Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Similars in
SciELO
Similars in
SciELO
Share
Revista mexicana de fitopatología
On-line version ISSN 2007-8080Print version ISSN 0185-3309
Rev. mex. fitopatol vol.36 n.1 Texcoco Jan./Apr. 2018
https://doi.org/10.18781/r.mex.fit.1707-2
Phytopathological notes
Variability and symptoms caused by Iris yellow spot virus in Nicotiana benthamiana
1 Posgrado en Fitosanidad-Fitopatología. Colegio de Postgraduados, Km 36.5 carretera México-Texcoco, C.P. 56230, Montecillo, Texcoco, Estado de México.
2 Campo Experimental Zacatepec, INIFAP. Km. 0.5 carretera Zacatepec-Galeana, C.P. 62780, Colonia Centro Zacatepec, Morelos.
3 Department of Plant Pathology and Nebraska Center for Virology, University of Nebraska-Lincoln, Lincoln, NE, USA.
In onion Allium cepa crops in the state of Morelos, Mexico, typical and severe symptoms associated with Iris yellow spot virus (IYSV) are observed. In this work the alterations caused by IYSV isolates from typical and severe symptoms in Nicotiana benthamiana, the differences in the N gene and their phylogeny were studied. Four typical and five severe isolates mechanically inoculated caused systemic infection. Severe isolates caused more severe symptoms in bioclimatic chamber. The N gene sequence of both isolates had 98-99% identity with the IYSV nucleoprotein and no changes in nucleotide sequence between them were observed. Both isolates were grouped with the IYSVBR genotype and had greater similarity with those reported in Canada and the United States.
Key words: Tospovirus; phylogeny; genomic changes
En cultivos de cebolla Allium cepa del estado de Morelos, México, se observan síntomas típicos y severos asociados a Iris yellow spot virus (IYSV). En esta investigación se estudiaron las alteraciones que ocasionan los aislamientos de IYSV procedentes de síntomas típicos y severos en Nicotiana benthamiana, las diferencias en el gen N y su filogenia. Cuatro aislamientos típicos y cinco severos inoculados mecánicamente causaron infección sistémica. En cámara bioclimática los aislamientos severos ocasionaron mayor severidad de síntomas. La secuencia del gen N de ambos aislamientos tuvo 98-99% de identidad con la nucleoproteína de IYSV y no se observaron cambios en la secuencia de nucleótidos entre ellos. Ambos aislamientos se agruparon con el genotipo IYSVBR y tuvieron mayor similitud con los reportados en Canadá y Estados Unidos.
Palabras clave: Tospovirus; filogenia; cambios genómicos
The most important viral disease that affects onion crops (Allium cepa) is yellow spot caused by the Iris yellow spot virus (IYSV) (Bunyaviridae:Tospovirus) (Kritzman et al., 2001), because it leads to major reductions in bulb size (Gent et al., 2004). Symptoms of IYSV on onion leaves and flower stems appear as elongated or diamond-shaped straw colored and dry-looking chlorotic spots, with or without green areas in the middle of the lesions (Gent et al., 2006). The IYSV genome consists of three negative-sense, single-strand RNA segments defined as large (L), medium (M) and small (S) (Bag et al., 2009, 2010). The S segment contains the N gene that encodes for capsid nucleoprotein that is commonly used to identify and classify tospoviruses, because of its degree of divergence (Pappu et al., 2006), as well as to study geographic distribution and genetic diversity (Iftikhar et al., 2014). IYSV is widely distributed worldwide (Gent et al., 2006) but in Mexico it has been reported since 2012 on onion crops in the state of Morelos, where it causes a disease known as “yellow spot” with 100% incidence and more than 90% severity (Ramírez et al., 2016). In Morelos, two types of symptoms associated with IYSV have been observed and, for this reason, the objective of this research was to describe the symptoms caused by both isolates on Nicotiana benthamiana, and to find out if there are differences in the N segment of the genome and its phylogenetic relation with other isolates. In the 2014-2015 cycle, onion plants showing symptoms of “yellow spot” associated