecto. Guo y Kemphues, utilizaron una cadena complementaria del RNA mensajero del gen par–1 para bloquear su expresión en C. elegans mediante la técnica conocida como "silenciamiento por antisentido". Ésta depende de la liberación de grandes cantidades en el citoplasma del transcrito pequeño complementario al RNAm blanco (secuencia de interferencia o antisentido).11 Inicialmente se pensó que era un bloqueo pasivo en el proceso de traducción, debido al apareamiento de bases entre el RNAm de interés y el RNA de interferencia. Posteriormente se encontró que tanto la cadena sentido como la antisentido podían inducir silenciamiento del RNAm, lo cual hizo necesario replantear esta hipótesis funcional.12
En 1998 la ruta del RNAi fue reconocida en C. elegans. En este trabajo Fire y cois, sugirieron que el RNAdc, que frecuentemente está presente en las células durante una infección viral, podría ser el iniciador o el precursor de la ruta del RNAi. Dicho estudio pretendía silenciar el RNAm blanco en forma pasiva microinyectando simultáneamente concentraciones bajas de secuencias complementarias de RNA sentido y su antisentido, lo que al hibridar formó complejos de RNAdc.13,14 Dado que es posible producir y purificar in vitro RNA e introducirlo directamente en C. elegans y el silenciamiento es heredado a la progenie por vía germinal, entonces se comprueba que el RNAdc es un potente activador de la ruta del RNAi, y el silenciamiento es aproximadamente 100 veces más efectivo que el RNA de cadena sencilia.15,16
El experimento anterior permite utilizar este inductor en grandes poblaciones de animales como tratamiento, simplemente alimentándolos con bacterias que expresen RNAdc. Tanto en Drosophila,11 como en algunas plantas18 y hongos,19,20 de manera natural existen genes que silencian, ya sea postranscripcionalmente, por co–supresión o por RNAi, sugiriendo que cada uno de estos mecanismos tiene un origen ancestral común en eucariontes.
]]> Poder utilizar la ruta del RNAi como herramienta experimental para silenciar genes específicos en modelos de mamíferos ha sido una tarea muy difícil. El problema principal radicaba en hacer que el RNAdc exógeno silenciara genes específicos sin que activara la respuesta del interferón mediante PKR, que es parte del funcionamiento normal de la célula en respuesta a agentes infecciosos y/o virales. No fue hasta que se descubrió que el siRNA dúplex puede programar al complejo de silenciamiento de RISC (del que se habla en las siguientes secciones), para inactivar genes específicos. En mamíferos el descubrimiento de este último proceso fue muy importante debido a que los RNAdc de más de 30 pares de bases activan la respuesta del interferón, mientras que los siRNA dúplex resultantes de la acción del complejo RISC contienen aproximadamente 19 pb, lo que permite utilizar esta herramienta como una metodología estándar para silenciar la expresión de genes específicos en células de mamíferos.21La diferencia entre el tipo de respuesta que es generada por la PKR y el RNAi, radica en su respectiva especificidad; la respuesta de PKR inhibe la expresión de manera global, mientras que el RNAi tiene un efecto específico sobre la expresión de un gen o genes determinados.22 De manera natural, el silenciamiento por la ruta del RNAi se origina a partir de transposones, de virus que producen RNAdc durante su replicación, así como de algunas secuencias repetidas que se transcriben bidireccionalmente.23–25
La ruta del RNAi también puede ser inducida por la expresión endógena de horquillas pequeñas de RNA (shRNAs por sus siglas en inglés: "short hairpin RNAs"). Éstos tienen una estructura similar a una clase de RNA pequeños altamente conservados denominados microRNA (miRNA), los cuales median la ruta del RNAi a través de un mecanismo de inhibición de la traducción. Veremos esto a continuación.26,27
BIOGÉNESIS Y TRÁFICO DE miRNA
De acuerdo con un convenio internacional, un microRNA se define como un RNA de cadena sencilla de 18 a 24 nucleótidos de longitud que es generado por la enzima Dicer, una RNAsa de tipo III, a partir de un transcrito endógeno que contiene una estructura de horquilla. Al momento de escribir este artículo, la base de datos de miRNA (disponible en: http://www.sanger.ac.uk/Software/Rfam/mirna) contiene 2,116 secuencias, de las cuales existen varias específicas para C. elegans, D. melanogaster, Dardo renio, Gallus gallus, A. thaliana y virus de Epstein Barr, entre otras. Para Homo sapiens existen 638 secuencias.
Aunque la mayoría de los miRNA están aislados a lo largo del genoma, algunos se encuentran formando clusters que son transcritos y expresados de manera coordinada. Esto indica una organización de tipo operón para algunos miRNA. Los miRNA se localizan en exones e intrones de RNA no codificante, así como intrones de mRNA codificante en la misma orientación que los mRNA, lo cual sugiere que algunos miRNA se podrían originar a partir de transcritos de intrones de pre–mRNA que sufrieron escisión.28
Las enzimas RNAsa de tipo III son una familia de ribonucleasas que reconocen específicamente al RNAdc y que se cree que están presentes en todas las células vivas. Una característica importante de las RNAsas de tipo III es que al digerir el RNAdc introducen dos nucleótidos en el sitio de digestión.29–30
Las células humanas expresan tres miembros de esta familia. La expresión de una de estas proteínas está restringida a la mitocondria, mientras que las otras dos se encuentran en citosol y núcleo. Éstas se denominan Drosha y Dicer y ambas juegan papeles diferentes pero fundamentales en el procesamiento de los miRNA: Drosha participa en el procesamiento de los RNA humanos en el núcleo, mientras que Dicer genera casi exclusivamente miRNA y siRNA.
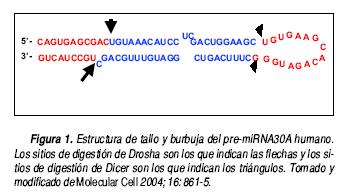
La biogénesis de los miRNA presenta varios pasos. Al inicio, los miRNA forman parte de una estructura de tallo y burbuja de aproximadamente 80 nt de longitud, que forma parte del pri–miRNA. Esta estructura es procesada en el núcleo por Drosha para liberar una horquilla más pequeña de aproximadamente 60 nt de longitud llamada pre–miRNA, la cual contiene un extremo cohesivo colgante. Aún no se determina la forma exacta en la cual Drosha discrimina estructuras de tallo y burbuja de pre–miRNA y posteriormente determina el sitio exacto de la digestión. (Figura 1).31
]]>
Esta enzima forma parte del complejo proteico de 600 kDa denominado "microprocesador". Otro de los componentes esenciales del microprocesador, además de Drosha, se denomina DGCR8 en humanos y Pasha en Drosophila. El microprocesador se presenta como un heterotetrámero y consiste principalmente en dos moléculas de Drosha y dos de DGCR8. La proteína DGCR8 contiene dos dominios de unión a RNAdc, sugiriendo que ésta puede jugar un papel importante en el reconocimiento o unión del complejo microprocesador.32
La Exportina 5 (Exp5) es un miembro de la familia de las karioferinas que son factores de transporte núcleo–citoplasma, éstos son dependientes de un co–factor denominado Ran (una GTPasa) para su funcionamiento. Exp5 forma un heterodímero nuclear con Ran–GTP y con los pre–miRNA que resultan del procesamiento de Drosha. De la misma manera que Drosha y el heterodímero Exp5/Ran–GTP se une a RNAs pequeños que tienen una hebra terminal de aproximadamente 16 bp y una cola cohesiva en el extremo 3', precisamente la estructura que presentan los pre–miRNA.33–37
PROCESAMIENTO CITOPLÁSMICO DE PRE–miRNA POR Dicer
En invertebrados, las enzimas relacionadas a Dicer humana juegan un papel muy importante en el RNAi, digiriendo RNAdc largos a RNA dúplex de aproximadamente 22 pb que contienen una cola cohesiva de 2 nt en 3'.38–41 Esta digestión es subsecuente, donde Dicer se une nuevamente a los extremos del RNAdc y vuelve a digerir aproximadamente 22 bases después del extremo hasta que el RNAdc queda totalmente digerido. Mientras que Dicer digiere al RNAdc que tiene extremos romos, el dominio PAZ de Dicer se une a los extremos de RNAdc que tiene colas cohesivas (como los generados por Dicer).42
Dicer también juega un papel importante en la generación de los miRNA maduros en vertebrados, donde el dominio PAZ de Dicer se une a la cola cohesiva presente en la base de la horquilla de pre–miRNA, y el dominio de unión a RNAdc de Dicer se une a la base de la burbuja y define la distancia de la digestión desde la base.43–46 El procesamiento por Dicer genera RNA dúplex de aproximadamente 22 pb, que contienen dos colas cohesivas en el extremo 3', lo cual es similar tanto en estructura como en función a los siR–NA dúplex que son generados por Dicer a partir de RNAdc largos. Después de la digestión, Dicer permanece asociada con el miRNA o siRNA dúplex intermediario de 22 pb y posteriormente es liberado por una helicasa (Figura 2).

