Amaranth is a plant of interest in farming due to its ability to adapt into arid and semi-arid climates, where it can grow and yield seed and substantial amounts of biomass. The seed, which is gluten-free and contains relatively high levels of lysine (Clouse et al. 2016), is widely appreciated in Mexico as a food and its leaves are suitable for human consumption and as forage because they provide high-quality protein (Barba de la Rosa et al. 2009). The amaranth species studied in the current investigation, Amaranthus hypochondriacus L., has been cultivated in the Valley of Mexico under semi-arid conditions. Despite its agronomic and nutritional attributes, little is known about the activity and diversity of microbial communities associated with amaranth roots.
One important function of all soil microorganisms comprises the decomposition of soil organic matter (SOM). Proteases are involved in the hydrolysis of peptide bonds to release nitrogen (N); phosphatases catalyze the hydrolysis of esters and anhydrides of phosphoric acid; cellulases hydrolyze cellulose into glucose; and chitinase is capable of hydrolyzing insoluble chitin into its oligo and monomeric components (Kuddus & Ahmad 2013, Gianfreda 2015). Dehydrogenases (DH) also play a significant role in decomposition of SOM.
Microbial activity can increase in certain rhizosphere microenvironments, such as rhizosheaths, which are highly stable structures shaped by the enmeshment of soil particles (sand, silt and organic matter) by root-hairs (Othman et al. 2004). A high proportion of active soil microorganisms in these microenvironments benefit from the cellular debris and compounds released by roots (Lesuffleur et al. 2007). In the rhizosheath, greater microbial activity would be expected than in bulk soil due to the accumulation of organic compounds (Gianfreda 2015). Rhizosheaths are commonly found in the roots of grasses, but have also been found in other cultivated plants, such as amaranth (Figure 1; Moreno-Espíndola et al. 2007). Since plant phenology influences quantity and quality of root exudates, which determine microbial communities in roots (Bencherif et al. 2016, Francioli et al. 2018), it is expected that plant phenology influences rhizosheath traits. Microorganisms living in the rhizosphere of sandy soils promoted the formation of stable macroaggregates (Moreno-Espíndola et al. 2007). Into this context, Moreno-Espíndola et al. (2013) reported the presence of a culturable bacterial community comprising 37 isolates from the amaranth rhizosheath. This community has high heterotrophic potential, which could be useful for improving the fertility of the regional farming systems. It has been demonstrated that soil bacteria (Bacillus and Burkholderia) promote growth and increase yield in grain of amaranth through higher plant P and N uptake (Pal 1998, Parra-Cota et al. 2014). For the development of sustainable systems, the understanding of the microbial processes in soils is required (Philippot et al. 2006), especially in soils in which rain-fed agriculture is practiced.
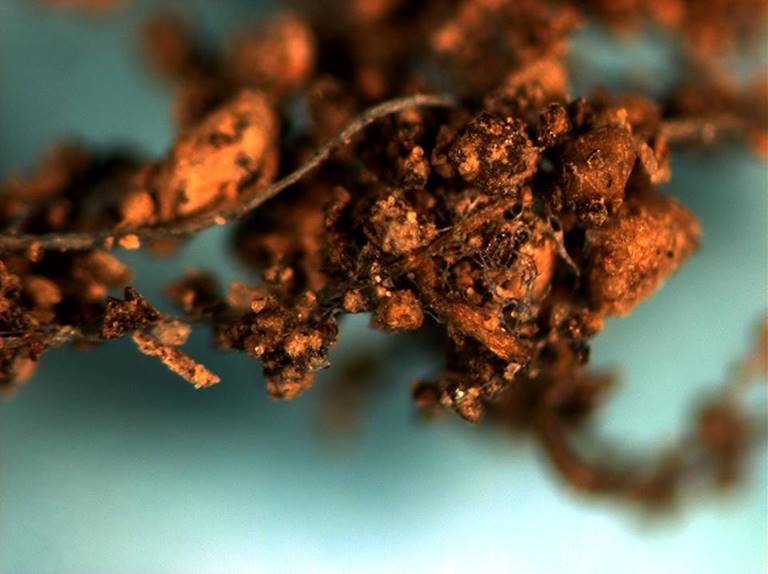
Figure 1 Rhizosheath of amaranth: Root-hairs adhering sand particles in a volcanic soil in the Valley of Mexico.