with IYSV were collected in the municipalities of Ayala, Axochiapan, Emiliano Zapata, Jojutla, Puente de Ixtla, Tlaquiltenango, Xochitepec and Zacatepec in the state of Morelos, Mexico. Onion leaf tissue was classified based on the severity of the symptoms observed, frozen with liquid nitrogen and stored at -80 °C. Nicotiana benthamiana plants were mechanically inoculated (Mandal et al., 2006) with onion leaf tissue that tested positive for Iris yellow spot virus using RT-PCR and showed typical or severe symptoms. Two N. benthamiana plants were used as negative controls, but they were only rubbed with a buffer and carborundum (silicon carbide). A group of five plants were inoculated with each of the typical or severe isolates and kept in a bioclimatic chamber at 25 °C average temperature and 91% average relative humidity (RH), 77% minimum RH and 99% maximum RH, and a photoperiod of 16 h light and 8 h darkness. Another group of plants was kept in the greenhouse at 25 °C average temperature (13 °C and 39 °C minimum and maximum temperature, respectively) and 50% average RH (27% and 71%, minimum and maximum HR, respectively). The plants were observed every other day for three weeks to record the incubation period and type of symptoms. Total RNA was extracted from 0.1 g of leaf tissue of onions plants collected in the field that showed symptoms associated with IYSV, and from 0.25 g of mechanically inoculated N. benthamiana leaves; RNA was extracted from the latter 10, 15 and 20 days post inoculation (dpi) with TriReagent® (Ambion®), following the manufacturer’s protocol. The concentration and quality of the RNA was determined using a Nanodrop® (Thermo Fisher Scientific, Inc), and its integrity through electrophoresis on 1% agarose gel stained with ethidium bromide (0.5 µg/ml). The cDNA synthesis was performed using M-MLV Reverse Transcriptase (Promega®), according to the manufacturer’s instructions. Primers IYSV917L and IYSV56U were selected to detect IYSV on onion because they amplify a segment of 896 bp corresponding to the N gene (Robène et al., 2006). A region of ribosomal gene 18S was amplified as control for the PCR reaction (Zamboni et al., 2008). To determine whether there were differences in the nucleotide sequence between the two IYSV isolates inoculated into N. benthamiana, primers IYSV-KO4 (5’-CTTAACTAACACAAATACTG-3’) and IYSV-KO6 (5’-AGAGCAATCGAGGTATAAAAC-3’) were designed to amplify the N gene using the Vector NTI™ Suite 8.0 program and based on the AF001387 sequence of IYSV stored in the GenBank. The obtained sequences were edited with the Chromas Lite 2.0 and Vector NTI™ Suite 8.0 programs and deposited in the GenBank database (KX434621-434623, KX443600-443604, KX443598 and KX443599). Multiple alignment was performed with the ClustalW program, and the phylogenetic analysis following the Neighbor-joining method (2000 replications) by using the MEGA6 program (Tamura et al., 2013). Seventy-eight onion plants showing “yellow spot” symptoms associated with IYSV were collected at 14 sites of the eight sampled municipalities. The collected plants belonged to six onion varieties: Stratus (33.3%), Carta Blanca (24.4%), Florentina (10.2%), Cirrus (6.4%), Cal 214 (6.4%) and Joya (2.6%); it was not possible to identify the variety of the remaining 16.7% of samples. The plants were divided into two groups according to the symptoms they showed: a) typical symptoms in the form of elongated yellow-to-whitish dry spots with a chlorotic halo (Figura 1B), and b) severe symptoms consisting of white to yellow elongated and dry-looking spots with or without a necrotic halo, that in most cases covered the leaf (Figura 1C). Four typical isolates and five severe isolates were selected for analysis.

Figure 1 Symptoms associated with Iris yellow spot virus in onion. A. Asymptomatic plant (Pa). B. Typical symptom (St). C. Severe symptom (Ss). D. PCR products obtained using the IYSV917L and IYSV56U primers that amplify a segment of the N gen of IYSV. -: negative control. MP: 100 bp molecular weight marker.