COMPLEJOS EFECTORES
La digestión por Dicer es seguida por la liberación del miRNA maduro o del siRNA, y también por la incorporación al complejo de RISC ("RNA–induced silencing complex", por sus siglas en inglés) o al de RITS ("RNA–induced initiation of gene transcriptional gene silencing", por sus siglas en inglés), cuyas funciones incluyen digestión del mRNA, supresión de la traducción, silenciamiento transcripcional y formación de heterocromatina.47
Existe una variedad de componentes que auxilian en el ensamblaje de RISC, particularmente aquellos que despliegan al siRNA o al miRNA y a través de esta acción determinan cuál cadena de estos RNA dúplex es la que se activa para el silenciamiento.48
Los complejos de RISC se componen principalmente de ribonucleoproteínas que contienen a miembros de la familia de proteínas Argonauta (Ago) con dominios PAZ–Piwi.49 De manera adicional, RISC contiene varios factores accesorios cuyas acciones son necesarias para la función efectora. La mayoría de los organismos tienen homólogos de la familia de Argonauta, Drosophila tiene cinco, el ratón tiene ocho y los nemátodos tienen al menos veinticuatro. Estas proteínas dan especificidad al complejo RISC y determinan la respuesta específica para cada siRNA o miRNA. El dominio Piwi interacciona directamente e inhibe a la región del dominio de unión de RNa–seIII/dcRNA de Dicer.50 Esto se ha interpretado como una interacción de transferencia de sustrato y se ha especulado que las interacciones de Piwi con Dicer pueden estimular la liberación del miRNA o del siRNA. Algunos otros componentes adicionales de RISC han sido identificados y aún no se ha descrito su función en la ruta del RNAi. Dentro de éstos se encuentran la proteína de unión a RNA VIG, el homólogo en Drosophila de la proteína X Frágil, dFXR, helicasas y Tudor–SN.51–56
El ensamblaje de RISC es uno de los pasos más complejos en la ruta del miRNA y del RNAi. Involucra a pequeños productores de RNA pequeño (DCR), RNA dúplex pequeño, el desdoblamiento de RNA dúplex de simétrico a asimétrico y el reclutamiento de diferentes proteínas Ago. Cada una de las diferentes proteínas Ago de RISC tienen funciones diferentes que están posiblemente determinadas por los dominios Piwi de la proteína Ago.57–60 Algunos dominios Piwi confieren actividad "slicer", mientras que otros no.
Dependiendo del tipo de proteínas Ago reclutadas a RISC, estos complejos pueden ser tentativamente divididos en dos tipos: RISC digestivo y RISC no digestivo. Un RISC digestivo tiene una función dual que dirige tanto la digestión del mRNA blanco como la represión traduccional, dependiendo de las características del apareamiento entre el RNA pequeño y el RNAm. Si la complementariedad entre el apareamiento del siRNA o miRNA y su RNAm blanco no es al 100%, el RNAm es físicamente inalcanzable por el centro activo de la endonucleasa (slicer) del RISC digestivo, ya que se forma una horquilla entre el blanco y los miRNA o siRNA.61–63 Por lo tanto, esto genera una represión traduccional en lugar de una digestión eficiente del RNAm blanco. En contraste, los RISC no digestivos no tienen actividad de endonucleasa (slicer) en el dominio Piwi de las proteínas Ago y pueden utilizar el RNAm blanco sólo para la represión de la traducción.64–68 Además de estos dos tipos de funcionamiento de RISC, existen otros tipos denominados RITS.
CONTROL DE LA ESTRUCTURA DE LA CROMATINA POR EL COMPLEJO RITS
El efecto de las rutas de silenciamiento del RNA no sólo está limitado a procesos citoplásmicos como el apagado del mRNA y la síntesis de proteínas. El inicio de la formación de la cromatina, tanto en los centrómeros como en otros sitios cromosómicos, ha mostrado que depende del aparato de silenciamiento del RNA y también de los RNA pequeños. Independientemente de su papel sobre la estructura del cromosoma durante el crecimiento y la división celular, la formación de heterocromatina inducida por RNAdc también parece generar silenciamiento transcripcional en respuesta a la expresión de transgenes o transposones. Dependiendo del organismo y del contexto celular específico, los RNA pequeños que derivan de secuencias repetidas o de transgenes pueden metilar el DNA, o bien, las histonas. Este fenómeno ha sido descrito en protozoarios, plantas, insectos y vertebrados; sin embargo, donde se ha descrito de manera principal es en Schizosaccharomyces pombe.69,74 En esta levadura Agol, el único homólogo de Argonauta, es ensamblado dentro del complejo de RITS. Este complejo además se forma de Chpl, una proteína de unión a centrómero que contiene un cromo–dominio y que es necesaria para la metilación de la histona H3 lisina 9 en los repetidos del centrómero. Y también este complejo se conforma por Tas3, una proteína de función desconocida. El RITS utiliza siRNA provenientes de transcritos centroméricos repetidos para guiar su localización en estos repetidos e inducir la formación de heterocromatina. Independientemente del mecanismo, identificar un complejo efector que contenga Ago indica que éste pudiera ser un mecanismo paralelo a miRISC o siRISC.75–79
]]>ACTIVACIÓN DIRIGIDA DE LA RUTA DE SILENCIAMIENTO DEL RNAi: siRNA Y miRNA
La ruta de procesamiento del RNAi ilustrada en la Figura 3, está conservada en la mayoría de las células eucariontes y puede utilizarse para promover la entrada de siRNA exógenos o sintéticos. Como se mencionó, en muchos vertebrados los RNAdc de mayor longitud pueden activar la respuesta de interferón, por lo que su uso no permite silenciar genes de manera dirigida en estos organismos. Para silenciar genes específicos ha sido necesario realizar transfecciones con siRNA dúplex sintético, que es muy parecido al miRNA dúplex, haciendo que actúe RISC. Aunque estos experimentos resultaron en la destrucción específica y efectiva de los mRNA blanco, los siRNA sintéticos son muy caros y la transfección del RNA es poco eficiente en todas las células de interés. Para superar este problema se han utilizado promotores dependientes de RNA polimerasa, desarrollando plásmidos que puedan expresar pequeñas estructuras de RNA de horquilla, análogas estructuralmente a los pre–miRNA. A partir de esto, una serie de trabajos posteriores han refinado los parámetros para diseñar las estructuras pequeñas de RNA de horquilla para obtener silenciamiento exitoso por RNAi. Actualmente se sabe que las secuencias de 25 a 29 pb favorecen el procesamiento del RNA y la expresión de siRNA.33,80
También se han desarrollado vectores virales basados en el genoma del virus de leucemia murina o de inmunodeficiencia humana que incorporan casetes de expresión con pequeñas estructuras de horquilla. Estos vectores infectan a las células blanco in vivo e in vitro, generando expresión estable de los siRNA y supresión a largo plazo del mRNA blanco. Otros vectores de expresión con pequeñas estructuras de horquilla están basados en lentivirus y han sido utilizados de manera eficiente y estable para suprimir la expresión de genes en ratones transgénicos.81,82
El primer miRNA descrito fue lin–4, que ayuda a controlar la duración de los diferentes estadios del desarrollo de la larva de C. elegans. Este gen no codifica ninguna proteína, sino que produce un par de RNA pequeños. La mayoría de los miRNA provienen de regiones del genoma muy distantes de los genes hasta ahora descritos, lo que sugiere que provienen de unidades de transcripción independientes y se ha mostrado que aproximadamente una cuarta parte de los miRNA descritos hasta ahora están localizados en intrones de pre mRNA.83 Otros miRNA se agrupan de manera secuencial a lo largo del genoma, con un arreglo en el patrón de expresión que implica una transcripción de tipo multicistrónico. La mayoría de los genes que codifican miRNA en nemátodos y en humanos están aislados, mientras que en Drosophila son secuenciales.84 Como se mencionó antes, no se conocen con precisión los blancos de los miRNA y, por lo tanto, tampoco todas sus funciones. Es posible que las secuencias blanco de los miRNA no se restrinjan sólo a RNAm, sino que algunos podrían hibridar con regiones de RNA no codificante.85–88 La identificación de secuencias blanco reguladoras de cada miRNA está aún muy lejana. Para identificarlas dentro de las rutas metabólicas celulares en un contexto biológico, probablemente será necesaria la intervención de otras ciencias como la bioquímica, la bioinformática y la genética.