The aim of this work was to evaluate the potential activity of six enzymes (dehydrogenase, protease, acid and alkaline phosphatases, cellulase and chitinase) involved in the transformation of organic matter in rhizosheath soil and surrounding rhizosheath soil (soil not adhered by roots) of amaranth plants grown in a sandy soil. Likewise, 37 bacterial isolates obtained from amaranth rhizosheath soil were molecularly identified, as microorganisms which potentially perform these enzyme activities.
Materials and methods
Soil and experimental site. The experimental site is located in the Valley of Mexico (Tulyehualco, 19º 15’N, 99º 13”W, 2,280 m in altitude). The soil is an Entisol (Soil Survey Staff 2003), and is composed of volcanic ash and 70, 140, and 790 g kg-1 of clay, silt, and sand, respectively. The predominant minerals are pumice and feldspars, the climate is semi-arid, with an average annual precipitation of 537 mm, a mean temperature of 17.9 ºC (De León-González et al. 2006). In 2009, an experimental field was established with amaranth, which was subdivided into three plots of 12 m2 each. Amaranth was sown under rain-fed conditions, with minimum tillage. An application of a single dose of fertilizer (80-40-00, N-P-K) at the beginning of the flowering period was carried out.
Soil sampling. Sampling of rhizosheath soil and surrounding rhizosheath soil (or loose soil), included four cropping periods of amaranth: sowing, flowering, harvest, and post-harvest (0, 45, 90, and 120 days after sowing). Three soil-root monoliths, one per plant, were obtained with a PVC cylinder (25.5 cm diameter; 37.5 cm length) used as a soil core sampler (Moreno-Espíndola et al. 2007) during each sampling period. Rhizosheath soil was separated from roots using a dissecting needle. The loose soil was sampled 3 cm out near the soil-root monolith. The samples were preserved in sterile plastic bags at 4 ºC.
Soil-organic carbon, microbial-biomass carbon, and total nitrogen. Soil organic carbon (SOC) was measured by the Walkley & Black (1934) method, and total nitrogen (TN), by digestion with H2SO4 (Bremner & Mulvaney 1982). Microbial biomass carbon (Cmic) was measured using the chloroform-fumigation (Vance et al. 1987), and an automatic carbon analyzer (Shimadzu, TOC-VCSN).
Soil pH and soil-water content. The pH of the soil samples was measured potentiometrically (HANNA HI250) in a water-soil suspension (2.5:1). At each phenological stage, soil water content (SWC) (g 100 g-1) was determined in situ using a neutron probe (CNP® MC-1DR), by averaging six measurements obtained by introducing the probe every 5 cm into a depth of 0-30 cm.
Enzymatic activities. The enzymatic activities considered for evaluation were those of dehydrogenase (DH), acid (ACP) and alkaline (ALP) phosphatase, cellulase (CEL), chitinase (CHI), and protease (PRO). All these enzymes participate in processes of soil organic matter (SOM) decomposition. Measurement of the potential of each enzyme activity was performed following the spectrophotometric methods described by García et al. (2003). To determine all absorbances, a Shimadzu Dual-beam Spectrophotometer (Shimadzu UV-1700) was employed. Corresponding calibration curves were generated for each enzymatic activity and each test included a control and four replicates.