Primers IYSV917L and IYSV56U amplified the expected fragment of ~ 900 bp in all the typical isolates, while in the case of severe isolates the amplicon was ~500 bp (Figure 1D). However, both shared 99% identity with the capsid nucleoprotein of the N segment of IYSV reported in the GenBank. These results show that the 900 and 500 bp amplicons are associated with typical or severe IYSV isolates, respectively. The severity of the symptoms induced by typical and severe IYSV isolates on N. benthamiana was different depending on the environmental conditions under which the plants were kept after inoculation, regardless of the inoculum source. When kept in a bioclimatic chamber, at 7 dpi the symptoms induced by four of the five severe isolates were systemic and consisted of necrotic leaf venation, petiole constriction, necrotic leaf lesions surrounded by a dark brown halo, chlorosis and wilting. At 22 dpi, severe wilting was observed similar to that reported by Bag et al. (2012) (Figure 2B). In the case of the typical isolates, one produced chlorotic and necrotic lesions with necrotic veins and petioles; two more produced chlorotic and necrotic lesions but did not affect veins and petioles; the fourth isolate did not induce symptoms. On the other hand, when kept in the greenhouse, four severe and two typical isolates produced systemic infection at 10 dpi in the form of chlorosis and leaf wrinkling. One typical and one severe isolate caused more necrotic lesions on the leaves (Figure 2B). The fact that not all the typical or severe isolates caused the same symptoms (or that some did not even cause visible alterations) during that evaluation period can be attributed to the mechanical inoculation process, because it is known that the mechanical transmission of IYSV is difficult (Bag and Pappu, 2009). In spite of the inconsistency of the first symptoms observed, in the greenhouse all the inoculated plants showed wilting at 30-35 dpi regardless of the isolate. On the other hand, in a study using Tobacco mosaic virus strains inoculated in Nicotiana tabacum, differences in symptoms associated with temperature were observed (Scholthof, 2008). It is known that the environment can naturally contribute to increasing or decreasing the frequency of virus populations (García et al., 2001). In the case of IYSV, no specific environmental conditions account for symptom variations. However, the genetic divergence among isolates, the host range and host response has been attributed to environmental adaptation (Pozzer et al., 1999).
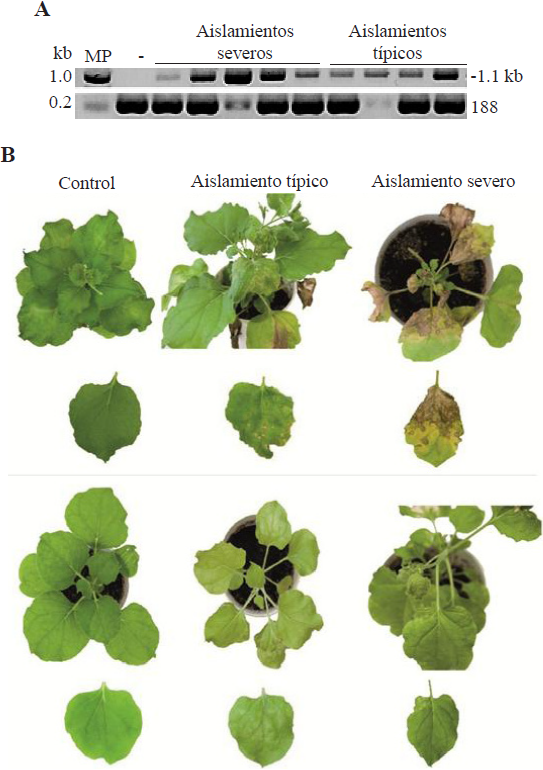

Figure 2 Figure 2. A. PCR products obtained using the IYSV-KO4 and IYSV-KO6 primers that amplify the N + UTR5’ gen from typical and severe IYSV isolates. MP: Molecular weight marker 1 kb (Promega), -: negative control asymptomatic plant. B. Symptoms in N. benthamiana plants and leaves mechanically inoculated with two Iris yellow spot virus isolates kept under two environmental conditions.
In all the typical and severe isolates analyzed in this research using RT-PCR and primers IYSV-KO4 and IYSV-KO6, the expected fragment of 1,100 bp was found that includes the N + UTR5’ gene (Figure 2A). A comparison of the sequences obtained from both isolates showed no differences between them; these results are similar to those reported by Bag et al. (2012). According to García et al. (2001), viral strains are groups of isolates with similar properties such as host range, transmission capacity and similar nucleotide sequences, among others. In Mexico, TSWV is the only tospovirus that has been biological and molecularly characterized. González (2014) studied three Mexican isolates of TSWV and found that the difference in severity may be associated with changes in the amino acids in the Nsm and NSs proteins, as well as those in the intergenic regions (IGR). This may be due to the fact that the tospovirus NSs protein acts a gene silencing suppressor (Ocampo et al., 2016), and, for this reason, it would be necessary to analyze other genes in molecular characterization studies. The sequences obtained with the IYSV-KO4 and IYSV-KO6 primers of the typical and severe isolates tested (Figure 2B) had 98 to 99% identity with the nucleoprotein of the capsid of IYSV