DIFERENCIAS ENTRE siRNA Y miRNA
Como hemos visto, los siRNA y los miRNA comparten algunas funciones y muchos de sus aspectos, incluyendo su composición química y mecanismos de acción. Además, parte de la biogénesis de miRNA y siRNA endógenos es compartida, todo lo cual hace difícil la distinción entre ellos. Sin embargo, algunas estriban en:
1. Los miRNA son procesados principalmente a partir de transcritos en horquilla, mientras que los siRNA provienen generalmente de grandes moléculas de RNA dúplex.
2. Cada molécula precursora de horquilla genera un solo RNA dúplex de miRNA:miRNA, mientras que cada siRNA precursor genera varios siRNA dúplex diferentes, provenientes de cada extremo del RNAdc.
]]> 3. Las secuencias de miRNA están relativamente conservadas en organismos relacionados, no así las secuencias endógenas de siRNA.4. Es posible que los siRNA endógenos realicen un "autosilenciamiento", ya que silencian el mismo locus, o uno muy similar al que les dio origen, mientras que los miRNA llevan a cabo un "heterosilenciamiento", ya que algunos se producen a partir de genes que pueden silenciar a varios genes blanco.63,89
LA RUTA DEL RNAi EN LA GENÉTICA DE MAMÍFEROS
En experimentos basados en RNAi, una de las primeras decisiones que hay que tomar es cuál tipo de RNA, siRNA o miRNA habrá de utilizarse para activar la supresión. Las principales ventajas del siRNA son: su alta eficacia para la liberación de secuencias dentro de la célula, lo que resulta en concentraciones altas del gen silenciador, además de la gran disponibilidad comercial de siRNA prevalidados. Entre las limitaciones de los siRNA destaca el hecho de que sus efectos son transitorios y dependientes de la tasa de división celular, ya que las células de mamíferos no tienen mecanismos para amplificar y propagar el RNAi (como las plantas y C. elegans), además de que algunas células son muy difíciles de transfectar y de que el proceso de transfección per se puede alterar la fisiología de la célula.90–92
En el caso de los miRNA la inversión es mucho mayor, ya que es necesario diseñar oligonucleótidos para clonarlos y secuenciarlos, lo cual es indispensable para producir una construcción óptima. Sin embargo, los miRNA son capaces de producir silenciamiento sostenido y expresarse abundantemente mediante transfección convencional o utilizando diversos vectores virales que permitan su integración estable en el genoma. Además, los vectores de expresión del miRNA pueden ser propagados indefinidamente.57
Ambos se han utilizado para determinar la función de diversos genes in vivo, principalmente en ratones. La primera demostración de silenciamiento en animales adultos mediada por RNAi, se hizo mediante la represión del gen reportero de luciferasa, mediante la transfección de plásmidos con siRNA y miRNA en el hígado de un ratón.93 En estudios subsecuentes se introdujeron siRNA y miRNA a varias células de diferentes formas, como inyección del ácido nucleico desnudo o mediante lipofección inhibiendo la expresión de diversos genes blanco.
El silenciamiento de genes a largo plazo in vivo se ha demostrado produciendo mosaicismo genético y transferencia a la línea germinal. Por ejemplo, el crecimiento de un tumor en un modelo xenogénico puede atenuarse agregando a las células un cásete de miRNA cuyo blanco sea el oncogén RAS, permitiendo que éste se active antes de reinyectar las células al animal. También se ha suprimido la expresión de genes específicos en células germinales y sus órganos productores.94–98
Algunos oligonucleótidos sintéticos suprimen el gen blanco de manera transmisible, basados en la acción de heredabilidad dominante de un cásete de expresión de miRNA. Con el éxito de estas estrategias surgieron muchos experimentos que incluyeron inyecciones nucleares, creación de quimeras mediante células germinales sometidas a ingeniería genética, y por transgénesis mediante la inyección de lentivirus en huevos fertilizados. En la actualidad, esta tecnología ya permite la creación de animales con silenciamiento inducible de casi cualquier gen.81,82,99,100
]]> EL RNAi COMO UNA HERRAMIENTA DE ANÁLISIS GENÓMICO GLOBAL
Paradójicamente los miRNA pueden también utilizarse para el análisis genómico global a partir de estudios a pequeña escala. Recientemente se utilizó una biblioteca de estos RNA dirigida a la familia de las enzimas de la desubicuitinación, encontrando que el gen supresor de tumor CYLD (gen de susceptibilidad a cilindromatosis) suprime la actividad de NF–kB. Este resultado originó diversas propuestas para el tratamiento de la cilindromatosis con fármacos ya existentes, y confirmó que los estudios genéticos no solamente generan avances en la biología, sino que podrían proveer avances prácticos en el tratamiento racional de enfermedades.101–104
Las bibliotecas de siRNA pueden construirse mediante síntesis química o por digestión enzimática de RNAdc largos. De manera alternativa, la construcción de varios vectores de expresión de miRNA que tengan cada uno un gen blanco, permite también la producción de bibliotecas. Recientemente, dos grupos publicaron la producción de bibliotecas a partir de oligonucleótidos sintéticos que cubren aproximadamente 10,000 genes únicos.105 Otro grupo de investigación creó una biblioteca de miRNA mediante productos de PCR. Por otro lado, también existen métodos para construir bibliotecas de miRNA basadas en la manipulación del RNA complementario.106 Todo esto está en desarrollo y pronto veremos sus resultados.
Los análisis a gran escala con bibliotecas de siRNA, pueden realizarse por medio de microarreglos imprimiendo diferentes miRNA o siRNA sobre laminillas de sílice para realizar trasfecciones inversas. Este tipo de transfecciones involucran a la deposición de complejos ácidos nucleicolípidos sobre una superficie sólida y las células que se plaquean sobre esta superficie "tomarán" el DNA o RNA encapsulado para regular la expresión o el silenciamiento del RNAm.107–109 Otra manera de valorar los efectos del miRNA es por medio de una transfección in situ, seleccionando las células transfectadas en cultivo ya sea mediante citosinas o bien por antibióticos y posteriormente realizar microarreglos de expresión para determinar la expresión diferencial en cada una de las células (Figura 4).110–112

En conclusión, el éxito del empleo del RNAi para analizar la función de un solo gen ha llevado a aplicar esta tecnología a gran escala. Ahora que los genomas del humano, ratón, rata y chimpancé están completamente secuenciados, el RNAi provee un mecanismo mediante el cual toda esta información puede ser traducida a generar definiciones funcionales para cada gen.
RNAi EN EL TRATAMIENTO Y DESCUBRIMIENTO DE NUEVOS FÁRMACOS
Como hemos visto, el RNAi ha comenzado a cambiar los paradigmas hasta ahora existentes en el proceso del descubrimiento de nuevos fármacos. Con los métodos de análisis a gran escala que se mencionaron, el RNAi cobra gran importancia, ya que puede enfocarse directamente a la búsqueda de los blancos de fármacos más promisorios.
]]> Sin embargo, desde que se describió por primera vez la utilización de RNAi en células de mamíferos, se han realizado muchos estudios para utilizarlos en el tratamiento de enfermedades. Su eficacia en la terapéutica dependerá de la especificidad de la inhibición a la que el gen blanco es sometido. En tal caso, las posibles secuencias blanco para tratar enfermedades serían: oncogenes, genes supresores o incluso polimorfismos de un solo nucleótido (SNP). Además, hay grandes esperanzas de poder algún día utilizar el RNAi en el tratamiento de enfermedades virales como la hepatitis C y las infecciones por virus de inmunodeficiencia humana (VIH), de lo cual hay resultados preliminares. No obstante el gran potencial de esta técnica en terapéutica, debemos mantener en mente su posible toxicidad, cuyas consecuencias no estamos en condiciones de predecir. Actualmente existen propuestas clínicas para utilizar miRNA sintéticos o vectores virales como tratamiento, pero ninguna ha sido aprobada.