Molecular identification of bacterial isolates from amaranth rhizosheath. While enzymatic activity was determined in soils, 37 bacteria were isolated from the amaranth rhizosheath, and physiologically characterized (Moreno-Espíndola et al. 2013). These 37 bacterial isolates were subcultured for molecular identification. Genomic DNA was extracted using the ZRFungal/Bacterial DNA MiniPrepTM kit according to the manufacturer’s instructions. DNA quality was analyzed in a 2 % agarose gel, and quantification was achieved using a Nano-Drop spectrophotometer (Thermo). PCR amplification of the 16S rDNA fragment was performed using primers rD1 (5´AAGGAGGTGATCCAGCC3´) and fD1 (5´AGAGTTTGATCCTGGCTCAG3´) (Laguerre et al. 1994). The PCR mix per sample was as follows: template DNA (50 ng); primers rD1 and fD1 (0.1 μM each); 10X buffer; dNTP (20 μM); MgCl2 (1.5 μM), and Fermentas Taq polymerase (2 U). PCR was performed in a Biometra thermocycler under the following conditions: initial denaturation at 95 ºC for 5 min; 35 cycles of denaturation at 94 ºC for 1 min; annealing at 55 ºC for 1 min, and extension at 72 ºC for 2 min; finally, an extension step at 72 ºC for 3 min. PCR products of approximately 1,500 bp were verified in a 2 % agarose gel and then purified using the QIAquick QiageneTM kit according to the manufacturer’s instructions. Last, 16 μL of the PCR products (150 ng μL-1) of each isolate were sequenced using the forward and reverse primers (Biotechnology Institute, UNAM). The sequences were edited and assembled using DNASTAR SEQman II ver. 5.06 software, and low-quality sequences were eliminated. Using the basic local alignment search tool (BLAST), the sequences were analyzed and compared with those registered in the National Center for Biotechnology Information (NCBI) database to identify their similarity. Finally, the sequences were submitted to the NCBI database.
Statistical analysis. To analyze the data, a two-way analysis of variance (ANOVA) assay was performed. The test considered rhizosheath soil and soil not adhered by roots (loose soil), and the cropping period (four dates: sowing; flowering; harvest, and post-harvest) as factors. When statistically significant differences were detected (p < 0.05), the Tukey mean comparison test was carried out. JMP® 8.0 statistical software was employed.
Results
Properties in the rhizosphere of Amaranthus hypochondriacus. Soil organic carbon (SOC) content and total nitrogen (TN) were significantly higher in rhizosheath soil compared with samples of loose soil, but non-differences were observed concerning microbial biomass carbon (Cmic) even if higher levels were present in loose soil (Table 1). pH and soil water content (SWC) did not exhibit differences between rhizosheath soil and loose soil, which was also observed concerning the six enzyme activities (Table 1). At sowing date roots were not yet developed; by analyzing the other three sampling seasons (flowering, harvest, and post-harvest where the plant roots remained into the soil), TN was always higher in rhizosheath than in loose soil, as well as SOC in flowering and post-harvest seasons; Cmic was three-times higher in loose than in rhizosheath soil in post-harvest season (Table 2). pH and SWC were measured without discriminate between both microenvironments; pH did not show differences between phenological stages, while SWC decreased from 20 g 100 g-1 in sowing to 4.9 g 100 g-1 in post-harvest. Concerning enzyme activity, DH exhibited highest values in post-harvest season compared with the previous two seasons, with differences between the two soil types at harvest and post-harvest stages. ACP demonstrated a tendency to increase as the farming season progressed, and no differences between rhizosphere and loose soils occurred, while ALP showed differences between the two soil types only at the flowering season. CEL showed low values in all three sampling seasons. CHI had similar values in the flowering and harvest seasons without differences between soil types, with a slight decrease in post-harvest season, during which rhizosheath soil was nearly double compared with loose soil. Finally, PRO exhibited differences between soil types at harvest season, with the higher mean for rhizosheath soil (Table 2).
Table 1 Properties and potential enzyme activities in rhizosheath of Amaranthus hypochondriacus, and loose soils (mean±standard error; n = 9). Different letters indicate different mean groups (p < 0.05).
| Rhizosheath | Loose soil | |
|---|---|---|
| Soil properties | ||
| Soil organic carbon (mg 100 g-1) | 0.71±0.05a | 0.41±0.03b |
| Microbial biomass carbon (mg kg-1) | 213.28±29.45a | 284.61±33.66a |
| Total nitrogen (mg 100 g-1) | 0.14±0.00a | 0.04±0.01b |
| pH | 7.1±0.03a | 7.1±0.03a |
| Soil water content (g 100 g-1) | 10.43±2.06a | 9.81±1.83a |
| Potential enzyme activity | ||
| Dehydrogenase (µM INTF g-1 h-1) | 5.01±0.63a | 5.31±0.40a |
| Acid phosphatase (µM p-nitrophenol g-1 h-1) | 0.96±0.04a | 0.93±0.05a |
| Alkaline phosphatase (µM p-nitrophenol g-1 h-1) | 1.16±0.06a | 1.28±0.05a |
| Cellulase (µM equivalent glucose g-1 h-1) | 0.01±0.00a | 0.00±0.00a |
| Chitinase (µM N-acetylglucosamine g-1 h-1) | 12.25±2.88a | 7.96±2.48a |
| Protease (µM tyrosine g-1 h-1) | 13.75±11.38a | 9.89±4.24a |
Table 2 Properties and potential enzyme activities in rhizosheath of Amaranthus hypochondriacus in every phenological stage, and loose soils (mean±standard error; n = 3). Different letters indicate different mean groups (Tukey; p < 0.05).
