Introduction
Salmonella spp. is the agent that causes gastrointestinal infections known as salmonelosis, a disease considered a public health problem worldwide (Kumar et al., 2019). In 2018 there were 124,277 cases of diseases caused by different serotypes of Salmonella in Mexico (SINAVE, 2018), while 103,289 cases were reported by October 2019 (SINAVE, 2019). The pathogen is usually transmitted by water intake and contaminated food, as meat and meat products, which cause severe human health problems when they do not comply with quality and hygiene standards (Fachmann et al., 2017).
Beef consumption in Mexico is only second to that of chicken (FIRA, 2017). By the end of 2018, Mexico was the sixth beef producer worldwide accounting for 3.2 % of the world beef consumption (USDA-FAS, 2018; SIAP, 2018).
In Mexico, the Ministry of Agriculture and Rural Development (SADER, Secretaría de Agricultura y Desarrollo Rural) is responsible for establishing criteria and inspecting the quality of livestock product for human consumption. Through the National Service of Health, Safety, and Food Quality (SENASICA, Servicio Nacional de Sanidad, Inocuidad y Calidad Agroalimentaria), it has determined a categorization of the slaughterhouses operating across the country. Depending on their compliance with international quality and safety standards, SENASICA classifies slaughterhouses as federal-inspection type (TIF, tipo inspección federal) when they follow SENASICA standards and are constantly monitored. In contrast, non-TIF slaughterhouses are not monitored and do not comply with the standards (FIRCO, 2016).
The aim of the present investigation was to analyze and compare the presence of Salmonella spp. in beef from TIF and non-TIF slaughterhouses commercialized in supermarkets, as well as non-TIF beef commercialized in local butchers in Tepic, Nayarit (Mexico).
Material and Methods
Beef samples from TIF (n = 10) and non-TIF (n = 10) slaughterhouses were collected at supermarkets, and non-TIF beef samples (n = 10) were obtained at local butchers in Tepic, Nayarit (Mexico). The samples were taken to the laboratory under asceptic conditions (sterile bags and cold chain) to be immediately used.
Samples were processed following NF Validation™ workflow by Applied Biosystems®. Samples (25 g) were homogenized in 225 mL tetrathionate broth (TT broth) (BD Bioxon®, Mexico) in a stomacher (BagMixer® 400W, France) at a constant speed (8 strokes/sec) for 1 min and incubated at 37 ºC for 20 h. They were then diluted in 15 g/L peptone water (BD Bioxon®, Mexico) at a 1:9 ratio and incubated at 37 ºC for 20 h.
To extract DNA, a PrepSEQ® Rapid Spin preparation kit was used according to the manufacturer’s instructions. Enriched dilutions (750 µL) were placed in spin columns and centrifuged at 12,000 x g for 3 min. The supernatant was discarded, 50 µL lysis buffer were added, and the pellets formed were homogenized. After incubation at 95 ºC for 12 min, samples were allowed to cool at room temperature and centrifuged at 12,000 x g for 1 min. Finally, 250 µL nuclease-free water were added to all samples for homogenization and centrifugation at 12,000 x g for 2 min. The supernatant containing the DNA was used to identify Salmonella spp. by qPCR.
To detect Salmonella spp., a MicroSEQ® Salmonella spp. detection kit was used according to the manufacturer’s instruction. The DNA samples (30 µL) were placed into tubes containing lyophilized beads and carefully mixed. Afterwards, a qPCR was carried out using a 7500 Fast real-time PCR system (Applied Biosystems®) and validated with an internal positive control (IPC) analyzed with each sample. Pathogen detection negative control was used as negative control while a Salmonella spp. culture (0.5 McFarland standard) was used as positive control.
Results were qualitatively interpreted as absence or presence of Salmonella using RapidFinder™ Express software. The Ct values (amplification threshold cycle) obtained were considered semiquantitative regarding contamination by Salmonella spp. in the positive samples.
The qualitative data obtained were analyzed from their relative frequencies using a chi-squared test. Semiquantitative data were analyzed with a Kruskal-Wallis, Dunn’s subtest. Data were analyzed with GraphPad Prism v6.0 (GraphPad software, Inc. USA). Statistical significance was considered when p < 0.05.
Results and Discussion
After the comparative analysis, there was no statistical difference (p > 0.05) between TIF and non-TIF samples (5/10 vs 7/10) from supermarkets regarding the presence of Salmonella spp. However, the bacteria were detected in 100 % of the samples acquired at local butchers (Figure 1).
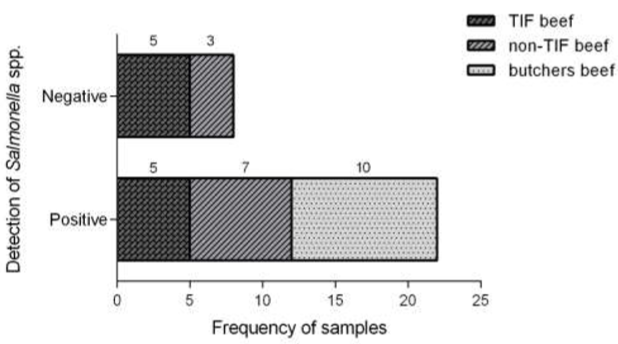
Figure 1 Frequency of TIF and non-TIF beef samples with Salmonella spp. commercialized in Tepic, Nayarit (Mexico). Frequency values were compared by chi-squared test. Different letters in each group indicate a significant statistical difference p < 0.05.
