1. Introduction
Childhood acute lymphoblastic leukemia (ALL) is the most common type of cancer and the second leading cause of mortality among Mexican children.1 Overall cure rates of newly diagnosed ALL patients is around 80%, while chemoresistance is one of the main challenges in the relapsed population.2-4 (3) Although much progress has been achieved in the detailed characterization of the molecular processes directly involved in drug tolerance of cancer cells,5,6 more research is needed to develop effective therapeutic strategies to resensitize chemoresistant cells.6
Vincristine is a vinca alkaloid, which interacts with tubulin disrupting microtubule polymerization and favoring the cell cycle arrest in the M phase which is followed by induction of apoptosis.7 It is used in several stages of the ALL treatment.3 It has been reported that the P-glycoprotein MDR1 actively pumps vincristine outside the cell reducing its therapeutic effect.8 Inactivation of intracellular vincristine by the myeloperoxidase has also been reported to contribute to resistance in some types of leukemia.9 Moreover, resistant leukemia, and other types of cancer cells, commonly show deregulated apoptosis4,6,10 and signaling pathways involved in survival.5,6,11 The complex state of resistance is the result of the concerted action of multiple interacting genes, proteins, and metabolites; and this scenario is well suited for characterization with the omic technologies. Proteomics studies provide an overview of the changes in the relative abundance of the proteins of a cell or tissue under different conditions.12 The characterization of proteins, as the final executors of cellular activities, may lead to identification of putative therapeutic targets and a better understanding of the pathological states.13 In the present work, we describe the changes in the proteome of a B-ALL cell line after adaptation to vincristine. Our results allowed the identification of targetable signaling and metabolic steps which may represent potential targets to resensitize leukemic cells to vincristine.
2. Methods
2.1. Growth conditions
The B-lineage pediatric ALL cell line CCRF-SB (ATCC CCL-120) was grown in RPMI-1640 with 10% (v/v) fetal bovine serum, 100 U/ml penicillin, 0.1 mg/ml streptomycin, 1% sodium pyruvate at 37 ◦C under an atmosphere with 5% CO2.
2.2. Vincristine exposition
Cell viability was estimated with the MTT assay in 96 well plates.14 To determine the IC50, vincristine was added to 7 x104 cells in 100 L of media per well at 0, 1, 2, 3, 4, 5, 6, 8, 10, 20 and 40 nM in triplicate for 48 hours (Figure 1A). Gradual exposition was made as follows: 3 x106 cells were cultured in 5 ml of media in the presence of 1 nM vincristine for five days, and then were exposed to 2 nM for the next five days, 3.5 nM for five days, 4.5 nM for three days, and 6 nM for four days (Figure 1B). In every step viable cells were enriched by centrifugation at 800 rpm 5 min, only the pelleted cells were transferred to the subsequent drug concentration. After 22 days of gradual exposition, cells proliferated in the presence of 6 nM vincristine, and cell viability was higher than 70%. Control cells were subjected to the same manipulations but cultured in the absence of vincristine.
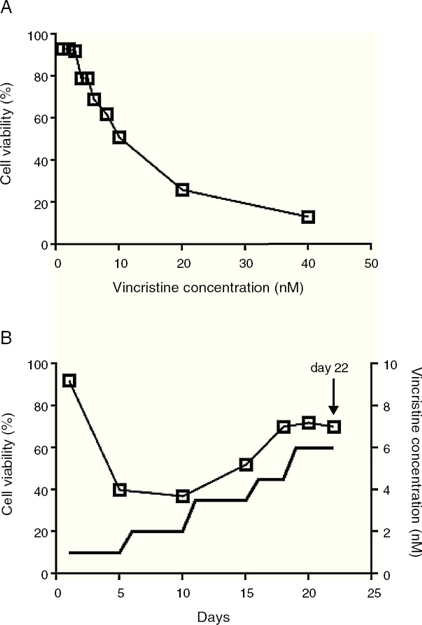
Figure 1 The CCRF-SB cell line is sensitive to vincristine. A. To obtain the IC50 (10 nM), cells were exposed to growing concentrations of vincristine for 48 h. B. Cell viability (□) and vincristine concentration (—) during the gradual adaptation protocol. The arrow shows the time point at which cells were harvested for proteomic studies.