| Sowing | Flowering | Harvest | Post-harvest | |||||
|---|---|---|---|---|---|---|---|---|
| Base line | Rhizosheath | Loose soil | Rhizosheath | Loose soil | Rhizosheath | Loose soil | ||
| Soil properties | ||||||||
| Soil organic carbon (mg 100 g-1) |
0.48±0.00 | 0.89±0.00a | 0.29±0.01b | 0.53±0.01a | 0.52±0.01a | 0.69±0.00a | 0.41±0.01b | |
| Microbial biomass carbon (mg kg-1) |
183.7±62.8 | 262.5±26.8a | 179.7±24.0a | 260.6±46.1a | 291.1±15.7a | 116.8±24.3b | 383.0±49.1a | |
| Total nitrogen (mg 100 g-1) |
0.11±0.00 | 0.16±0.00a | 0.10±0.00b | 0.16±0.00a | 0.01±0.00b | 0.11±0.00a | 0.00±0.00b | |
| pH | 7.9±0.03 | 7.1±0.04a | 7.1±0.02a | 7.2±0.04a | ||||
| Soil water content (g 100 g-1) |
20.4±0.24 | 17.5±0.90a | 7.9±0.06b | 4.9±0.05c | ||||
| Potential enzyme activity | ||||||||
| Dehydrogenase (µM INTF g-1 h-1) |
6.94±0.07 | 5.03±0.26a | 5.45±0.82a | 2.04±0.36b | 3.91±0.24a | 7.97±0.28a | 6.56±0.37b | |
| Acid phosphatase (µM p-nitrophenol g-1 h-1) |
0.42±0.10 | 0.81±0.01a | 0.72±0.06a | 0.86±0.00a | 0.93±0.04a | 1.21±0.02a | 1.12±0.02a | |
| Alkaline phosphatase (µM p-nitrophenol g-1 h-1) |
0.80±0.06 | 1.06±0.04b | 1.28±0.03a | 0.94±0.03a | 1.10±0.07a | 1.48±0.02a | 1.46±0.03a | |
| Cellulase (µM equivalent glucose g-1 h-1) |
0.064±0.00 | 0.014±0.00a | 0.006±0.00a | 0.008±0.00a | 0.012±0.00a | 0.010±0.00a | 0.005±0.00a | |
| Chitinase (µM N-acetylglucosamine g-1 h-1) |
13.28±3.11 | 13.16±6.51a | 5.44±2.57a | 13.80±5.33a | 13.04±6.43a | 9.78±4.69a | 5.41±2.55a | |
| Protease (µM tyrosine g-1 h-1) |
8.51±0.99 | 25.62±6.02a | 17.48±11.28a | 12.79±1.03a | 2.18±1.77b | 2.85±2.28a | 10.00±5.22a | |
Correlation analysis between all variables measured in this study is shown in Table 3. ACP and ALP activities showed negative and significant correlation with soil pH and SWC. CEL activity had positive and significant correlation with pH and SWC. Finally, a positive correlation between PRO activity and SWC was observed. None of the six enzymatic activities showed significant correlation with Cmic, SOC, or TN (Table 3).