RNAi EN EL TRATAMIENTO DE PACIENTES CON VIH
El uso de la combinación de dos o tres fármacos en el tratamiento del VIH ha ayudado enormemente a los pacientes infectados. Sin embargo, a pesar del aparente éxito de los nuevos agentes antirretrovirales, persisten problemas debido principalmente al surgimiento de variantes virales resistentes a dichos agentes y a la emergente toxicidad resultante de las mezclas de los diferentes fármacos. De ahí el gran interés en la búsqueda de nuevos tratamientos antirretrovirales. El VIH fue el primer agente infeccioso estudiado mediante RNAi, probablemente debido a que su patrón de expresión y ciclo de vida están siendo conocidos. Se han utilizado varias secuencias sintéticas de siRNA y miRNA para silenciar los RNA de expresión temprana y tardía del VIH en líneas celulares y en células hematopoyéticas.113–114 Aunque la inhibición de secuencias de VIH en cultivos celulares por RNAi ha sido efectiva hasta ahora, el reto actual es la alta tasa de mutación del virus que, además de permitirle evadir la respuesta inmune, podría permitir también a las secuencias mutantes escapar del silenciamiento. De ahí que otra alternativa terapéutica en esta enfermedad es la supresión de co–factores celulares requeridos para la infección por VIH. Algunos cofactores celulares como el receptor de VIH CD4 y otros, han sido suprimidos en líneas celulares de linfocitos T y células germinales derivadas de macrófagos mediante RNAi, resultando en inhibición de la replicación del VIH o de la invasión celular.113
La liberación de siRNA y miRNA dentro de las células infectadas con VIH es un problema. Dado que la acción de los siRNA no persiste largos periodos de tiempo en la célula, y considerando que las células a tratar son linfocitos T, monocitos y macrófagos, estos RNA pequeños deben ser introducidos en las células varias veces durante años para tener éxito en el control de la infección.100,115–117 Además, cualquier método de transfección que no tenga 100% de eficacia permite la persistencia de clonas infectadas que a largo plazo, al expandirse, permiten la persistencia del virus. Por otro lado, la utilización de vectores virales de miRNA también causa problemas debido a la respuesta inmune que el mismo vector puede causar en el individuo.100,118,119 De ahí que uno de los métodos que han sido mayormente utilizados sea aislar células T de los pacientes para ser transfectadas, crecidas e introducidas en los mismos pacientes. Para lo anterior, también se han utilizado células precursoras transfectadas con vectores que contienen los genes terapéuticos, las cuales generarán células hematopoyéticas resistentes al virus. 115,120,121 Las células madre son adquiridas del paciente y transducidas ex vivo antes de transferencia adoptiva o reconstitución in vivo. Debido a que el RNAi ha demostrado ser más exitoso que algunas sondas antisentido o ribozimas que se utilizan actualmente, se cree que esta tecnología podrá ser utilizada en humanos en los próximos años.115
RNAi EN EL TRATAMIENTO DE LAS HEPATITIS VIRALES
La hepatitis producida por los virus de hepatitis B (VHB) y C (VHC) es un problema muy importante de salud a nivel mundial, ya que millones de individuos se infectan anualmente. Hoy en día existe una vacuna contra el VHB pero el tratamiento solamente funciona para prevenir la infección viral y no existe todavía vacuna para VHC. De ahí que la hepatitis causada por estos dos agentes virales sea un blanco potencial de terapia por RNAi.122
El VHC infecta aproximadamente a 3% de la población mundial y es el factor etiológico más importante de enfermedad crónica del hígado que puede progresar a cirrosis hepática y a carcinoma hepático. El genoma del VHC es una cadena positiva de RNA que se procesa postranscripcionalmente para producir por lo menos diez proteínas. En algunos estudios se han transfectado regiones subgenómicas o replicones con el genoma del VHC a células derivadas de hepatocarcinoma para estudiar nuevos fármacos antivirales. En otros trabajos se ha inyectado siRNA antiVHC en ratones para digerir secuencias de VHC utilizando una construcción de VHC–luciferasa, como gen reportero, en hepatocitos de ratón in vivo.123,126
En estudios in vivo se utilizaron siRNA para tratar la hepatitis fulminante inducida por un anticuerpo agonista antifas en ratones. Los siRNA anti–fas se inyectaron a ratones previamente tratados con el anticuerpo. El 82% de los ratones sobrevivió por 10 días, mientras que los controles murieron a los tres días. Los ratones que padecían hepatitis autoinmune también mejoraron después del tratamiento con siRNA. De ahí que el siRNA podría disminuir la severidad de algunas enfermedades interfiriendo con la respuesta inflamatoria más que con el agente infeccioso mismo. De la misma manera que con el VIH, la liberación de los siRNA o de los vectores de miRNA es el reto actual a vencer para poder lograr un posible tratamiento del VHC, ya que el método de inyección intravenosa utilizada en ratones no es factible en el tratamiento de la hepatitis humana. Aunque en ratones el material genético puede introducirse en los hepatocitos utilizando catéteres, no existen métodos de liberación efectiva y sin riesgos para su uso en humanos.127,129
]]>RNAi Y CÁNCER
Algunos estudios sugieren que el RNAi puede ser utilizado como tratamiento potencial en cáncer. Aunque no existen protocolos de RNAi en proceso, los antecedentes son estudios en que se utilizó DNA antisentido. El primer oligonucleótido antisentido utilizado para el tratamiento del cáncer fue el Genasense (Genta, Inc.), que tiene como blanco al gen antiapoptótico BCL–2, su uso, en combinación con quimioterapia convencional, ha dado resultados alentadores en melanoma. Sin embargo, su uso como fármaco recientemente aprobado por la FDA ha sido postergado sin explicar la causa.130,131 Es innegable que el uso potencial del RNAi para tratar cáncer metastático dependerá de encontrar buenos genes blanco en las células.
Aparentemente la expresión de los miRNA es regulada tanto por el tipo de célula como por su estadio de progresión. En 2003 se reportó que este aspecto de especificidad se mantiene aún en cáncer, ya que al medir la expresión de 217 miRNA humanos en muestras de cáncer encontraron que el patrón de expresión de estos RNA pequeños varía en los diferentes tipos de tumor. Es importante mencionar que los patrones de expresión de este pequeño grupo de miRNA definen e identifican mejor los diferentes tipos de cáncer que los datos de expresión generados por microarreglos de cDNA con 16,000 clonas, lo que lo ubica como una novedosa estrategia para clasificar los diferentes tipos de cáncer.132,133
Como se esperaba, la expresión de algunos miRNA está relacionada directamente con el desarrollo y con el tejido en que se desarrollan. Los tumores de un mismo origen embrionario tienen patrones de expresión similares (por ejemplo el cáncer gástrico, el hepatocarcinoma y el cáncer de colon, que derivan del endodermo). Las leucemias tienen un patrón muy diferente al de los tumores sólidos y están subagrupadas de acuerdo con anormalidades genéticas. Es posible que más observaciones ayuden a mejorar el diagnóstico de los cánceres poco diferenciados con orígenes desconocidos y aporten mejores opciones para el tratamiento.
Considerando que en algunos linfomas hay copias extras de un fragmento del cromosoma 13 (13q31–32), se determinó que en esta región que se amplifica existe un gen que se llama cl3orf25 que codifica el precursor de siete miRNA. He et al. encontraron que los miRNA derivados de cl3orf25 se sobrexpresan en células de linfoma que tienen copias extra del cl3orf25. Para identificar las consecuencias biológicas de la sobrexpresión de este miRNA, se utilizó un modelo murino de linfoma inducido por el oncogen myc. Se infectaron células precursoras hematopoyéticas de este ratón con un retrovirus que contenía una región del cluster de miRNA, y se inyectaron en ratones. En éstos, la leucemia se desarrolló en 51 días, mientras que en los controles, inyectados con un vector vacío, la leucemia se desarrolló en un periodo de 3 a 6 meses. El mecanismo de funcionamiento no ha sido aún descrito y tampoco se ha determinado cuál de los componentes del cluster de cl3orf25 es el causante. Sin embargo, este estudio identifica al primer oncogen que no codifica ninguna proteína.134
En cáncer, frecuentemente se sobreexpresa el gen regulador de la trascripción C–MYC. En un trabajo reciente de O'Donell et al., se utilizó una línea celular de linfoma que expresa el gen C–MYC. En estas células se activó este transcrito utilizando un inductor y se detectó que el aumento en la expresión de C–MYC resulta en expresión de seis miRNA. De éstos, dos están codificados por el cluster de cl3orf25 y el resto por clusters de los cromosomas 7 y X.135
Es evidente la necesidad de definir las rutas biológicas alteradas en cáncer que son reguladas por miRNA. Esto resulta bastante complejo debido a las funciones duales de los mismos miRNA como las previamente descritas. La identificación de las redes reguladoras de los miRNA es uno de los retos actuales, pero esto se apoya en la evidencia de que la regulación de la expresión génica es esencial en el estudio del crecimiento, desarrollo y sobrevivencia de los organismos multicelulares.