2.3. Proteomic methods
Cells were washed with cold PBS and resuspended in 500 μL lysis buffer (4% SDS, 100 mM DTT, 100 mM TRIS pH 8.6 and a protease/phosphatase inhibitor cocktail, Thermo Scientific). Complete cell disruption was achieved by 20 cycles of 2 seconds sonication at 50% amplitude in a sonics 130-Watt Ultrasonic Processor and 5 seconds ice. After reduction (30 min at 40 ◦C) and alkylation (200 mM iodoacetamide, 30 min at room temperature in the darkness), 50 g of protein were precipitated overnight with nine volumes of ethanol at -20 ◦C and washed twice with 90% ethanol. The dried pellet was dissolved in 50 mM guanidinium chloride, 20 mM TRIS and digested with mass spectrometry grade trypsin (1:50, Promega) overnight at 37 ◦C. The resulting peptides were desalted (sep-pak C18, cartridges, Waters), dried and kept at -80 ◦C until used.
Just before injection, peptides were dissolved in 50 L of 0.1% formic acid. 5 L of each sample were injected in triplicate to an Ultimate 3000 nanoHPLC system with a 1 cm Acclaim PepMap100 C18 pre-column and a 50 cm Acclaim PepMap100 C18 column. Peptides were eluted with a non-linear gradient from 5% to 90% acetonitrile with 0.1% formic acid in 240 min. The HPLC system was coupled to the ESI-Ion Trap mass spectrometer Amazon speed (Bruker Daltonics) operated in positive mode, 400-1400 m/z, the 20 most abundant ions were fragmented every 50 msec, single charged ions were excluded. The ion list was constructed with the Data Analysis software (Bruker Daltonics) and used for subsequent protein identification with the Mascot search engine using the SwissProt human database, two missed trypsin cleavages, carbamidomethylation as fixed modification, methionine oxidation as variable modification, and a peptide mass tolerance of 0.6 Da. Only those proteins identified in the three replicates were considered as valid identifications. Venn diagrams were drawn with the website from the University of Southern Mississippi (http://genevenn.sourceforge.net/) and functional categories were analyzed with the statistical overrepresentation test of the PANTHER classification system15 (http://pantherdb.org) in which the analyzed list contained those proteins detected exclusively in the presence of vincristine whereas the reference list contained the proteins detected in both groups (with and without vincristine), the Bonferroni correction for multiple testing was not used. Only the functional categories with a fold enrichment value equal or higher than three were considered for further analysis.
3. Results
3.1. After gradual adaptation, the CCRF-SB cell line grew in the presence of 6 nM vincristine
Sudden vincristine exposition for 48 h resulted in IC50 of 10 nM (Figure 1A), which is similar to previous reports using other types of sensitive cell lines.9,16,17 However, gradual exposition allowed growth and cell duplication at 6 nM. The adaptation procedure depicted in Figure 1B resulted to be highly reproducible and allowed that cell density doubled during the last four days (from about 6x105 to 1.5 x106 cells/ml) keeping cell viability above 70%. These adapted cells were used for proteomic studies.
3.2. Approximately 15% of the proteome is differentially expressed after adaptation to vincristine
The proteomic approach used resulted in 849 and 858 valid identifications in the control and adapted cells respectively (Figure 2). As expected, most of the proteins (approximately 85%) were present in both conditions. However, 126 proteins were detected exclusively in the absence of vincristine and 135 in the presence of this drug. This result indicated that during the adaptation process, significant changes in the proteome took place.
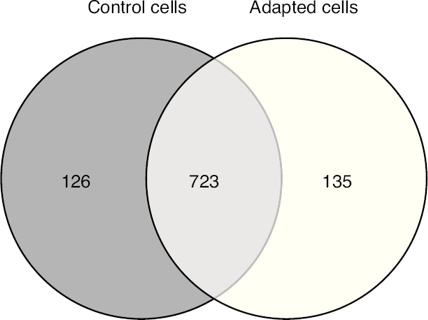
Figure 2 Venn diagram of valid protein identications with and without gradual exposition of the CCRF-SB cell line to 6 nM vincristine.
To identify the cellular processes that mainly contributed to drug tolerance, the statistical overrepresentation test was made using the PANTHER classification system15. The 135 proteins identified exclusively in the presence of vincristine were compared with the 723 proteins common to both groups (Figure 2). This resulted in the identification of 17 overrepresented pathways and functions (Figure 3) to which 30 different proteins were assigned (Table 1). Interestingly, 43% of the proteins in Table 1 and 53% of the pathways in Figure 3 were directly involved in signal transduction, whereas 13% of the proteins were involved in regulation of signaling and 20% in oxidative phosphorylation.
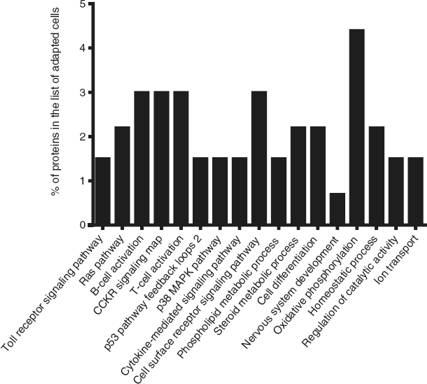
Figure 3 Functional categories overrepresented in adapted cells. The analysis was made with the statistical overrepresentation test of the PANTHER classication system as described in the Proteomic methods section.