Table 3 Pearson correlation coefficient of overall soil variables measured in rhizosheath and loose soils of Amaranthus hypochondriacus (n = 12).
| pH | SWC | SOC | TN | DH | ACP | ALP | CEL | CHI | PRO | Cmic | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| pH | 1.00 | ||||||||||
| SWC | 0.53* | 1.00 | |||||||||
| SOC | -0.13 | 0.07 | 1.00 | ||||||||
| TN | 0.08 | 0.43 | 0.48* | 1.00 | |||||||
| DH | 0.43 | 0.06 | 0.01 | -0.19 | 1.00 | ||||||
| ACP | -0.66** | -0.84** | 0.18 | -0.30 | 0.15 | 1.00 | |||||
| ALP | -0.46* | -0.54* | -0.11 | -0.41 | 0.39 | 0.77** | 1.00 | ||||
| CEL | 0.91** | 0.59* | -0.02 | 0.14 | 0.30 | -0.70** | -0.63* | 1.00 | |||
| CHI | 0.14 | 0.10 | 0.26 | 0.21 | -0.11 | -0.06 | -0.30 | 0.28 | 1.00 | ||
| PRO | -0.12 | 0.46* | 0.22 | 0.37 | -0.14 | -0.16 | -0.09 | -0.07 | 0.11 | 1.00 | |
| Cmic | -0.25 | -0.22 | -0.12 | -0.42 | -0.22 | 0.24 | 0.08 | -0.22 | -0.06 | 0.12 | 1.00 |
SWC, soil water content; SOC, soil organic carbon; TN, total nitrogen; DH, dehydrogenase; ACP, acid phosphatase; ALP, alkaline phosphatase; CEL, cellulase; CHI, chitinase; PRO, protease; Cmic, microbial biomass carbon; *, **, significant at p < 0.05 and p < 0.01, respectively.
Molecular identification of bacterial isolates. Partial sequences of the 16S rDNA segments of each bacterial isolate were submitted to the GenBank database at NCBI (Table 4). Predominant genera found during the four sampling seasons included Bacillus (29 isolates), followed by Enterobacter (3 isolates), Streptomyces (2 isolates), Stenotrophomonas (1 isolate), Pseudomonas (1 isolate), and Arthrobacter (1 isolate). Streptomyces was detected only during the sowing season. In summary, the flowering season showed highest bacterial diversity, whereas the harvest season had lowest bacterial diversity.
Table 4 Molecular identification based on the 16S rDNA segment of 37 bacterial strains from rhizosheath soil of Amaranthus hypochondriacus and their potential roles.
| Strain | Accesion number in the GenBank | Highest homology (accesion number in the GenBank) | Potential role in the soil (references) |
|---|---|---|---|
| Sowing | |||
| AN1E1 | KC336020 | Streptomyces zaomyceticus (EU593685) | Degradation of recalcitrant organic matter, cellulolytic activity (a, b, e) |
| AN3E1a | KC336021 | Bacillus sp. (AB773240) | P solubilization (a, b, e) |
| AN4E1a | KC336022 | Bacillus sp. (JN132107) | P solubilization (a, b, e) |
| AN5E1b | KC336024 | Bacillus sp. (HQ600985) | P solubilization (a, b, e) |
| AS1E1 | KC336025 | Bacillus cereus (EU661712) | P solubilization, PGPR (a, b, e) |
| AS2E1b | KC336027 | Streptomyces sp. (JQ812096) | Degradation of recalcitrant organic matter, cellulolytic activity (a, b, e) |
| AS4E1b | KC336029 | Bacillus sp. (JN872500) | P solubilization (a, b, e) |
| AS5E1a | KC336030 | Bacillus cereus (JX855262) | P solubilization, PGPR (a, b, e) |
| Flowering | |||
| AN1E2 | KC336032 | Bacillus sp. (JX402435) | P solubilization, PGPR (a, b, e) |
| AN2E2 | KC336033 | Bacillus sp. (JQ396537) | P solubilization, PGPR (a, b, e) |
| AN3E2 | KC336034 | Enterobacter sp. (EU331414) | PGPR (a, c) |
| AN4E2 | KC336035 | Stenotrophomonas sp. (AY689048) | Chitinolytic activity, degradation of xenobiotics (a, b) |
| AN5E2 | KC336036 | Bacillus cereus (EF488087) | P solubilization, PGPR (a, b) |