RNAi, VIRUS DE PAPILOMA HUMANO Y CÁNCER CERVICOUTERINO
]]> El cáncer cervicouterino (CC) es la segunda causa de muerte en mujeres por neoplasia en México. El virus de papiloma humano (VPH) es considerado el factor etiológico más importante, ya que se encuentra presente en 99% de todas las muestras. La infección por VPH a la célula hace que se traduzcan dos oncoproteínas virales denominadas E6 y E7, las cuales se unen a las proteínas celulares p53 y RB respectivamente, inactivándolas. Recientemente, diversos grupos se han dado a la tarea de diseñar una vacuna contra este virus; sin embargo, hasta la fecha no existe ninguna de estas vacunas, ya sea preventiva o de tratamiento disponible en el mercado.Basándose en que la inhibición de factores antiapoptóticos podría tener gran importancia en terapias antitumorales, trabajos recientes reportan que en líneas celulares derivadas de CC e infectadas con VPH cuando se utiliza siRNA para el oncogen E6, se acumula p53, se transactiva el gen p21 (controlador del ciclo celular) y se reduce el crecimiento celular, además de que se restablece la actividad de las rutas supresoras de tumor. Mientras que al inhibir E7 se induce la muerte celular abriendo la posibilidad de utilizar siRNA en el tratamiento del CC. Por otro lado, trabajos recientes indican que cuando se utiliza siRNA contra las oncoproteínas virales E6 y E7, además de los efectos antes mencionados, se incrementa el efecto terapéutico del cisplatino, que es el tratamiento más empleado en CC, como resultado directo del incremento en los niveles de p53. Sin embargo, aunque el panorama resulte bastante similar al de VIH y VHC, es muy evidente que el estudio del cáncer cervicouterino mediante RNAi aún no ha tenido la atención suficiente, convirtiéndolo en un blanco de investigación.136–139
CONCLUSIONES
En un periodo de tiempo relativamente corto, y a partir de su descubrimiento en modelos animales, la ruta de silenciamiento del RNAi es ya considerada una poderosa herramienta para la determinación de la función de los genes de mamíferos. Conforme aumente nuestro conocimiento sobre la biología y bioquímica de este mecanismo conservado de regulación génica, mejorará nuestra habilidad para utilizar al RNAi como herramienta experimental. Con el auge de los estudios de RNAi en diferentes modelos animales, en un futuro cercano podrá establecerse la mejor estrategia para utilizar el RNAi en terapia.
Queda una pregunta en el aire: ¿podrá el RNAi revolucionar el tratamiento de enfermedades de la misma manera en que ha revolucionado a la investigación básica en la búsqueda de la función de los genes?
AGRADECIMIENTO
A la M. en C. Patricia Mendoza Lorenzo, por su valiosa participación en la revisión del manuscrito.
]]> REFERENCIAS
1. Eldering E, Spek CA, Aberson HL, Grummels A, Derks IA, de Vos AF, et al. Expression profiling via novel multiplex assay allows rapid assessment of gene regulation in defined signalling pathways. Nucleic Acids Res 2003; 31: el53. [ Links ]
2. Cullen BR. Derivation and function of small interfering RNAs and microRNAs. Virus Res 2004; 102: 3–9. [ Links ]
3. Cullen BR. RNAi the natural way. Nat Genet 2005; 37: 1163–5. [ Links ]
4. Couzin J. Molecular biology. RNAi shows cracks in its armor. Science 2004; 306: 1124–5. [ Links ]
5. Novina CD, Sharp PA. The RNAi revolution. Nature 2004; 430: 161–4. [ Links ]
6. Whither RNAi? Nat.Cell Biol. 2003; 5: 489–90. [ Links ]
7. Scherr M, Eder M. RNAi in functional genomics. Curr Opin Mol Ther 2004; 6: 129–35. [ Links ]
8. Tomari Y, Zamore PD. Perspective: machines for RNAi. Genes Dev. 2005; 19: 517–29. [ Links ]
9. Chalk AM, Warfinge RE, Georgii–Hemming P, Sonnhammer EL. siRNAdb: a database of siRNA sequences. Nucleic Acids Res 2005; 33: D131–D134. [ Links ]
10. Friedrich I, Ben–Bassat H, Levitzki A. Activation of dsRNA Dependent Protein Kinase PKR in Karpas299 Does Not Lead to Cell Death. Cancer Biol Ther 2005; 4: 734–9. [ Links ]
11. Piano F, Schetter AJ, Morton DG, Gunsalus KC, Reinke V, Kim SK, et al. Gene clustering based on RNAi phenotypes of ovary–enriched genes in C. elegans. Curr Biol 2002; 12: 1959–64. [ Links ]
12. Lipardi C, Wei Q, Paterson BM. RNAi as random degradative PCR: siRNA primers convert mRNA into dsRNAs that are degraded to generate new siRNAs. Cell 2001; 107: 297–307. [ Links ]
13. Grishok A, Pasquinelli AE, Conte D, Li N, Parrish S, Ha I, et al. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 2001; 106: 23–34. [ Links ]
14. Parrish S, Fleenor J, Xu S, Mello C, Fire A. Functional anatomy of a dsRNA trigger: differential requirement for the two trigger strands in RNA interference. Mol Cell 2000; 6: 1077–87. [ Links ]
15. Gao G, Raikar S, Davenport B, Mutapcic L, Montgomery R, Kuzmin E, et al. Cross–species RNAi: Selected Ascaris suum dsRNAs can sterilize Caenorhabditis elegans. Mol Biochem Parasitol 2006; 146: 124–8. [ Links ]
16. Tabara H, Yigit E, Siomi H, Mello CC. The dsRNA binding protein RDE–4 interacts with RDE–1, DCR–1, and a DExH–box helicase to direct RNAi in C. elegans. Cell 2002; 109: 861–71. [ Links ]
17. Yang D, Lu H, Erickson JW. Evidence that processed small dsRNAs may mediate sequence–specific mRNA degradation during RNAi in Drosophila embryos. Curr Biol 2000; 10: 1191–200. [ Links ]
18. Buchón N, Vaury C. RNAi: a defensive RNA–silencing against viruses and transposable elements. Heredity 2006; 96: 195–202. [ Links ]
19. Hansen KR, Ibarra PT, Thon G. Evolutionary–conserved telomere–linked helicase genes of fission yeast are repressed by silencing factors, RNAi components and the telomere–binding protein Tazl. Nucleic Acids Res 2006; 34: 78–88. [ Links ]
20. Hutvagner G, Zamore PD. RNAi: nature abhors a double–strand. Curr Opin Genet Dev 2002; 12: 225–32. [ Links ]
21. Schwenzer R, Zimmermann G, Fotin M, Wajant H, Grell M. No RISC, no fun: assembly of receptor–induced signalling complexes in the tumor necrosis factor system. Eur Cytokine Netw 2000; 11: 519–20. [ Links ]
22. Tavernarakis N, Wang SL, Dorovkov M, Ryazanov A, Driscoll M. Heritable and inducible genetic interference by double–stranded RNA encoded by transgenes. Nat Genet 2000; 24: 180–3. [ Links ]
23. Kato M, Kakutani T. [Control of transposons by DNA methylation and RNAi]. Tanpakushitsu Kakusan Koso 2004; 49: 2097–102. [ Links ]
24. Kuhlmann M, Borisova BE, Kaller M, Larsson P, Stach D, Na J, et al. Silencing of retrotransposons in Dictyostelium by DNA methylation and RNAi. Nucleic Acids Res. 2005; 33: 6405–17. [ Links ]
25. Svoboda P, Stein P, Anger M, Bernstein E, Hannon GJ, Schultz RM. RNAi and expression of retrotransposons MuERV–L and IAP in preimplantation mouse embryos. Dev Biol 2004; 269: 276–85. [ Links ]