4. Discussion
This study addressed the question of which are the cellular processes that sensitive leukemic cells induced to achieve tolerance to vincristine. To this end, the B-ALL cell line CCRF-SB was gradually exposed until cell proliferation was observed in the presence of 6 nM vincristine, and the corresponding proteomic profile was compared to that of cells grown in the absence of the chemotherapeutic drug.
Chemoresistance may be intrinsic or acquired.18 The ability to tolerate high concentrations of chemotherapeutics of an intrinsically resistant cancer cell is not developed as a result of an exposition to the drugs; instead it is the result of genetic abnormalities the cell carries before exposition.18 By contrast, acquired chemoresistance is developed after the cancer cell is exposed to the drug and may be the result of molecular evolution of resistant clones.19 Experimental settings to study acquired chemoresistance include the comparison of matched paired samples at diagnosis and after relapse20 or the comparison of sensitive cell lines and resistant sublines which are obtained after prolonged exposure (several months) to the drug.21 It is likely that resistant clones may have evolved during the gradual adaptation protocol depicted in Figure 1B. However, the low growth rate, the short time of exposure (22 days) and the fact that it was highly reproducible when it was assayed from frozen stocks of sensitive cells suggest that the observed tolerance to vincristine was the result of induction of cellular processes to counteract the cytotoxic effect of the drug rather than the result of events involving mutation, selection, and evolution of resistant clones.
The cytotoxic effects of drugs are dependent on the concentration and the time of exposure. In many studies, cells have been exposed to relatively high concentrations for short times (24 or 48 h). In such experimental setting, at concentrations close to the IC50, cells are not growing actively but remain viable and quiescent. Our results showed that even at concentration as low as 1 nM for five days, a reduction in cell viability was observed (Figure 1B). The gradual adaptation protocol used allowed cell division even after four days of exposition to 6 nM. By contrast, in cells exposed to 6 nM vincristine for 48 h without previous adaptation, no cellular division was observed (data not shown). In this regard, our results may be complementary to previous studies and may reflect some aspects of leukemic cells in vivo during advanced stages of therapy.
Successful adaptation was also reflected in the absence of overrepresented categories typically associated with cell death (Figure 3).
Interestingly, 43% of the proteins included in the over-represented categories of adapted cells (namely MP2K1, MK14, RB1, RAC1, VAV, CASP3, ZAP70, VRK1, IMA4, IMA3, RAP1A, PIPSL, and STK25) were involved in signaling pathways like Toll receptor signaling pathway, Ras pathway, B- and T-cell activation, CCKR signaling map, p53 pathway feedback loops, p38 MAPK pathway, cytokine-mediated signaling pathway, cell surface receptor signaling pathway (Table 1); some of them have been reported to be important for growth and proliferation of lymphocytes and chemoresistance.22-24 (23) For example, the MP2K1, MK14 and RAC1 proteins positively regulate the Ras pathway, and the MP2K1 protein is an essential component of the MAP kinase pathway. These signal transduction pathways have been associated with tumorigenesis and chemoresistance.22,25,26 Inhibition of the MP2K1 protein with PD-0325901, BAY86-9766, trametinib, selumetinib, pimasertib or GDC-0973 has been studied in advanced clinical trials.22 The MK14 protein has been reported to be closely associated with drug resistance in leukemia,27 and the RAC1 protein has been proposed as a therapeutic target for gefitinib-resistant non-small-cell lung cancer and pancreatic cancer cells,28-30 (29) and its overexpression has been associated with poor prognosis in non-small-cell lung cancer.30 Deregulation of the VAV protein has been reported in neuroblastoma, melanoma, pancreatic, lung and breast cancers, and B-cell chronic lymphocytic leukemia.31 Furthermore, the proteins ZYX, AT2A2, CK5P3 and OXSR1, which were assigned to categories like Nervous system development, Homeostatic process and Regulation of catalytic activity (Table 1), are modulators of signaling proteins.32,33 By contrast, in the control cells, any pathway resulted to be overrepresented when the statistical over-representation test was made comparing the 126 proteins detected exclusively in the absence of vincristine with the proteins detected in both groups (with and without vincristine). This difference may indicate that some signaling proteins and pathways may be crucial for adaptation to vincristine. In this regard, some drugs have been developed to target signaling pathways and to overcome drug tolerance.22,25,34 Further research is necessary to explore this possibility during the gradual adaptation to vincristine.