gene N. The in silico analysis of the sequences obtained (data not shown) using the IYSV-KO4 and IYSV-KO6 primers of both isolates through RFLP with HinfI showed a restriction pattern that includes a fragment larger than 308 bp which corresponds to the IYSVBR genotype associated with isolates from Asia (Iftikhar et al., 2014). On the other hand, it was found that IYSV isolates from the state of Zacatecas corresponded to the IYSVNL genotype associated with isolates from North America (Velásquez-Valle et al., 2016). The presence of chlorotic lesions on leaves with small green islands in the middle has also been reported in Zacatecas and Michoacán (Velásquez-Valle et al., 2016; Ávila et al., 2017), while in Morelos there is no record of this type of symptom (Ramírez-Rojas et al., 2016). These results suggest that there are IYSV variants in Mexico that are biological and molecularly different. Regarding the phylogenetic analysis, isolates from Morelos were found to be more similar to those reported in Canada, United States and New Zealand and not to those from Asia, where the IYSVBR genotype is mainly concentrated. Iftikhar et al. (2014) mentioned in their study that there are exceptions where the IYSVBR genotypes are grouped with IYSVNL, and vice versa; however, those results may provide information on how the virus spreads across countries (Smith et al., 2006). Also, typical and severe isolates are concentrated in a clade that is divided into three groups: group I consists mainly of severe isolates, while groups II y III consist of moderate isolates (data not shown). Abad et al. (2003) and Bag et al. (2012) studied isolates from different states of western United States. In the first study, greater genetic diversity was found among isolates that were grouped with other isolates from different countries, while in the second study, the isolates were grouped in the same clade. Based on these results, the fact that different groups of IYSV isolates have developed in Zacatecas and Morelos suggests that there is greater genetic diversity of the virus in Mexico (Smith et al., 2006). Such diversity may affect the efficiency of virus transmission by vector thrips, the range of host weed species, the severity and incidence of plants with symptoms and, in general, the spatio-temporal disease progress in both states (Ávila et al., 2017).
In conclusion, the typical and severe Iris yellow spot virus isolates analyzed in the present research infected N. benthamiana and caused similar symptoms at different severity levels under two environmental conditions. The analysis of the N gene did not show changes in the sequence of nucleotides between both isolates.
Literatura citada
Abad JA, Speck J, Mohan SK and Moyer JW. 2003. Diversity of the Iris yellow spot virus N gene in the USA. Phytopathology 93: S1. Disponible en línea: https://apsjournals.apsnet.org/doi/pdf/10.1094/PHYTO.2003.93.6.S1 [ Links ]
Ávila-Alistac N, Ramírez-Rojas S, Lozoya-Saldaña H, Rebollar-Alviter A, Guzmán-Plazola RA. 2017. Alternate hosts of Iris yellow spot virus and trips on onion crops in Morelos and Michoacan, Mexico. Revista Mexicana de Fitopatología 35: 242-262. https://doi.org/10.18781/R.MEX.FIT.1701-1 [ Links ]
Bag S, Druffel KL and Pappu HR. 2010. Structure and genome organization of the large RNA of Iris yellow spot virus (genus Tospovirus, family Bunyaviridae). Archives of Virology 155:275-279. https://doi.org/10.1007/s00705-009-0568-5 [ Links ]
Bag S, Druffel KL, Salewsky T and Pappu HR. 2009. Nucleotide sequence and genome organization of the medium RNA of Iris yellow spot virus from the United States. Archives of Virology 154:715-718. https://doi.org/10.1007/s00705-009-0349-1 [ Links ]
Bag S, Schwartz HF and Pappu HR. 2012. Identification and characterization of biologically distinct isolates of Iris ye-llow spot virus (genus Tospovirus, family Bunyaviridae), a serious pathogen of onion. European Journal of Plant Pathology 134: 97-104. https://doi.org/10.1007/s10658-012-0026-1 [ Links ]
Bag S and Pappu HR. 2009. Symptomatology of Iris yellow spot virus in selected indicator hosts. Online. Plant Health Progress. https://doi.org/10.1094/PHP-2009-0824-01-BR [ Links ]