26. Dinh A, Mo YY. Alternative approach to generate shRNA from cDNA. Biotechniques 2005; 38: 629–32. [ Links ]
27. Paddison PJ, Hannon GJ. siRNAs and shRNAs: skeleton keys to the human genome. Curr Opin Mol Ther 2003; 5: 217–24. [ Links ]
28. Ying SY, Lin SL. Current perspectives in intronic micro RNAs (miRNAs). J Biomed Sci 2006; 13: 5–15. [ Links ]
29. Filippov V, Solovyev V, Filippova M, Gill SS. A novel type of RNase III family proteins in eukaryotes. Gene 2000; 245: 213–21. [ Links ]
30. Knight SW, Bass BL. A role for the RNase III enzyme DCR–1 in RNA interference and germ line development in Caenorhabditis elegans. Science 2001; 293: 2269–71. [ Links ]
31. Zeng Y, Cullen BR. Efficient processing of primary microRNA hairpins by Drosha requires flanking nonstructured RNA sequences. J Biol Chem 2005; 280: 27595–603. [ Links ]
32. Han Z, Saam JR, Adams HP, Mango SE, Schumacher JM. The C. elegans Tousled–like kinase (TLK–1) has an essential role in transcription. Curr Biol 2003; 13: 1921–9. [ Links ]
33. Bohnsack MT, Czaplinski K, Gorlich D. Exportin 5 is a RanGTP–dependent dsRNA–binding protein that mediates nuclear export of pre–miRNAs. RNA 2004; 10: 185–91. [ Links ]
34. Kim VN. MicroRNA precursors in motion: exportin–5 mediates their nuclear export. Trends Cell Biol 2004; 14: 156–9. [ Links ]
35. Yi R, Doehle BP, Qin Y, Macara IG, Cullen BR. Overexpres–sion of exportin 5 enhances RNA interference mediated by short hairpin RNAs and microRNAs. RNA 2005; 11: 220–6. [ Links ]
36. Yi R, Qin Y, Macara IG, Cullen BR. Exportin–5 mediates the nuclear export of pre–microRNAs and short hairpin RNAs. Genes Dev 2003; 17: 3011–6. [ Links ]
37. Zeng Y, Cullen BR. Structural requirements for pre–microRNA binding and nuclear export by Exportin 5. Nucleic Acids Res 2004; 32: 4776–85. [ Links ]
38. Hiraguri A, Itoh R, Kondo N, Nomura Y, Aizawa D, Murai Y, et al. Specific interactions between Dicer–like proteins and HYLl/DRB–family dsRNA–binding proteins in Arabidopsis thaliana. Plant Mol Biol 2005; 57: 173–88. [ Links ]
39. Kim DH, Behlke MA, Rose SD, Chang MS, Choi S, Rossi JJ. Synthetic dsRNA Dicer substrates enhance RNAi potency and efficacy. Nat Biotechnol 2005; 23: 222–6. [ Links ]
40. Vermeulen A, Behlen L, Reynolds A, Wolfson A, Marshall WS, Karpilow J, et al. The contributions of dsRNA structure to Dicer specificity and efficiency. RNA 2005; 11: 674–82. [ Links ]
41. Zhang H, Kolb FA, Brondani V, Billy E, Filipowicz W. Human Dicer preferentially cleaves dsRNAs at their termini without a requirement for ATP. EMBO J 2002; 21: 5875–85. [ Links ]
42. Tahbaz N, Kolb FA, Zhang H, Jaronczyk K, Filipowicz W, Hobman TC. Characterization of the interactions between mammalian PAZ PIWI domain proteins and Dicer. EMBO Rep 2004; 5: 189–94. [ Links ]
43. Cornejo MF, Maloberti P, Neuman I, Cano F, Castilla R, Castillo F, et al. An arachidonic acid–preferring acyl–CoA synthetase is a hormone–dependent and obligatory protein in the signal transduction pathway of steroidogenic hormones. J Mol Endocrinol 2005; 34: 655–66. [ Links ]
44. Lingel A, Simon B, Izaurralde E, Sattler M. Structure and nucleic–acid binding of the Drosophila Argonaute 2 PAZ domain. Nature 2003; 426: 465–9. [ Links ]
45. Tijsterman M, Okihara KL, Thijssen K, Plasterk RH. PPW–1, a PAZ/PIWI protein required for efficient germline RNAi, is defective in a natural isolate of C. elegans. Curr Biol 2002; 12: 1535–40. [ Links ]
46. Yan KS, Yan S, Farooq A, Han A, Zeng L, Zhou MM. Structure and conserved RNA binding of the PAZ domain. Nature 2003; 426: 468–74. [ Links ]
47. Jaronczyk K, Carmichael JB, Hobman TC. Exploring the functions of RNA interference pathway proteins: some functions are more RISCy than others? Biochem J 2005; 387: 561–71. [ Links ]
48. Hutvagner G. Small RNA asymmetry in RNAi: function in RISC assembly and gene regulation. FEBS Lett 2005; 579: 5850–7. [ Links ]
49. Miyoshi K, Tsukumo H, Nagami T, Siomi H, Siomi MC. Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev 2005; 19: 2837–48. [ Links ]
50. Kolb FA, Zhang H, Jaronczyk K, Tahbaz N, Hobman TC, Filipowicz W. Human dicer: purification, properties, and interaction with PAZ PIWI domain proteins. Methods Enzymol 2005; 392: 316–36. [ Links ]
51. Carthew RW. RNA interference: the fragile X syndrome connection. Curr Biol 2002; 12: R852–R854. [ Links ]
52. Caudy AA, Myers M, Hannon GJ, Hammond SM. Fragile X–related protein and VIG associate with the RNA interference machinery. Genes Dev 2002; 16: 2491–6. [ Links ]
53. Handa V, Saha T, Usdin K. The fragile X syndrome repeats form RNA hairpins that do not activate the interferon–inducible protein kinase, PKR, but are cut by Dicer. Nucleic Acids Res 2003; 31: 6243–8. [ Links ]
54. Ishizuka A, Siomi MC, Siomi H. A Drosophila fragile X protein interacts with components of RNAi and ribosomal proteins. Genes Dev. 2002; 16: 2497–508. [ Links ]
55. Jin P, Zarnescu DC, Ceman S, Nakamoto M, Mowrey J, Jongens TA, et al. Biochemical and genetic interaction between the fragile X mental retardation protein and the microRNA pathway. Nat Neurosci 2004; 7: 113–7. [ Links ]
56. Scadden AD. The RISC subunit Tudor–SN binds to hyper–edited double–stranded RNA and promotes its cleavage. Nat Struct Mol Biol 2005; 12: 489–96. [ Links ]
57. Lin SL, Chang D, Ying SY. Asymmetry of intronic pre–miRNA structures in functional RISC assembly. Gene 2005; 356: 32–8. [ Links ]
58. Maniataki E, Mourelatos Z. A human, ATP–independent, RISC assembly machine fueled by pre–miRNA. Genes Dev 2005; 19: 2979–90. [ Links ]
59. Schwenzer R, Zimmermann G, Fotin M, Wajant H, Grell M. No RISC, no fun: assembly of receptor–induced signalling complexes in the tumor necrosis factor system. Eur.Cytokine Netw. 2000; 11: 519–20. [ Links ]
60. Tomari Y, Du T, Haley B, Schwarz DS, Bennett R, Cook HA, et al. RISC assembly defects in the Drosophila RNAi mutant armitage. Cell 2004; 116: 831–41. [ Links ]
61. regory RI, Chendrimada TP, Cooch N, Shiekhattar R. Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 2005; 123: 631–40. [ Links ]
62. Chiu YL, Dinesh CU, Chu CY, Ali A, Brown KM, Cao H, et al. Dissecting RNA–interference pathway with small molecules. Chem Biol 2005; 12: 643–8. [ Links ]
63. Okamura K, Ishizuka A, Siomi H, Siomi MC. Distinct roles for Argonaute proteins in small RNA–directed RNA cleavage pathways. Genes Dev 2004; 18: 1655–66. [ Links ]
64. Akusjarvi G, Svensson C, Nygard O. A mechanism by which adenovirus virus–associated RNAI controls translation in a transient expression assay. Mol Cell Biol 1987; 7: 549–51. [ Links ]
65. Mathews MB. Control of translation in adenovirus–infected cells. Enzyme 1990; 44: 250–64. [ Links ]
66. Sen GL, Wehrman TS, Blau HM. mRNA translation is not a prerequisite for small interfering RNA–mediated mRNA cleavage. Differentiation 2005; 73: 287–93. [ Links ]