The lack of significant difference between TIF and non-TIF beef suggests that the critical control and contamination prevention points followed in supermarkets are key to lowering the risks related to safety. In addition, TIF slaughterhouses make sure to follow processes that include animal comfort and sacrifice, as well as beef storage (FIRCO, 2017). Therefore, the fact that non-TIF beef distributed in supermarkets has a similar degree of contamination to that of TIF beef indicated that handling at final distribution points is of great importance. Contrastingly, there is high contamination in the beef from local butchers, commercial outlets that usually lack a program of hygiene practices to handle foods, such as temperature control and other safety measures (rodent control and other vectors).
On the other hand, the amplification threshold cycle (Ct) was one of the most relevant parameters in this investigation since it is inversely proportional to the concentration of the pathogen’s DNA; therefore, it can be considered an indirect measure of the degree of contamination in the beef. Additionally, the data distribution obtained from TIF beef was very close to the limit value of Ct established for the negative samples (Ct > 35). Five of these samples even showed Ct values above this value and were then considered negative samples. The remaining five samples of TIF beef were positive to Salmonella spp., while only one of the samples showed a Ct value of 21.90, the other 4 showed Ct values just below 35 (Figure 2) and were therefore slightly contaminated with Salmonella spp. Less sensitive techniques could even consider these samples as negative (false negatives).
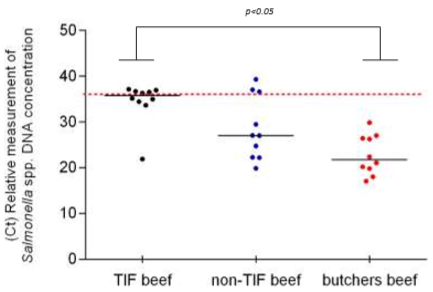
Figure 2 Distribution of Ct values obtained in TIF and non-TIF beef to identify Salmonella spp. The value of the median is shown in each group (n = 10). Ct value is inversely proportional to the ammount of Salmonella DNA found in the samples. Data were compared with Kruskal-Wallis and Dunn’s tests.
On the other hand, the median of non-TIF beef distributed in supermarkets was 27.03 and only 3 samples were above the cut-off point (negative samples). The 10 samples from local butchers were below the cut-off point, 100 % of them positive to Salmonella spp., and the median was 21.69 (Figure 2).
A study by Narváez-Bravo et al. (2013) reported the presence of S. enterica in 49.2 % of pre-gutted beef but only in 6 % of it by the end of the process. This stresses the relevance of hygiene measures implemented at slaughterhouses. On the other hand, Pérez-Montaño (2012) analyzed 505 samples of bovine carcasses and reported the presence of Salmonella spp. in 15.5 % of them. The most abundant serotypes were S. Give and S. Typhimurium in 24.4 % and 17.9 % of the samples, respectively. In addition, Hernández-San Juan (2007) reported the presence of S. enterica in 13.8 % of the analyzed bovine carcasses. The three studies above mentioned demonstrate that beef commercialized in Mexico is often contaminated with Salmonella, infringing the regulations established by SENASICA.
In Mexico, COFEPRIS is the government committee in charge of monitoring Salmonella spp. at points of sale. This organization has determined in its normativity that beef products ready to sell must be negative to Salmonella (NOM-213-SSA1-2002; NOM-114-SSA1-1994). However, given the socioeconomic and cultural conditions in Mexico, these regulations are difficult to implement. In a study published by Parrilla et al. (2014) on beef products, the presence of samples positive to Salmonella was 2.5 % while contaminated samples reached 36 %. The most common S. enterica serotypes were: Derby (29.6 %), London (16.8 %), Give (7.3 %), Anatum (7.0 %), Agona (6.7 %), and Infanti (5.1 %).
Additionally, several research groups have carried out microbiological monitoring of beef commercialized in several states of Mexico, as Tabasco, Mexico City, Hidalgo, Jalisco, and Nuevo Leon. Samples positive to Salmonella spp. were reported to reach 1.2 % - 30 % (Esquivel-Hernández & Nava-Morales, 2017).
Besides the probable health risks that beef contamination poses, there is a lack of international consensus on the methods to detect the microorganism. In this context, manual or automatized microbiological methods (based on biochemical profile samples) are used for routine determinations in Mexico, even though molecular methods based on the identification of gene fragments have proven to be more sensitive and specific. This has been demonstrated in the study by Yañez et al. (2008), which reports up to 20 % difference in detection limits between qPCR and a traditional method. The molecular method yielded results in 24 h, while results were obtained in 4 days following the traditional method. Also, molecular strategies allow for developing methods based on multiplex PCR, for the simultaneous detection and quantification of several pathogens in a faster way (García-López et al., 2009).
According to the results obtained in the present research work, the degree of contamination with Salmonella spp. in TIF beef was lower, both in number of samples and Ct values, as compared against the other two groups. Alarmingly, 50 % of the TIF samples were contaminated with Salmonella, which indicates that consumption of TIF beef does not ensure security and safety of the products. However, this investigation cannot identify whether the contamination with Salmonella spp. in this type of beef originates from the TIF slaughterhouse or the final handling of the product.











 texto en
texto en 