The cell-adhesion and mechanotransducer protein ZYX was assigned to the Nervous system development category (Table 1). However, it has been reported to participate in signal transduction pathways leading to cell migration, proliferation, and tumourigenesis.32 The ZYX protein is upregulated in human breast cancer and positively correlates with metastasis. It has been proposed as a potential therapeutic target in breast cancer and as an early detection biomarker in non-small cell lung cancer.32,35
Verrills et al. reported the proteomic changes of the T-lineage ALL cell line CCRF-CEM after exposition to 2, 4 and 8 nM vincristine for 24 h using two-dimensional polyacrylamide electrophoresis and MALDI-TOF mass spectrometry.17 They found that vincristine-induced changes in the expression of 39 proteins and that a resistant subline differentially expressed 42 proteins mainly involved in cytoskeleton metabolism.17 Other differentially expressed proteins resulted to be regulators of apoptosis, chaperones, gene regulators and ribosomal proteins. The involvement of alterations in the metabolism of actin and tubulin during the cellular response to vincristine was confirmed in a subsequent study by the same group using primary cells engrafted into NOD/SCID mice and intraperitoneal vincristine (0.5 mg/kg every seven days).36 Results shown in Figure 3 and Table 1 were based in comparisons of those proteins identified exclusively in the adapted cells versus the shared proteins (Figure 2). As cytoskeletal metabolism is very active in every cell, many cytoskeletal regulating proteins were among the 723 shared proteins; this may have prevented the overrepresentation of cytoskeletal metabolism in the adapted cells. To detect changes in the relative abundance of the shared proteins, a quantitative analysis is required. However, the microtubule-associated protein 4 (MAP4) was assigned to the overrepresented pathway Cell differentiation (Table 1). This protein promotes microtubule assembly and may reflect a more active cytoskeletal metabolism in the adapted cells compared to the control cells. Further research is necessary to explore this possibility.
After the combined effect of signaling pathways, Oxidative phosphorylation had the second highest percentage of proteins among the overrepresented pathways (Table 1). This suggested that a higher ATP production is needed to deal with the presence of vincristine. In this regard, modulation of the metabolic peculiarities of cancer cells has been proposed as a promising therapeutic strategy.37,38 The inhibitors metformin, rotenone, -tocopheryl succinate, benzylisothiocyanate, oligomycin and resveratrol have been used to target mitochondrial energetic metabolism.38 Our results suggest that these compounds may be good candidates to overcome tolerance to vincristine.
Detection of the TFR1 and THTM proteins resulted in the overrepresentation of the Ion transport category in the adapted cells (Table 1 and Figure 3). The protein TFR1 may have contributed to B-cell growth through iron uptake.39 Interestingly, this protein has been implicated in resistance to tamoxifen in a subgroup of ER+/luminal-like breast cancer.40 Calcium is translocated from the cytosol to the sarcoplasmic reticulum lumen by the AT2A2 protein (assigned to the Homeostatic process category in Table 1). Inhibition of this protein has been proposed as a promising strategy to overcome multidrug-resistance in leukemic cells.33 Moreover, calcium signaling modulates the activity of the THTM protein which participates in the production of hydrogen sulfide (Table 1). Interestingly, this compound has been reported to be involved in chemoresistance of lung adenocarcinoma cells, and inhibition of hydrogen sulfide-producing enzymes has been proposed as a strategy to sensitize resistant cells.41
DNA repair protein RAD50 was assigned to the overrepresented category Homeostatic process (Table 1). As part of the MRN complex, this protein has been proposed as a predictor for poor prognosis and chemoresistance in gastric cancer.42
Unexpectedly, p-glycoprotein was not detected in our proteomic analysis of adapted cells. This may be due to the low relative abundance of this protein compared to other cellular proteins. To achieve higher sensitivity, peptide mixture can be fractionated in a previous HPLC run before the nanoHPLC-ESI-MS analysis.
Although monoculture of cell lines is a useful model for in vitro studies, the influence of the bone marrow microenvironment in chemoresistance43 cannot be studied. In this regard, the co-culture of bone marrow stromal cells and leukemic cells44 will certainly give more information of the involved mechanisms.
As a general rule, findings from the omic analyses must be confirmed with traditional techniques like Western blot, flow cytometry, ELISA or RT-PCR. Confirmation would be necessary only for those few genes of interest in a particular study. Although we used stringent criteria for protein identification in the MASCOT search engine, only those proteins identified in the three analytical replicates were considered as valid identifications, and only pathways with a fold enrichment value higher or equal than three were considered as overrepresented. Our results must be validated if they are to be used in further studies aimed to deepen into the molecular mechanisms involved in tolerance to vincristine.











 text new page (beta)
text new page (beta)