67. Wilson IW, Praszkier J, Pittard AJ. Molecular analysis of RNAI control of repB translation in IncB plasmids. J Bacteriol 1994; 176: 6497–508. [ Links ]
68. Zhao Z, Fang LL, Johnsen R, Baillie DL. ATP–binding cassette protein E is involved in gene transcription and translation in Caenorhabditis elegans. Biochem Biophys Res Commun 2004; 323: 104–11. [ Links ]
69. Allshire R. Molecular biology. RNAi and heterochromatin a hushed–up affair. Science 2002; 297: 1818–9. [ Links ]
70. Martienssen RA, Zaratiegui M, Goto DB. RNA interference and heterochromatin in the fission yeast Schizosaccharomyces pombe. Trends Genet 2005; 21: 450–6. [ Links ]
71. Matzke M, Matzke AJ. RNAi extends its reach. Science 2003; 301: 1060–1. [ Links ]
72. Qian Z, Xuan B, Hong J, Hao Z, Wang L, Huang W. Expression and purification of the carboxyl terminus domain of Schizosaccharomyces pombe dicer in Escherichia coli. Protein Pept Lett 2005; 12: 311–4. [ Links ]
73. Sigova A, Rhind N, Zamore PD. A single Argonaute protein mediates both transcriptional and posttranscriptional silencing in Schizosaccharomyces pombe. Genes Dev 2004; 18: 2359–67. [ Links ]
74. Yamada T, Fischle W, Sugiyama T, Allis CD, Grewal SI. The nucleation and maintenance of heterochromatin by a histone deacetylase in fission yeast. Mol Cell 2005; 20: 173–85. [ Links ]
75. Carette JE, Overmeer RM, Schagen FH, Alemany R, Barski OA, Gerritsen WR, et al. Conditionally replicating adenoviruses expressing short hairpin RNAs silence the expression of a target gene in cancer cells. Cancer Res 2004; 64: 2663–7. [ Links ]
76. Motamedi MR, Verdel A, Colmenares SU, Gerber SA, Gygi SP, Moazed D. Two RNAi complexes, RITS and RDRC, physically interact and localize to noncoding centromeric RNAs. Cell 2004; 119: 789–802. [ Links ]
77. Noma K, Sugiyama T, Cam H, Verdel A, Zofall M, Jia S, et al. RITS acts in cis to promote RNA interference–mediated transcriptional and post–transcriptional silencing. Nat Genet 2004; 36: 1174–80. [ Links ]
78. Verdel A, Jia S, Gerber S, Sugiyama T, Gygi S, Grewal SI, et al. RNAi–mediated targeting of heterochromatin by the RITS complex. Science 2004; 303: 672–6. [ Links ]
79. Verdel A, Moazed D. Labeling and characterization of small RNAs associated with the RNA interference effector complex RITS. Methods Enzymol 2005; 392: 297–307. [ Links ]
80. Pekarik V. Design of shRNAs for RNAi–A lesson from pre–miRNA processing: possible clinical applications. Brain Res Bull 2005; 68: 115–20. [ Links ]
81. An DS, Xie Y, Mao SH, Morizono K, Kung SK, Chen IS. Efficient lentiviral vectors for short hairpin RNA delivery into human cells. Hum Gene Ther 2003; 14: 1207–12. [ Links ]
82. Bbas–Terki T, Blanco–Bose W, Deglon N, Pralong W, Aebischer P. Lentiviral–mediated RNA interference. Hum Gene Ther 2002; 13: 2197–201. [ Links ]
83. Ying SY, Lin SL. Current perspectives in intronic micro RNAs (miRNAs). J Biomed Sci 2006; 13: 5–15. [ Links ]
84. Hayashita Y, Osada H, Tatematsu Y, Yamada H, Yanagisawa K, Tomida S, et al. A polycistronic microRNA cluster, miR–17–92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res 2005; 65: 9628–32. [ Links ]
85. Bagga S, Bracht J, Hunter S, Massirer K, Holtz J, Eachus R, et al. Regulation by let–7 and lin–4 miRNAs results in target mRNA degradation. Cell 2005; 122: 553–63. [ Links ]
86. Hsu PW, Huang HD, Hsu SD, Lin LZ, Tsou AP, Tseng CP, et al. miRNAMap: genomic maps of microRNA genes and their target genes in mammalian genomes. Nucleic Acids Res 2006; 34: D135–D139. [ Links ]
87. Jones–Rhoades MW, Bartel DP. Computational identification of plant microRNAs and their targets, including a stress–induced miRNA. Mol Cell 2004; 14: 787–99. [ Links ]
88. Zhang Y. miRU: an automated plant miRNA target prediction server. Nucleic Acids Res 2005; 33: W701–W704. [ Links ]
89. Ma ZL, Yang HY, Tien P. [Progress of miRNA and its functions in eukaryotes]. Yi.Chuan Xue Bao 2003; 30: 693–6. [ Links ]
90. Abramson J, Rozenblum G, Pecht I. Stable knockdown of MAFA expression in RBL–2H3 cells by siRNA retrovirus–delivery system. Immunol Lett 2004; 92: 179–84. [ Links ]
91. Agrawal N, Malhotra P, Bhatnagar RK. siRNA–directed silencing of transgene expressed in cultured insect cells. Biochem Biophys Res Commun 2004; 320: 428–34. [ Links ]
92. Aigner A. Gene silencing through RNA interference (RNAi) in vivo: Strategies based on the direct application of siRNAs. J Biotechnol 2006. [ Links ]
93. Wu MT, Wu RH, Hung CF, Cheng TL, Tsai WH, Chang WT. Simple and efficient DNA vector–based RNAi systems in mammalian cells. Biochem Biophys Res Commun 2005; 330: 53–9. [ Links ]
94. Cheng AM, Byrom MW, Shelton J, Ford LP. Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic Acids Res 2005; 33: 1290–7. [ Links ]
95. Doench JG, Petersen CP, Sharp PA. siRNAs can function as miRNAs. Genes Dev 2003; 17: 438–42. [ Links ]
96. Kasschau KD, Xie Z, Allen E, Llave C, Chapman EJ, Krizan KA, et al. Pl/HC–Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA unction. Dev Cell 2003; 4: 205–17. [ Links ]
97. Morris JP, McManus MT. Slowing down the Ras lane: miRNAs as tumor suppressors? Set STKE 2005; 2005: e41. [ Links ]
98. Xie Z, Allen E, Fahlgren N, Calamar A, Givan SA, Carrington JC. Expression of Arabidopsis MIRNA genes. Plant Physiol 2005; 138: 2145–54. [ Links ]
99. Arrighi JF, Pion M, Wiznerowicz M, Geijtenbeek TB, Garcia E, Abraham S, et al. Lentivirus–mediated RNA interference of DC–SIGN expression inhibits human immunodeficiency virus transmission from dendritic cells to T cells. J Virol 2004; 78: 10848–55. [ Links ]
100. Banerjea A, Li MJ, Bauer G, Remling L, Lee NS, Rossi J, et al. Inhibition of HIV–1 by lentiviral vector–transduced siR–NAs in T lymphocytes differentiated in SCID–hu mice and CD34+ progenitor cell–derived macrophages. Mol Ther 2003; 8: 62–71. [ Links ]
101. Bhattacharya S, Ray RM, Johnson LR. Decreased apoptosis in polyamine depleted IEC–6 cells depends on Akt–mediated NF–kappaB activation but not GSK3beta activity. Apoptosis 2005; 10: 759–76. [ Links ]
102. Brummelkamp TR, Nijman SM, Dirac AM, Bernards R. Loss of the cylindromatosis tumour suppressor inhibits apoptosis by activating NF–kappaB. Nature 2003; 424: 797–801. [ Links ]
103. Kovalenko A, Chable–Bessia C, Cantarella G, Israel A, Wallach D, Courtois G. The tumour suppressor CYLD negatively regulates NF–kappaB signalling by deubiquitination. Nature 2003; 424: 801–5. [ Links ]
104. Trompouki E, Hatzivassiliou E, Tsichritzis T, Farmer H, Ashworth A, Mosialos G. CYLD is a deubiquitinating enzyme that negatively regulates NF–kappaB activation by TNFR family members. Nature 2003; 424: 793–6. [ Links ]