| AS1E2 | KC336037 | Pseudomonas sp. (HQ224640) | PGPR (a, b) |
| AS2E2b | KC336039 | Enterobacter sp. (HQ677832) | PGPR (a, c) |
| AS3E2a | KC336040 | Enterobacter sp. (GU272374) | PGPR (a, c) |
| AS4E2a | KC336042 | Bacillus sp. (JN210907) | P solubilization, PGPR (a, b, e) |
| AS5E2a | KC336044 | Bacillus sp. (KC121044) | P solubilization, PGPR (a, b) |
| Harvest | |||
| AN1E3 | KC336046 | Bacillus subtilis (HM744709) | P solubilization, biological control, amylase and cellulase production (a, b, e) |
| AN2E3 | KC336047 | Bacillus pumilus (JX847116) | P solubilization, PGPR (a, b, e) |
| AN3E3a | KC336048 | Bacillus pumilus (JX625990) | P solubilization, PGPR (a, b, e) |
| AN4E3 | KC336050 | Bacillus pumilus (JN082266) | P solubilization, PGPR (a, b, e) |
| AN5E3 | KC336051 | Bacillus pumilus (HQ218989) | P solubilization, PGPR (a, b, e) |
| AS1E3 | KC336052 | Bacillus megaterium (AB738793) | P solubilization, PGPR (a, b, e) |
| AS2E3 | KC336053 | Bacillus megaterium (AB738793) | P solubilization, PGPR (a, b, e) |
| AS3E3 | KC336054 | Bacillus aryabhattai (JX460818) | P solubilization, PGPR (a, b, e) |
| AS4E3 | KC336055 | Bacillus megaterium (JX312585) | P solubilization, PGPR (a, b, e) |
| Post-harvest | |||
| AN1E4b | KC336057 | Bacillus sp. (JX897963) | P solubilization, PGPR (a, b) |
| AN2E4b | KC336059 | Bacillus sp. (JX566587) | P solubilization, PGPR (a, b) |
| AN3E4b | KC336061 | Bacillus cereus (JX273683) | P solubilization, PGPR (a, b, e) |
| AN4Eb | KC336062 | Arthrobacter sp. (JF768708) | Degradation of xenobiotics (d) |
| AN5E4b | KC336064 | Bacillus sp. (JX266343) | P solubilization, PGPR (a, b) |
| AS1E4 | KC336065 | Bacillus sp. (HM104462) | P solubilization, PGPR (a, b) |
| AS2E4 | KC336066 | Bacillus megaterium (AB738793) | P solubilization, PGPR (a, b, e) |
| AS3E4 | KC336067 | Bacillus sp. (JF309237) | P solubilization, PGPR (a, b) |
| AS4E4a | KC336068 | Bacillus megaterium (HM357355) | P solubilization, PGPR (a, b, e) |
| AS5E4 | KC336070 | Bacillus aryabhattai (JX460818) | P solubilization, PGPR (a, b, e) |
Strain names were reported by Moreno-Espíndola et al. (2013); PGPR, plant growth-promoting rhizobacteria. References: a) Andrade (2004), b) Yang et al. (2012), c) Schütz et al. (2003), d) Vaishampayan et al. (2007), e) Yasir et al. (2009).
Discussion
The similarity in the general potential enzyme activity in rhizosheath and loose soils indicate biochemical likeness in both microenvironments in the amaranth rhizosphere. This could be explained by the dominant mineral in soil, pumice, which favor equality in water-retention capacity in both soil types (Segura-Castruita et al. 2005), and, by the presence of abundant hyphae in association with the amaranth roots in the same soil (Moreno-Espíndola et al. 2007), which increases the influence zone of the roots. Othman et al. (2004) reported that in the rhizosheaths of two grasses (Panicum turgidum and Stipagrostis scoparia), SOC and TN were two-fold higher than in loose soil. This result is consistent with the fact that grasses are not exposed to mechanical tillage, which allows the accumulation of exudates and organic matter. In our study, the amaranth crop was established with minimum tillage, which favor microbial growth and potential enzymatic activity in the loose soil located in the vicinity of the main root zone.