García-Arenal F, Fraile A and Malpica JM. 2001. Variability and genetic structure of plant virus populations. Annual Review of Phytopathology 39: 157-86. Disponible en línea: http://www.annualreviews.org/doi/pdf/10.1146/annu-rev.phyto.39.1.157 [ Links ]
Gent DH, du Toit LJ, Fichtner SF, Mohan SK, Pappu HR and Schwartz HF. 2006. Iris yellow spot virus: An emerging threat to onion bulb and seed production. Plant Disease 90: 1468-1480. https://doi.org/10.1094/PD-90-1468 [ Links ]
Gent DH, Schwartz HF and Khosla R. 2004. Distribution and incidence of Iris yellow spot virus in Colorado and its relation to onion plant population and yield. Plant Disease 88: 446-452. https://doi.org/10.1094/PDIS.2004.88.5.446 [ Links ]
González PBE. 2014. Caracterización biológica y molecular del Tomato spotted wilt virus (TSWV) en aislamientos mexicanos. Tesis Doctoral. Instituto de Fitosanidad-Fitopalogía. Colegio de Postgraduados. Disponible en línea: http://colposdigital.colpos.mx:8080/jspui/handle/10521/2271 [ Links ]
Iftikhar R, Bag S, Ashfaq M and Pappu HR. 2014. First report of Iris yellow spot virus infecting onion in Pakistan. Plant Disease 97: 1517. https://doi.org/10.1094/PDIS-05-13-0502-PDN [ Links ]
Kritzman A, Lampel M, Raccah B and Gera A. 2001. Distribution and transmission of Iris yellow spot virus. Plant Disease 85: 838-842. https://doi.org/10.1094/PDIS.2001.85.8.838 [ Links ]
Mandal B, Pappu HR, Csinos AS and Culbreath AK. 2006. Response of peanut, pepper, tobacco, and tomato cultivars to two biologically distinct isolates of Tomato spotted wilt virus. Plant Disease 90: 1150-1155. http://dx.doi.org/10.1094/PD-90-1150 [ Links ]
Ocampo OT, Gabriel PSM, Bacheller N, Uiterwaal S, Knapp A, Hennen A, Ochoa-Martinez DL and Garcia-Ruiz H. 2016. Antiviral RNA silencing suppression activity of Tomato spotted wilt virus NSs protein. Genetics and Molecular Research 15(2): gmr.15028625. DOI: 10.4238/gmr.15028625 [ Links ]
Pappu HR, duToit LJ, Schwartz HF and Mohan SK. 2006. Sequence diversity of the nucleoprotein gene of Iris yellow spot virus (genus Tospovirus, family Bunyaviridae) isolates from the western region of the United States. Archives of Virology 151: 1015-1023. http://dx.doi.org/10.1007/s00705-005-0681-z [ Links ]
Pozzer L, Bezerra IC, Kormelink R, Prins M, Peters D, Resende RO and de Ávila AC. 1999. Characterization of a tospovirus isolate of Iris yellow spot virus associated with a disease in onion fields in Brazil. Plant Disease 83: 345-350. http://dx.doi.org/10.1094/PDIS.1999.83.4.345 [ Links ]
Ramírez-Rojas S, Ornelas-Ocampo K, Osuna-Canizalez FJ, Bartolo-Reyes JC, Varela-Loza V, Hernández-Romano J y Ochoa-Martínez DL. 2016. Detection of Iris yellow spot virus in onion plants from Tepalcingo, Morelos state, Mexico. Revista Mexicana de Fitopatología 34: 309-315. http://dx.doi.org/10.18781/R.MEX.FIT.1604-1 [ Links ]
Robène-Soustrade I, Hostachy B, Roux-Cuvelier M, Minatchy J, Hédont M, Pallas R, Couteau A, Cassam N and Wuster G. 2006. First report of Iris yellow spot virus in onion bulb- and seed-production fields in Réunion Island. Plant Pathology 55: 288. http://dx.doi.org/10.1111/j.1365-3059.2005.01262.x [ Links ]
Scholthof KBG. 2008. Tobacco Mosaic Virus: The Beginning of Plant Pathology. Online. APSnet Features. DOI:10.1094/APSnetFeatures-2008-0408 [ Links ]
Smith TN, Jones RAC and Wylie SJ. 2006. Genetic diversity of the nucleocapsid gene of Iris yellow spot virus. Australian Plant Pathology 35: 359. http://dx.doi.org/10.1071/AP06031 [ Links ]
Tamura K, Stecher G, Peterson D, Filipski A and Kumar S. 2013. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Molecular Biology and Evolution 30: 2725-2729. http://dx.doi.org/10.1093/molbev/mst197 [ Links ]
Velásquez-Valle R, Reveles-Torres LR, Salas-Muñoz S, Mauricio-Castillo JA and Pappu HR. 2016. First confirmed report of Iris yellow spot virus in onion nurseries in Zacatecas, Mexico. Plant Disease 100: 1509. http://dx.doi.org/10.1094/PDIS-01-16-0061-PDN [ Links ]
Zamboni A, Pierantoni L and De Franceschi P. 2008. Total RNA extraction from strawberry tree (Arbutus unedo) and several other woody plants. iForest-Biogeosciences and Forestry 1: 122-125. http://dx.doi.org/10.3832/ifor0465-0010122. [ Links ]
Received: July 13, 2017; Accepted: October 05, 2017











 text in
text in 