105. Silva JM, Li MZ, Chang K, Ge W, Golding MC, Rickles RJ, et al. Second–generation shRNA libraries covering the mouse and human genomes. Nat Genet 2005; 37: 1281–8. [ Links ]
106. Lai EC, Wiel C, Rubin GM. Complementary miRNA pairs suggest a regulatory role for miRNA:miRNA duplexes. RNA 2004; 10: 171–5. [ Links ]
107. Amarzguioui M. Improved siRNA–mediated silencing in refractory adherent cell lines by detachment and transfection in suspension. Biotechniques 2004; 36: 766–8, 770. [ Links ]
108. Beale G, Hollins AJ, Benboubetra M, Sohail M, Fox SP, Benter I, et al. Gene silencing nucleic acids designed by scanning arrays: anti–EGFR activity of siRNA, ribozyme and DNA enzymes targeting a single hybridization–accessible region using the same delivery system. J Drug Target 2003; 11: 449–56. [ Links ]
109. Chang HS, Lin CH, Chen YC, Yu WC. Using siRNA technique to generate transgenic animals with spatiotemporal and conditional gene knockdown. Am J Pathol 2004; 165: 1535–41. [ Links ]
110. Chen S, Wang G, Makrigiorgos GM, Price BD. Stable siRNA–mediated silencing of ATM alters the transcriptional profile of HeLa cells. Biochem Biophys Res Commun 2004; 317: 1037–44. [ Links ]
111. Yoshikawa T, Uchimura E, Kishi M, Funeriu DP, Miyake M, Miyake J. Transfection microarray of human mesenchymal stem cells and on–chip siRNA gene knockdown. J Control Release 2004; 96: 227–32. [ Links ]
112. Zhelev Z, Bakalova R, Ohba H, Ewis A, Ishikawa M, Shinohara Y, et al. Suppression of bcr–abl synthesis by siRNAs or tyrosine kinase activity by Glivec alters different oncogenes, apoptotic/antiapoptotic genes and cell proliferation factors (microarray study). FEBS Lett 2004; 570: 195–204. [ Links ]
113. Couturier JP, Root–Bernstein RS. HIV may produce inhibitory microRNAs (miRNAs) that block production of CD28, CD4 and some interleukins. J Theor Biol 2005; 235: 169–84. [ Links ]
114. Yeung ML, Bennasser Y, Le SY, Jeang KT. siRNA, miRNA and HIV: promises and challenges. Cell Res 2005; 15: 935–46. [ Links ]
115. Akkina R, Banerjea A, Bai J, Anderson J, Li MJ, Rossi J. siRNAs, ribozymes and RNA decoys in modeling stem cell–based gene therapy for HIV/AIDS. Anticancer Res 2003; 23: 1997–2005. [ Links ]
116. Andersen JL, Zimmerman ES, DeHart JL, Murala S, Ardon O, Blackett J, et al. ATR and GADD45alpha mediate HIV–1 Vpr–induced apoptosis. Cell Death Differ 2005; 12: 326–34. [ Links ]
117. Bagasra O, Prilliman KR. RNA interference: the molecular immune system. J Mol Histol 2004; 35: 545–53. [ Links ]
118. Takaku H. Gene silencing of HIV–1 by RNA interference. Antivir Chem Chemother 2004; 15: 57–65. [ Links ]
119. Van den HC, Eggermont K, Nuttin B, Debyser Z, Baekelandt V. Lentiviral vector–mediated delivery of short hairpin RNA results in persistent knockdown of gene expression in mouse brain. Hum Gene Ther 2003; 14: 1799–807. [ Links ]
120. Dave RS, Pomerantz RJ. Antiviral effects of human immunodeficiency virus type 1–specific small interfering RNAs against targets conserved in select neurotropic viral strains. J Virol 2004; 78: 13687–96. [ Links ]
121. Forstemann K, Tomari Y, Du T, Vagin VV, Denli AM, Bratu DP, et al. Normal microRNA maturation and germ–line stem cell maintenance requires Loquacious, a double–stranded RNA–binding domain protein. P Lo S Biol 2005; 3: e236. [ Links ]
122. Cai DC, Ren H. [Advance in research of anti–HBV and anti–HCV through RNAi]. Zhonghua Gan Zang Bing Za Zhi 2004; 12: 126–8. [ Links ]
123. Azkur AK, Kim B, Suvas S, Lee Y, Kumaraguru U, Rouse BT. Blocking mouse MMP–9 production in tumor cells and mouse cornea by short hairpin (sh) RNA encoding plasmids. Oligonucleotides 2005; 15: 72–84. [ Links ]
124. Bron R, Eickholt BJ, Vermeren M, Fragale N, Cohen J. Functional knockdown of neuropilin–1 in the developing chick nervous system by siRNA hairpins phenocopies genetic ablation in the mouse. Dev Dyn 2004; 230: 299–308. [ Links ]
125. Calegari F, Haubensak W, Yang D, Huttner WB, Buchholz F. Tissue–specific RNA interference in postimplantation mouse embryos with endoribonuclease–prepared short interfering RNA. Proc Nati Acad Sci USA 2002; 99: 14236–40. [ Links ]
126. Cui XS, Li XY, Jeong YJ, Jun JH, Kim NH. Gene Expression of Cox 5a, 5b, or 6b 1, and their Roles in Preimplantation Mouse Embryos. Biol Reprod 2006. [ Links ]
127. Davern TJ. Increasing the RISC for HCV. Gastroenterol 2003; 125: 1546–8. [ Links ]
128. Goh PY, Tan YJ, Lim SP, Tan YH, Lim SG, Fuller–Pace F, et al. Cellular RNA helicase p68 relocalization and interaction with the hepatitis C virus (HCV) NS5B protein and the potential role of p68 in HCV RNA replication. J Virol 2004; 78: 5288–98. [ Links ]
129. Inglot M, Gladysz A, Rymer W. [Experimental therapy in HCV infection]. Przegl Epidemiol 2005; 59: 525–33. [ Links ]
130. Albertella MR, Lau A, O'Connor MJ. The overexpression of specialized DNA polymerases in cancer. DNA Repair (Amst) 2005; 4: 583–93. [ Links ]
131. Fu GF, Lin XH, Han QW, Fan YR, Xu YF, Guo D, et al. RNA Interference Remarkably Suppresses bcl–2 Gene Expression in Cancer Cells in Vitro and in Vivo. Cancer Bio L Ther 2005; 4. [ Links ]
132. Nishigaki M, Aoyagi K, Danjoh I, Fukaya M, Yanagihara K, Sakamoto H, et al. Discovery of aberrant expression of RRAS by cancer–linked DNA hypomethylation in gastric cancer using microarrays. Cancer Res 2005; 65: 2115–24. [ Links ]
133. Williams NS, Gaynor RB, Scoggin S, Verma U, Gokaslan T, Simmang C, et al. Identification and validation of genes involved in the pathogenesis of colorectal cancer using cDNA microarrays and RNA interference. Clin Cancer Res 2003; 9: 931–46. [ Links ]
134. Ota A, Tagawa H, Karnan S, Tsuzuki S, Karpas A, Kira S, et al. Identification and characterization of a novel gene, C13orf25, as a target for 13q31–q32 amplification in malignant lymphoma. Cancer Res 2004; 64: 3087–95. [ Links ]
135. O'Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT. c–Myc–regulated microRNAs modulate E2F1 expression. Nature 2005; 435: 839–43. [ Links ]
136. Finzer P, Krueger A, Stohr M, Brenner D, Soto U, Kuntzen C, et al. HDAC inhibitors trigger apoptosis in HPV–positive cells by inducing the E2F–p73 pathway. Oncogene 2004; 23: 4807–17. [ Links ]
137. Hougardy BM, Maduro JH, van der Zee AG, de Groot DJ, van den Heuvel FA, de Vries EG, et al. Proteasome inhibitor MG132 sensitizes HPV–positive human cervical cancer cells to rhTRAIL–induced apoptosis. Int J Cancer 2005. [ Links ]
138. Jiang M, Milner J. Selective silencing of viral gene expression in HPV–positive human cervical carcinoma cells treated with siRNA, a primer of RNA interference. Oncogene 2002; 21: 6041–8. [ Links ]
139. Tang S, Tao M, McCoy JP, Zheng ZM. Short–term induction and long–term suppression of HPV16 oncogene silencing by RNA interference in cervical cancer cells. Oncogene 2005. [ Links ] ]]>