When comparing potential enzymatic activity between rhizosheath and loose soils for each cropping period, significant statistical differences were found. These results confirm the role of DH activity as a sensitive agronomic indicator (Fuentes-Ponce et al. 2016); in the current study, DH changed with phenological stage and soil microenvironment. Likewise, higher ALP activity during the flowering period in loose soil in comparison with rhizosheath soil coincides with the larger area of influence attributed to this enzyme, which is produced only by microorganisms and not by roots (Spohn & Kuzyakov 2013). Activities of DH and both phosphatases were highest during the post-harvest stage. One factor that may intervene in higher enzyme activity during post-harvest period is the soil-atmosphere gas exchange. It is assumed that, as the soil dries the water-free pores are filled with air and further shrinkage of root and post-harvest root decay increase the number of pores that can be filled with air. Such conditions favor aerobic respiration and increase DH activity during post-harvest. The effects of these changes on rhizosphere conditions over time were also observed in ACP and ALP, which increased as each period elapsed, while CEL behavior was contrariwise. In the current study, the slow degradation rate of cellulose-rich residues is due to a decrease in soil moisture during the dry season and low temperatures during winter. These climatic conditions can explain the low levels of CEL activity, which are mainly driven by anaerobic bacteria. Moreno-Espíndola et al. (2007) reported high soil porosity in this same soil, which could increase O2 availability, resulting in possible inhibition of cellulolytic microbial groups, hence, of their activity. Moreover, Yaroslavtsev et al. (2009) reported that a decrease in soil moisture promoted CHI activity in sandy soils. In contrast, we found a decrease in CHI activity when SWC decreased. SOC and TN results showed that rhizosheaths possessed more favorable conditions for soil organic matter (SOM) accumulation.
In a semi-arid region, plant species increased the organic matter content in their rhizospheres, where enzymes such as phosphatase and urease increased their activity in contrast to a soil without vegetation; rhizosphere induces the synthesis of such enzymes (Garcia et al. 2005). Other semi-arid degraded soil, amended with compost, improved soil water holding capacity, which in turn increased activity of β-glucosidase, phosphatase and DH (Hernández et al. 2015). These results reinforce the need to study the rhizosphere footprint of plant species and the microbial ecology process (York et al. 2016).
In the microbiome identified, the Bacillus genus was the most abundant. During the crop-sowing stage CEL activity was highest, which was consistent with the presence of Streptomyces isolates that play a significant role in aerobic cellulose degradation (Andrade 2004). High cellulose accumulation in soil derives from root tissues of amaranth from the last post-harvest period. During the post-harvest season when phosphatase activities (ALP and ACP) were highest, the bacterial species Arthrobacter spp., Bacillus cereus, B. megaterium, and B. aryabhattai were found. These species have been reported as phosphate-solubilizing bacteria (Yang et al. 2012).
In the present study, highest bacterial diversity was observed during the flowering period which can be related, in part, with the unique fertilization dose applied to the soil, while, during crop harvest, Bacillus was the sole genus present. Stenotrophomonas has been reported as rhizosphere inhabitant cohabiting with Bacillus and Pseudomonas, and their presence is influenced by plant type and by carbon sources (Kapoor & Mukerji 2006). These genera have also been reported as plant growth-promoting rhizobacteria (PGPR) because they produce and release growth hormones (Schütz et al. 2003). The soil bacterial strains identified in the current study have a potential in biotechnological uses (i.e., native microbial consortia useful for biological soil fertility) and for cropping amaranth and other plants under the concept of sustainability (Scotti et al. 2015) in semi-arid conditions.
In conclusion, in the rhizosphere of Amaranthus hypochondriacus grown in a pumiceous sandy soil, potential enzymatic activities in the rhizosheath and loose soils were similar, which must be considered a unique rhizosphere environment. Dehydrogenase and acid phosphatase activities are highly sensitive to changes in the crop phenology. Acid and alkaline phosphatase, cellulose and protease activities correlated to changes in soil moisture. The behavior of phosphatases and dehydrogenase activities suggests a process of active degradation of organic materials and increased dynamic SOM during the post-harvest period. In the amaranth rhizosphere, native bacteria are involved in the breakdown of SOM and they possess potential for biotechnological applications for agriculture as inoculants for the design of novel biological inputs. Other molecular biology methods should be employed for the study of total microbial diversity.











 nueva página del texto (beta)
nueva página del texto (beta)


